Project
ENCODE: H3K4me3 ChIP-Seq of primary human cells
Navigation
Downloads
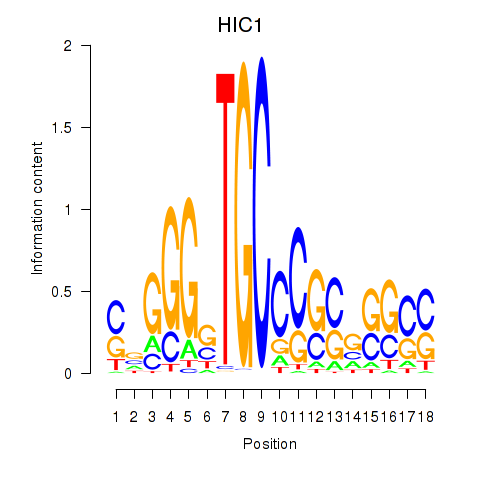
Results for HIC1
Z-value: 2.51
Transcription factors associated with HIC1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HIC1
|
ENSG00000177374.8 | HIC ZBTB transcriptional repressor 1 |
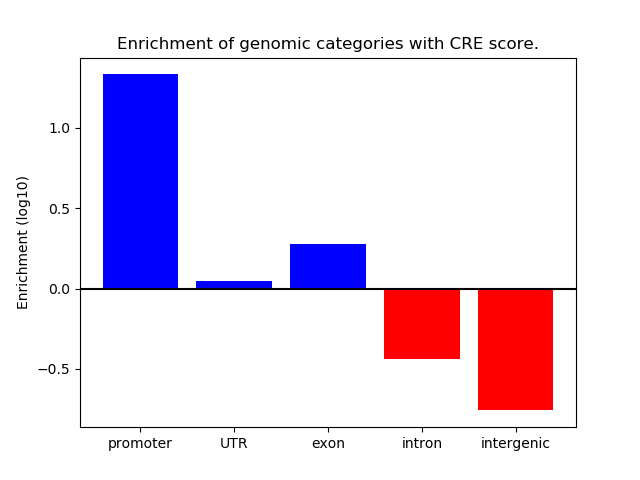
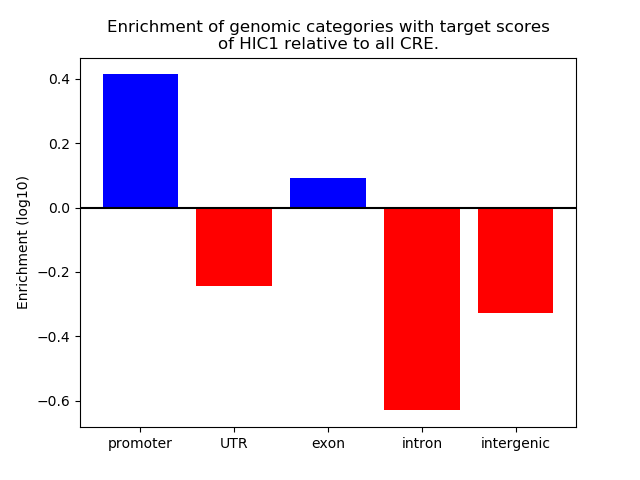
Correlations of motif activity and signal intensity at CREs associated with the motif's TFs:
This plot shows correlation between observed signal intensity of a CRE associated with the transcription factor across all samples and activity of the motif.
For each TF, only the top 5 correlated CREs are shown.
| CRE | Gene | Distance | Association probability | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|---|---|
| chr17_1956777_1956953 | HIC1 | 583 | 0.520406 | -0.35 | 3.6e-01 | Click! |
| chr17_1961527_1961722 | HIC1 | 2020 | 0.156240 | -0.24 | 5.3e-01 | Click! |
| chr17_1960560_1960711 | HIC1 | 1031 | 0.310092 | -0.22 | 5.8e-01 | Click! |
| chr17_1956964_1957351 | HIC1 | 291 | 0.778601 | -0.21 | 5.8e-01 | Click! |
| chr17_1961906_1962057 | HIC1 | 2377 | 0.137114 | 0.20 | 6.1e-01 | Click! |
Activity of the HIC1 motif across conditions
Conditions sorted by the z-value of the HIC1 motif activity
Move your cursor over a bar to see sample name and corresponding Z-value.
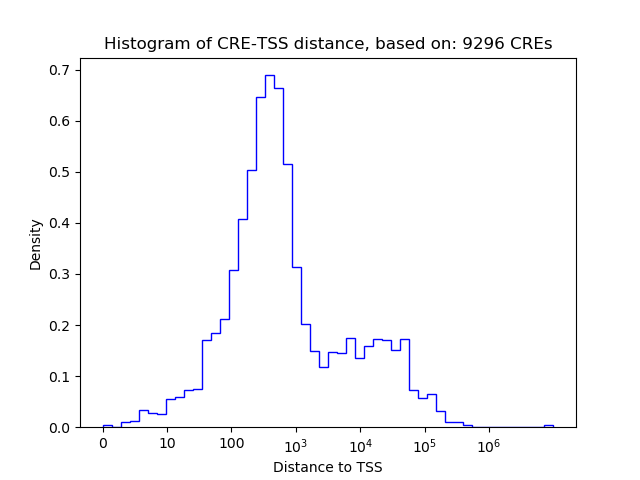
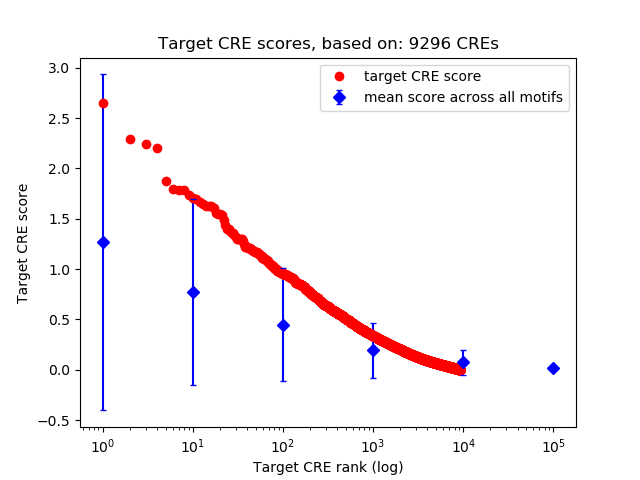
Top target CREs of the motif:
| Cis Regulatory Element (CRE) | Target Score | Top associated gene | Gene Info | Distance of CRE to TSS | CRE/Gene association probability |
|---|---|---|---|---|---|
| chr1_14924992_14926197 | 2.65 |
KAZN |
kazrin, periplakin interacting protein |
381 |
0.93 |
| chr16_85203170_85204140 | 2.29 |
CTC-786C10.1 |
|
1227 |
0.55 |
| chr20_55204347_55205219 | 2.25 |
TFAP2C |
transcription factor AP-2 gamma (activating enhancer binding protein 2 gamma) |
425 |
0.81 |
| chr8_99439239_99439631 | 2.21 |
KCNS2 |
potassium voltage-gated channel, delayed-rectifier, subfamily S, member 2 |
185 |
0.95 |
| chr2_54086905_54087527 | 1.88 |
GPR75-ASB3 |
GPR75-ASB3 readthrough |
46 |
0.37 |
| chr12_3308931_3310293 | 1.80 |
TSPAN9 |
tetraspanin 9 |
729 |
0.75 |
| chr16_65155038_65155536 | 1.78 |
CDH11 |
cadherin 11, type 2, OB-cadherin (osteoblast) |
546 |
0.88 |
| chr2_160918347_160919586 | 1.78 |
PLA2R1 |
phospholipase A2 receptor 1, 180kDa |
155 |
0.98 |
| chr11_94473290_94474384 | 1.73 |
RP11-867G2.8 |
|
316 |
0.92 |
| chr13_110958781_110959436 | 1.71 |
COL4A1 |
collagen, type IV, alpha 1 |
370 |
0.6 |
| chr1_39980649_39981517 | 1.69 |
RP11-69E11.4 |
|
9465 |
0.14 |
| chr3_8809480_8810320 | 1.66 |
OXTR |
oxytocin receptor |
463 |
0.76 |
| chr2_39893102_39893556 | 1.65 |
TMEM178A |
transmembrane protein 178A |
270 |
0.95 |
| chr6_123317343_123317959 | 1.63 |
CLVS2 |
clavesin 2 |
183 |
0.97 |
| chr11_70244216_70245100 | 1.63 |
CTTN |
cortactin |
11 |
0.53 |
| chr8_687260_687856 | 1.62 |
ERICH1-AS1 |
ERICH1 antisense RNA 1 |
93 |
0.97 |
| chr4_183369422_183369983 | 1.61 |
TENM3 |
teneurin transmembrane protein 3 |
450 |
0.89 |
| chr14_62279468_62280189 | 1.56 |
SNAPC1 |
small nuclear RNA activating complex, polypeptide 1, 43kDa |
50753 |
0.17 |
| chr19_14315927_14317134 | 1.55 |
LPHN1 |
latrophilin 1 |
451 |
0.77 |
| chr17_37856067_37857084 | 1.55 |
ERBB2 |
v-erb-b2 avian erythroblastic leukemia viral oncogene homolog 2 |
242 |
0.86 |
| chr16_14396121_14396821 | 1.54 |
ENSG00000207639 |
. |
1353 |
0.42 |
| chr16_2521208_2521831 | 1.49 |
NTN3 |
netrin 3 |
19 |
0.93 |
| chr17_64187141_64188685 | 1.44 |
CEP112 |
centrosomal protein 112kDa |
60 |
0.98 |
| chr2_96012468_96013004 | 1.40 |
KCNIP3 |
Kv channel interacting protein 3, calsenilin |
260 |
0.94 |
| chr11_119019412_119020504 | 1.40 |
ABCG4 |
ATP-binding cassette, sub-family G (WHITE), member 4 |
5 |
0.94 |
| chr2_27717607_27718081 | 1.38 |
FNDC4 |
fibronectin type III domain containing 4 |
268 |
0.82 |
| chr13_96296247_96296898 | 1.36 |
DZIP1 |
DAZ interacting zinc finger protein 1 |
372 |
0.89 |
| chr17_43483002_43483407 | 1.36 |
ARHGAP27 |
Rho GTPase activating protein 27 |
96 |
0.95 |
| chr3_64430574_64431219 | 1.34 |
PRICKLE2 |
prickle homolog 2 (Drosophila) |
162 |
0.97 |
| chr20_39994731_39995211 | 1.32 |
EMILIN3 |
elastin microfibril interfacer 3 |
496 |
0.81 |
| chr8_49782515_49783288 | 1.30 |
SNAI2 |
snail family zinc finger 2 |
51087 |
0.18 |
| chr11_75378366_75379067 | 1.30 |
MAP6 |
microtubule-associated protein 6 |
763 |
0.58 |
| chr17_29036450_29037561 | 1.30 |
ENSG00000241631 |
. |
7212 |
0.17 |
| chr16_86542600_86543872 | 1.30 |
FOXF1 |
forkhead box F1 |
897 |
0.62 |
| chr4_187644173_187644789 | 1.30 |
FAT1 |
FAT atypical cadherin 1 |
528 |
0.87 |
| chr4_20255612_20255816 | 1.28 |
SLIT2 |
slit homolog 2 (Drosophila) |
275 |
0.96 |
| chr14_105830576_105830747 | 1.25 |
PACS2 |
phosphofurin acidic cluster sorting protein 2 |
18795 |
0.13 |
| chr9_71939475_71940274 | 1.22 |
FAM189A2 |
family with sequence similarity 189, member A2 |
386 |
0.91 |
| chr2_242497891_242499073 | 1.22 |
BOK-AS1 |
BOK antisense RNA 1 |
90 |
0.79 |
| chrX_39548256_39548819 | 1.22 |
ENSG00000263730 |
. |
28067 |
0.24 |
| chr15_99640520_99640898 | 1.21 |
SYNM |
synemin, intermediate filament protein |
2289 |
0.24 |
| chr6_147829082_147829740 | 1.21 |
SAMD5 |
sterile alpha motif domain containing 5 |
652 |
0.84 |
| chr2_236403404_236403896 | 1.21 |
AGAP1 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 1 |
899 |
0.65 |
| chr5_7396416_7397106 | 1.20 |
ADCY2 |
adenylate cyclase 2 (brain) |
440 |
0.88 |
| chr9_35489468_35490886 | 1.19 |
RUSC2 |
RUN and SH3 domain containing 2 |
53 |
0.97 |
| chr6_46702576_46703631 | 1.19 |
PLA2G7 |
phospholipase A2, group VII (platelet-activating factor acetylhydrolase, plasma) |
24 |
0.97 |
| chr12_312502_313284 | 1.18 |
RP11-283I3.2 |
|
85 |
0.95 |
| chr17_9018882_9019405 | 1.18 |
NTN1 |
netrin 1 |
47109 |
0.15 |
| chr15_51633704_51634417 | 1.17 |
GLDN |
gliomedin |
234 |
0.91 |
| chr20_6032553_6033205 | 1.17 |
LRRN4 |
leucine rich repeat neuronal 4 |
1816 |
0.35 |
| chr14_101034010_101035263 | 1.17 |
BEGAIN |
brain-enriched guanylate kinase-associated |
192 |
0.93 |
| chr16_73082076_73082698 | 1.16 |
ZFHX3 |
zinc finger homeobox 3 |
113 |
0.97 |
| chr16_15952283_15952487 | 1.16 |
MYH11 |
myosin, heavy chain 11, smooth muscle |
1495 |
0.35 |
| chr18_2906194_2907483 | 1.15 |
RP11-737O24.5 |
|
14126 |
0.16 |
| chr18_6729547_6730123 | 1.15 |
ARHGAP28 |
Rho GTPase activating protein 28 |
14 |
0.5 |
| chr8_8749576_8750657 | 1.14 |
MFHAS1 |
malignant fibrous histiocytoma amplified sequence 1 |
1039 |
0.53 |
| chr8_27491561_27492240 | 1.13 |
SCARA3 |
scavenger receptor class A, member 3 |
202 |
0.93 |
| chr20_13976647_13977145 | 1.13 |
SEL1L2 |
sel-1 suppressor of lin-12-like 2 (C. elegans) |
193 |
0.78 |
| chr4_119273101_119273655 | 1.11 |
PRSS12 |
protease, serine, 12 (neurotrypsin, motopsin) |
780 |
0.76 |
| chr2_9347772_9348251 | 1.11 |
ASAP2 |
ArfGAP with SH3 domain, ankyrin repeat and PH domain 2 |
1117 |
0.65 |
| chr2_159825200_159826543 | 1.11 |
TANC1 |
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 1 |
688 |
0.73 |
| chr10_64564066_64565253 | 1.10 |
ADO |
2-aminoethanethiol (cysteamine) dioxygenase |
143 |
0.92 |
| chr1_66258526_66259233 | 1.10 |
PDE4B |
phosphodiesterase 4B, cAMP-specific |
15 |
0.99 |
| chr1_48462552_48463006 | 1.10 |
TRABD2B |
TraB domain containing 2B |
212 |
0.96 |
| chr18_499765_500605 | 1.09 |
COLEC12 |
collectin sub-family member 12 |
537 |
0.79 |
| chr5_113697124_113698006 | 1.09 |
KCNN2 |
potassium intermediate/small conductance calcium-activated channel, subfamily N, member 2 |
451 |
0.9 |
| chr13_21295723_21296224 | 1.08 |
ENSG00000265710 |
. |
18131 |
0.18 |
| chr22_20119477_20120671 | 1.08 |
ZDHHC8 |
zinc finger, DHHC-type containing 8 |
603 |
0.53 |
| chr9_140500205_140501463 | 1.07 |
ARRDC1 |
arrestin domain containing 1 |
679 |
0.56 |
| chr20_30457693_30458808 | 1.06 |
DUSP15 |
dual specificity phosphatase 15 |
125 |
0.67 |
| chrX_8699533_8700279 | 1.06 |
KAL1 |
Kallmann syndrome 1 sequence |
321 |
0.94 |
| chr3_123168721_123169253 | 1.05 |
ADCY5 |
adenylate cyclase 5 |
382 |
0.9 |
| chr10_21462508_21463481 | 1.04 |
NEBL-AS1 |
NEBL antisense RNA 1 |
51 |
0.59 |
| chr7_32110576_32110834 | 1.04 |
PDE1C |
phosphodiesterase 1C, calmodulin-dependent 70kDa |
231 |
0.97 |
| chr17_74581180_74582274 | 1.04 |
ST6GALNAC2 |
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 2 |
483 |
0.62 |
| chr3_55520023_55520469 | 1.03 |
WNT5A |
wingless-type MMTV integration site family, member 5A |
1085 |
0.48 |
| chr4_6472295_6472644 | 1.03 |
PPP2R2C |
protein phosphatase 2, regulatory subunit B, gamma |
1704 |
0.48 |
| chr19_41195400_41196566 | 1.02 |
NUMBL |
numb homolog (Drosophila)-like |
44 |
0.96 |
| chr8_69242814_69243988 | 1.02 |
C8orf34 |
chromosome 8 open reading frame 34 |
56 |
0.82 |
| chr16_67282110_67283381 | 1.02 |
SLC9A5 |
solute carrier family 9, subfamily A (NHE5, cation proton antiporter 5), member 5 |
108 |
0.89 |
| chr1_205648469_205649627 | 1.02 |
SLC45A3 |
solute carrier family 45, member 3 |
539 |
0.74 |
| chr1_180881914_180882730 | 1.00 |
KIAA1614 |
KIAA1614 |
3 |
0.98 |
| chr7_29846053_29846716 | 1.00 |
WIPF3 |
WAS/WASL interacting protein family, member 3 |
282 |
0.94 |
| chr3_192126971_192127690 | 1.00 |
FGF12 |
fibroblast growth factor 12 |
492 |
0.87 |
| chr10_35102956_35103650 | 0.99 |
PARD3 |
par-3 family cell polarity regulator |
946 |
0.5 |
| chr9_35791682_35791961 | 0.99 |
NPR2 |
natriuretic peptide receptor B/guanylate cyclase B (atrionatriuretic peptide receptor B) |
330 |
0.75 |
| chr11_126872809_126873319 | 0.99 |
KIRREL3-AS3 |
KIRREL3 antisense RNA 3 |
259 |
0.89 |
| chr9_91793130_91793737 | 0.99 |
SHC3 |
SHC (Src homology 2 domain containing) transforming protein 3 |
249 |
0.95 |
| chr11_63530349_63531105 | 0.98 |
ENSG00000264519 |
. |
34816 |
0.12 |
| chr14_38063567_38064244 | 0.98 |
FOXA1 |
forkhead box A1 |
334 |
0.84 |
| chr5_37839777_37839928 | 0.98 |
GDNF |
glial cell derived neurotrophic factor |
64 |
0.98 |
| chr5_38845474_38846010 | 0.97 |
OSMR |
oncostatin M receptor |
218 |
0.96 |
| chr17_76164337_76165654 | 0.97 |
SYNGR2 |
synaptogyrin 2 |
244 |
0.87 |
| chr2_211089377_211090268 | 0.97 |
ACADL |
acyl-CoA dehydrogenase, long chain |
393 |
0.85 |
| chr6_88875424_88876640 | 0.97 |
CNR1 |
cannabinoid receptor 1 (brain) |
46 |
0.99 |
| chr5_128796390_128797012 | 0.96 |
ADAMTS19-AS1 |
ADAMTS19 antisense RNA 1 |
319 |
0.77 |
| chr1_34630625_34630811 | 0.96 |
CSMD2 |
CUB and Sushi multiple domains 2 |
157 |
0.94 |
| chr16_691284_692401 | 0.96 |
FAM195A |
family with sequence similarity 195, member A |
29 |
0.92 |
| chr5_76372662_76373578 | 0.96 |
ENSG00000238961 |
. |
3276 |
0.17 |
| chr14_73706275_73706778 | 0.95 |
PAPLN |
papilin, proteoglycan-like sulfated glycoprotein |
193 |
0.92 |
| chrX_100545955_100546357 | 0.95 |
TAF7L |
TAF7-like RNA polymerase II, TATA box binding protein (TBP)-associated factor, 50kDa |
168 |
0.93 |
| chr18_35146776_35147412 | 0.95 |
CELF4 |
CUGBP, Elav-like family member 4 |
1094 |
0.67 |
| chr9_127539201_127540134 | 0.95 |
OLFML2A |
olfactomedin-like 2A |
117 |
0.95 |
| chr13_44360523_44361050 | 0.95 |
ENOX1 |
ecto-NOX disulfide-thiol exchanger 1 |
258 |
0.95 |
| chr2_36583449_36584760 | 0.95 |
CRIM1 |
cysteine rich transmembrane BMP regulator 1 (chordin-like) |
490 |
0.89 |
| chr11_110582275_110583137 | 0.95 |
ARHGAP20 |
Rho GTPase activating protein 20 |
67 |
0.99 |
| chr1_167408487_167409140 | 0.95 |
RP11-104L21.2 |
|
19085 |
0.19 |
| chr11_27740485_27740934 | 0.95 |
BDNF |
brain-derived neurotrophic factor |
585 |
0.82 |
| chr6_87647008_87647409 | 0.95 |
HTR1E |
5-hydroxytryptamine (serotonin) receptor 1E, G protein-coupled |
184 |
0.97 |
| chr17_10101159_10101987 | 0.95 |
GAS7 |
growth arrest-specific 7 |
295 |
0.92 |
| chr11_118478413_118479735 | 0.94 |
PHLDB1 |
pleckstrin homology-like domain, family B, member 1 |
716 |
0.54 |
| chr19_8400013_8400372 | 0.94 |
KANK3 |
KN motif and ankyrin repeat domains 3 |
7954 |
0.1 |
| chr18_60263508_60264066 | 0.93 |
ENSG00000243549 |
. |
54047 |
0.13 |
| chr22_44727371_44727573 | 0.93 |
KIAA1644 |
KIAA1644 |
18741 |
0.22 |
| chr6_125282837_125283421 | 0.93 |
RNF217 |
ring finger protein 217 |
562 |
0.67 |
| chr16_51184635_51185066 | 0.93 |
SALL1 |
spalt-like transcription factor 1 |
302 |
0.89 |
| chr10_15411519_15412658 | 0.93 |
FAM171A1 |
family with sequence similarity 171, member A1 |
970 |
0.7 |
| chr12_31079520_31079921 | 0.92 |
TSPAN11 |
tetraspanin 11 |
38 |
0.99 |
| chr5_173738719_173739174 | 0.92 |
CTC-229L21.1 |
|
138842 |
0.05 |
| chr21_46825103_46825740 | 0.92 |
COL18A1 |
collagen, type XVIII, alpha 1 |
369 |
0.83 |
| chr4_75718841_75720131 | 0.92 |
BTC |
betacellulin |
410 |
0.91 |
| chr3_123752157_123752801 | 0.92 |
ENSG00000238512 |
. |
28452 |
0.16 |
| chr15_39873424_39874712 | 0.92 |
THBS1 |
thrombospondin 1 |
774 |
0.66 |
| chr4_55096373_55096991 | 0.92 |
PDGFRA |
platelet-derived growth factor receptor, alpha polypeptide |
193 |
0.97 |
| chr19_709173_709581 | 0.91 |
PALM |
paralemmin |
276 |
0.84 |
| chrX_110038687_110039773 | 0.91 |
CHRDL1 |
chordin-like 1 |
56 |
0.99 |
| chr6_46458824_46459774 | 0.91 |
RCAN2 |
regulator of calcineurin 2 |
200 |
0.78 |
| chr2_120189452_120190154 | 0.91 |
TMEM37 |
transmembrane protein 37 |
358 |
0.88 |
| chr6_78172796_78173983 | 0.91 |
HTR1B |
5-hydroxytryptamine (serotonin) receptor 1B, G protein-coupled |
101 |
0.99 |
| chr4_146403033_146403938 | 0.91 |
SMAD1 |
SMAD family member 1 |
153 |
0.96 |
| chr2_24232585_24233394 | 0.90 |
MFSD2B |
major facilitator superfamily domain containing 2B |
22 |
0.97 |
| chr7_94536809_94537187 | 0.90 |
PPP1R9A |
protein phosphatase 1, regulatory subunit 9A |
11 |
0.99 |
| chr7_149389592_149390001 | 0.89 |
KRBA1 |
KRAB-A domain containing 1 |
22076 |
0.2 |
| chr14_105452072_105453062 | 0.89 |
C14orf79 |
chromosome 14 open reading frame 79 |
437 |
0.8 |
| chr6_72129322_72130506 | 0.88 |
ENSG00000207827 |
. |
16590 |
0.22 |
| chr2_236401822_236402641 | 0.87 |
AGAP1 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 1 |
502 |
0.83 |
| chr9_14312608_14313176 | 0.87 |
NFIB |
nuclear factor I/B |
774 |
0.71 |
| chr1_934275_935357 | 0.87 |
HES4 |
hes family bHLH transcription factor 4 |
545 |
0.57 |
| chr22_41633466_41634617 | 0.87 |
CHADL |
chondroadherin-like |
1584 |
0.26 |
| chr2_236578912_236579455 | 0.87 |
AGAP1 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 1 |
935 |
0.7 |
| chr15_69590968_69591548 | 0.86 |
PAQR5 |
progestin and adipoQ receptor family member V |
28 |
0.66 |
| chr7_1705694_1706073 | 0.86 |
ELFN1 |
extracellular leucine-rich repeat and fibronectin type III domain containing 1 |
21872 |
0.17 |
| chr6_56707814_56708209 | 0.86 |
DST |
dystonin |
2 |
0.95 |
| chr5_72921316_72921781 | 0.86 |
ARHGEF28 |
Rho guanine nucleotide exchange factor (GEF) 28 |
435 |
0.84 |
| chr7_150812372_150812612 | 0.86 |
AGAP3 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 3 |
727 |
0.51 |
| chr1_112532383_112532987 | 0.86 |
KCND3 |
potassium voltage-gated channel, Shal-related subfamily, member 3 |
908 |
0.69 |
| chr16_88450130_88450579 | 0.86 |
ZNF469 |
zinc finger protein 469 |
43525 |
0.15 |
| chr2_206546644_206547002 | 0.85 |
NRP2 |
neuropilin 2 |
401 |
0.91 |
| chr2_128431846_128432683 | 0.85 |
LIMS2 |
LIM and senescent cell antigen-like domains 2 |
381 |
0.83 |
| chr1_231298564_231298974 | 0.85 |
TRIM67 |
tripartite motif containing 67 |
53 |
0.98 |
| chr5_159343463_159343902 | 0.85 |
ADRA1B |
adrenoceptor alpha 1B |
108 |
0.98 |
| chr11_70962576_70962972 | 0.85 |
SHANK2 |
SH3 and multiple ankyrin repeat domains 2 |
849 |
0.72 |
| chr12_80083529_80084306 | 0.85 |
PAWR |
PRKC, apoptosis, WT1, regulator |
57 |
0.98 |
| chr10_72432567_72432920 | 0.85 |
ADAMTS14 |
ADAM metallopeptidase with thrombospondin type 1 motif, 14 |
184 |
0.96 |
| chr22_46546744_46547896 | 0.85 |
PPARA |
peroxisome proliferator-activated receptor alpha |
467 |
0.75 |
| chr12_57522507_57523183 | 0.85 |
STAT6 |
signal transducer and activator of transcription 6, interleukin-4 induced |
38 |
0.64 |
| chr18_11149170_11149978 | 0.84 |
PIEZO2 |
piezo-type mechanosensitive ion channel component 2 |
987 |
0.72 |
| chr16_19125601_19126223 | 0.84 |
ITPRIPL2 |
inositol 1,4,5-trisphosphate receptor interacting protein-like 2 |
658 |
0.65 |
| chr1_62784626_62784933 | 0.84 |
KANK4 |
KN motif and ankyrin repeat domains 4 |
193 |
0.95 |
| chr10_75670529_75671818 | 0.84 |
PLAU |
plasminogen activator, urokinase |
257 |
0.89 |
| chr3_12329537_12330195 | 0.84 |
PPARG |
peroxisome proliferator-activated receptor gamma |
466 |
0.88 |
| chr2_243030793_243031816 | 0.84 |
AC093642.5 |
|
460 |
0.62 |
| chr16_74440490_74441404 | 0.84 |
CLEC18B |
C-type lectin domain family 18, member B |
14343 |
0.14 |
| chr18_10454512_10454665 | 0.84 |
APCDD1 |
adenomatosis polyposis coli down-regulated 1 |
37 |
0.98 |
| chr21_44495024_44495938 | 0.84 |
CBS |
cystathionine-beta-synthase |
460 |
0.8 |
| chr2_201171688_201172904 | 0.83 |
SPATS2L |
spermatogenesis associated, serine-rich 2-like |
701 |
0.76 |
| chr12_96183483_96184147 | 0.83 |
NTN4 |
netrin 4 |
89 |
0.96 |
| chr7_108095607_108096848 | 0.83 |
NRCAM |
neuronal cell adhesion molecule |
538 |
0.84 |
| chr14_38052851_38053204 | 0.83 |
FOXA1 |
forkhead box A1 |
11212 |
0.23 |
| chr15_83875578_83876391 | 0.83 |
HDGFRP3 |
Hepatoma-derived growth factor-related protein 3 |
786 |
0.64 |
| chr1_40236298_40236878 | 0.82 |
OXCT2 |
3-oxoacid CoA transferase 2 |
432 |
0.76 |
| chr8_23260918_23261191 | 0.82 |
LOXL2 |
lysyl oxidase-like 2 |
535 |
0.78 |
| chr4_4388085_4389202 | 0.82 |
NSG1 |
Homo sapiens neuron specific gene family member 1 (NSG1), transcript variant 3, mRNA. |
189 |
0.96 |
| chr3_142607235_142608274 | 0.82 |
PCOLCE2 |
procollagen C-endopeptidase enhancer 2 |
13 |
0.98 |
| chr19_460012_460518 | 0.82 |
SHC2 |
SHC (Src homology 2 domain containing) transforming protein 2 |
731 |
0.5 |
| chr9_104248314_104249564 | 0.82 |
TMEM246 |
transmembrane protein 246 |
460 |
0.79 |
| chr19_45843770_45844284 | 0.81 |
KLC3 |
kinesin light chain 3 |
5 |
0.95 |
| chr19_46270942_46271877 | 0.80 |
AC074212.5 |
|
236 |
0.49 |
| chr6_19838575_19839711 | 0.80 |
RP1-167F1.2 |
|
168 |
0.95 |
| chr1_119543832_119544021 | 0.80 |
TBX15 |
T-box 15 |
11747 |
0.25 |
| chr9_90588787_90589959 | 0.80 |
CDK20 |
cyclin-dependent kinase 20 |
29 |
0.98 |
| chr4_87515623_87516236 | 0.80 |
PTPN13 |
protein tyrosine phosphatase, non-receptor type 13 (APO-1/CD95 (Fas)-associated phosphatase) |
44 |
0.93 |
| chr4_24913981_24914389 | 0.80 |
CCDC149 |
coiled-coil domain containing 149 |
323 |
0.94 |
| chr19_1566819_1567637 | 0.80 |
MEX3D |
mex-3 RNA binding family member D |
301 |
0.76 |
| chr4_20256302_20256578 | 0.79 |
SLIT2 |
slit homolog 2 (Drosophila) |
103 |
0.98 |
| chr14_67999720_68000517 | 0.79 |
PLEKHH1 |
pleckstrin homology domain containing, family H (with MyTH4 domain) member 1 |
100 |
0.78 |
| chr4_5053247_5053862 | 0.79 |
STK32B |
serine/threonine kinase 32B |
385 |
0.9 |
| chr10_71562195_71562717 | 0.79 |
COL13A1 |
collagen, type XIII, alpha 1 |
276 |
0.92 |
| chr5_31193803_31195028 | 0.78 |
CDH6 |
cadherin 6, type 2, K-cadherin (fetal kidney) |
558 |
0.85 |
| chr13_27131903_27132772 | 0.78 |
WASF3 |
WAS protein family, member 3 |
450 |
0.89 |
| chr16_1664599_1665551 | 0.78 |
CRAMP1L |
Crm, cramped-like (Drosophila) |
434 |
0.69 |
| chr3_55522456_55523178 | 0.78 |
WNT5A-AS1 |
WNT5A antisense RNA 1 |
1090 |
0.43 |
| chr17_79373625_79374766 | 0.78 |
ENSG00000266392 |
. |
383 |
0.6 |
| chr5_131562588_131563831 | 0.78 |
P4HA2 |
prolyl 4-hydroxylase, alpha polypeptide II |
138 |
0.96 |
| chr21_45139141_45140358 | 0.78 |
PDXK |
pyridoxal (pyridoxine, vitamin B6) kinase |
756 |
0.66 |
| chr6_105627392_105627796 | 0.78 |
POPDC3 |
popeye domain containing 3 |
141 |
0.93 |
| chrX_50556610_50557434 | 0.78 |
SHROOM4 |
shroom family member 4 |
22 |
0.99 |
| chr2_45395914_45396553 | 0.78 |
SIX2 |
SIX homeobox 2 |
159664 |
0.04 |
| chr16_51185618_51185964 | 0.78 |
SALL1 |
spalt-like transcription factor 1 |
605 |
0.76 |
| chr15_83316430_83317127 | 0.77 |
CPEB1 |
cytoplasmic polyadenylation element binding protein 1 |
50 |
0.76 |
Network of associatons between targets according to the STRING database.

Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.1 | GO:0050923 | regulation of negative chemotaxis(GO:0050923) |
| 0.8 | 2.4 | GO:0060677 | ureteric bud elongation(GO:0060677) |
| 0.7 | 2.1 | GO:0055098 | response to low-density lipoprotein particle(GO:0055098) |
| 0.7 | 3.5 | GO:0060206 | estrous cycle phase(GO:0060206) |
| 0.6 | 1.2 | GO:0002072 | optic cup morphogenesis involved in camera-type eye development(GO:0002072) |
| 0.6 | 2.3 | GO:0046322 | negative regulation of fatty acid oxidation(GO:0046322) |
| 0.5 | 1.4 | GO:0070141 | response to UV-A(GO:0070141) |
| 0.4 | 2.2 | GO:0090162 | establishment of epithelial cell polarity(GO:0090162) |
| 0.4 | 1.2 | GO:0003099 | positive regulation of the force of heart contraction by chemical signal(GO:0003099) |
| 0.4 | 1.2 | GO:0010757 | negative regulation of plasminogen activation(GO:0010757) |
| 0.4 | 0.8 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.4 | 0.8 | GO:0060083 | smooth muscle contraction involved in micturition(GO:0060083) |
| 0.4 | 1.1 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.4 | 1.1 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.4 | 1.1 | GO:0071692 | sequestering of extracellular ligand from receptor(GO:0035581) sequestering of TGFbeta in extracellular matrix(GO:0035583) protein localization to extracellular region(GO:0071692) maintenance of protein location in extracellular region(GO:0071694) extracellular regulation of signal transduction(GO:1900115) extracellular negative regulation of signal transduction(GO:1900116) |
| 0.4 | 0.4 | GO:0060686 | negative regulation of prostatic bud formation(GO:0060686) |
| 0.4 | 0.4 | GO:0072179 | nephric duct formation(GO:0072179) |
| 0.3 | 0.3 | GO:0072047 | proximal/distal pattern formation involved in nephron development(GO:0072047) specification of nephron tubule identity(GO:0072081) |
| 0.3 | 1.7 | GO:0048875 | surfactant homeostasis(GO:0043129) chemical homeostasis within a tissue(GO:0048875) |
| 0.3 | 0.3 | GO:0090103 | cochlea morphogenesis(GO:0090103) |
| 0.3 | 1.3 | GO:0072070 | loop of Henle development(GO:0072070) |
| 0.3 | 1.7 | GO:0045662 | negative regulation of myoblast differentiation(GO:0045662) |
| 0.3 | 1.0 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.3 | 1.0 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.3 | 1.0 | GO:0010566 | regulation of ketone biosynthetic process(GO:0010566) regulation of aldosterone metabolic process(GO:0032344) regulation of aldosterone biosynthetic process(GO:0032347) |
| 0.3 | 0.6 | GO:0060839 | endothelial cell fate commitment(GO:0060839) |
| 0.3 | 2.2 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.3 | 0.9 | GO:0009648 | photoperiodism(GO:0009648) |
| 0.3 | 0.9 | GO:0071502 | cellular response to temperature stimulus(GO:0071502) |
| 0.3 | 0.9 | GO:0021631 | optic nerve morphogenesis(GO:0021631) |
| 0.3 | 0.3 | GO:0032351 | negative regulation of hormone metabolic process(GO:0032351) negative regulation of hormone biosynthetic process(GO:0032353) |
| 0.3 | 0.9 | GO:0009756 | carbohydrate mediated signaling(GO:0009756) |
| 0.3 | 0.3 | GO:0072048 | pattern specification involved in kidney development(GO:0061004) renal system pattern specification(GO:0072048) |
| 0.3 | 0.9 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) 4-hydroxyproline metabolic process(GO:0019471) peptidyl-proline hydroxylation(GO:0019511) |
| 0.3 | 0.8 | GO:0032236 | obsolete positive regulation of calcium ion transport via store-operated calcium channel activity(GO:0032236) |
| 0.3 | 1.1 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.3 | 1.1 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.3 | 1.1 | GO:0008218 | bioluminescence(GO:0008218) |
| 0.3 | 0.5 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.3 | 0.3 | GO:0021798 | forebrain dorsal/ventral pattern formation(GO:0021798) |
| 0.3 | 0.8 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) myosin filament assembly(GO:0031034) striated muscle myosin thick filament assembly(GO:0071688) |
| 0.3 | 1.3 | GO:0042219 | cellular modified amino acid catabolic process(GO:0042219) |
| 0.3 | 0.3 | GO:0061364 | Roundabout signaling pathway(GO:0035385) apoptotic process involved in luteolysis(GO:0061364) |
| 0.2 | 0.7 | GO:0048845 | venous blood vessel morphogenesis(GO:0048845) |
| 0.2 | 1.7 | GO:0007185 | transmembrane receptor protein tyrosine phosphatase signaling pathway(GO:0007185) |
| 0.2 | 1.2 | GO:0021873 | forebrain neuroblast division(GO:0021873) |
| 0.2 | 0.7 | GO:0071883 | adrenergic receptor signaling pathway(GO:0071875) activation of MAPK activity by adrenergic receptor signaling pathway(GO:0071883) |
| 0.2 | 0.5 | GO:0060129 | thyroid-stimulating hormone-secreting cell differentiation(GO:0060129) |
| 0.2 | 0.2 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 0.2 | 0.7 | GO:0021932 | hindbrain radial glia guided cell migration(GO:0021932) |
| 0.2 | 0.7 | GO:0022605 | oogenesis stage(GO:0022605) |
| 0.2 | 0.7 | GO:0032331 | negative regulation of chondrocyte differentiation(GO:0032331) |
| 0.2 | 0.9 | GO:0021830 | interneuron migration from the subpallium to the cortex(GO:0021830) cerebral cortex GABAergic interneuron migration(GO:0021853) cerebral cortex GABAergic interneuron development(GO:0021894) interneuron migration(GO:1904936) |
| 0.2 | 0.2 | GO:0045915 | positive regulation of catecholamine metabolic process(GO:0045915) positive regulation of dopamine metabolic process(GO:0045964) |
| 0.2 | 0.6 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.2 | 0.6 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.2 | 0.4 | GO:0002689 | negative regulation of leukocyte chemotaxis(GO:0002689) |
| 0.2 | 1.6 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.2 | 0.4 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.2 | 0.8 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.2 | 0.6 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.2 | 0.4 | GO:0045978 | negative regulation of nucleoside metabolic process(GO:0045978) negative regulation of ATP metabolic process(GO:1903579) |
| 0.2 | 0.2 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.2 | 0.6 | GO:0070874 | negative regulation of glycogen metabolic process(GO:0070874) |
| 0.2 | 0.4 | GO:0007500 | mesodermal cell fate determination(GO:0007500) |
| 0.2 | 0.6 | GO:0035588 | adenosine receptor signaling pathway(GO:0001973) G-protein coupled purinergic receptor signaling pathway(GO:0035588) |
| 0.2 | 0.6 | GO:0035610 | C-terminal protein deglutamylation(GO:0035609) protein side chain deglutamylation(GO:0035610) |
| 0.2 | 0.2 | GO:0010837 | regulation of keratinocyte proliferation(GO:0010837) |
| 0.2 | 1.2 | GO:0007351 | tripartite regional subdivision(GO:0007351) anterior/posterior axis specification, embryo(GO:0008595) |
| 0.2 | 1.4 | GO:0036296 | response to increased oxygen levels(GO:0036296) response to hyperoxia(GO:0055093) |
| 0.2 | 0.2 | GO:2000980 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.2 | 0.2 | GO:0009214 | cyclic nucleotide catabolic process(GO:0009214) |
| 0.2 | 0.5 | GO:0035234 | ectopic germ cell programmed cell death(GO:0035234) |
| 0.2 | 0.5 | GO:0040037 | negative regulation of fibroblast growth factor receptor signaling pathway(GO:0040037) |
| 0.2 | 0.5 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.2 | 0.5 | GO:0021563 | glossopharyngeal nerve development(GO:0021563) glossopharyngeal nerve morphogenesis(GO:0021615) |
| 0.2 | 0.2 | GO:0060059 | embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.2 | 0.2 | GO:0045992 | negative regulation of embryonic development(GO:0045992) |
| 0.2 | 1.2 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.2 | 0.5 | GO:0016103 | diterpenoid catabolic process(GO:0016103) terpenoid catabolic process(GO:0016115) retinoic acid catabolic process(GO:0034653) |
| 0.2 | 0.3 | GO:0051541 | elastin metabolic process(GO:0051541) |
| 0.2 | 0.5 | GO:0045112 | integrin biosynthetic process(GO:0045112) |
| 0.2 | 0.5 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.2 | 0.3 | GO:0003148 | outflow tract septum morphogenesis(GO:0003148) |
| 0.2 | 1.0 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.2 | 0.6 | GO:0014012 | peripheral nervous system axon regeneration(GO:0014012) |
| 0.2 | 0.3 | GO:0090273 | regulation of somatostatin secretion(GO:0090273) |
| 0.2 | 0.8 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.2 | 0.2 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.2 | 0.5 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.2 | 0.2 | GO:0010664 | striated muscle cell apoptotic process(GO:0010658) cardiac muscle cell apoptotic process(GO:0010659) regulation of striated muscle cell apoptotic process(GO:0010662) negative regulation of striated muscle cell apoptotic process(GO:0010664) regulation of cardiac muscle cell apoptotic process(GO:0010665) negative regulation of cardiac muscle cell apoptotic process(GO:0010667) |
| 0.2 | 0.5 | GO:0045079 | negative regulation of chemokine biosynthetic process(GO:0045079) |
| 0.2 | 0.8 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.2 | 0.8 | GO:0051005 | negative regulation of lipoprotein lipase activity(GO:0051005) |
| 0.2 | 0.3 | GO:0090598 | male genitalia morphogenesis(GO:0048808) male anatomical structure morphogenesis(GO:0090598) |
| 0.1 | 0.4 | GO:0034201 | response to oleic acid(GO:0034201) |
| 0.1 | 0.6 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 0.1 | 0.6 | GO:0030146 | obsolete diuresis(GO:0030146) |
| 0.1 | 0.1 | GO:0060841 | venous blood vessel development(GO:0060841) |
| 0.1 | 0.6 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 2.0 | GO:0032400 | melanosome localization(GO:0032400) |
| 0.1 | 0.6 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.1 | 0.1 | GO:0060748 | tertiary branching involved in mammary gland duct morphogenesis(GO:0060748) |
| 0.1 | 0.4 | GO:0032060 | bleb assembly(GO:0032060) |
| 0.1 | 0.7 | GO:0001946 | lymphangiogenesis(GO:0001946) lymph vessel morphogenesis(GO:0036303) |
| 0.1 | 1.0 | GO:1900006 | positive regulation of dendrite morphogenesis(GO:0050775) positive regulation of dendrite development(GO:1900006) |
| 0.1 | 0.3 | GO:0048865 | stem cell fate commitment(GO:0048865) stem cell fate determination(GO:0048867) |
| 0.1 | 0.7 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.1 | 0.4 | GO:0001705 | ectoderm formation(GO:0001705) |
| 0.1 | 0.4 | GO:0021781 | glial cell fate commitment(GO:0021781) |
| 0.1 | 0.4 | GO:0051305 | chromosome movement towards spindle pole(GO:0051305) |
| 0.1 | 0.5 | GO:0032288 | myelin assembly(GO:0032288) |
| 0.1 | 0.4 | GO:0008215 | spermine metabolic process(GO:0008215) |
| 0.1 | 0.8 | GO:0030033 | microvillus assembly(GO:0030033) microvillus organization(GO:0032528) |
| 0.1 | 0.9 | GO:1904893 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.1 | 0.1 | GO:0006910 | phagocytosis, recognition(GO:0006910) |
| 0.1 | 0.3 | GO:0060214 | endocardium development(GO:0003157) endocardium morphogenesis(GO:0003160) endocardium formation(GO:0060214) |
| 0.1 | 0.4 | GO:0046886 | positive regulation of hormone metabolic process(GO:0032352) positive regulation of hormone biosynthetic process(GO:0046886) |
| 0.1 | 0.9 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.1 | 0.1 | GO:0031946 | regulation of glucocorticoid biosynthetic process(GO:0031946) |
| 0.1 | 0.2 | GO:0060438 | trachea development(GO:0060438) |
| 0.1 | 0.5 | GO:0030917 | midbrain-hindbrain boundary development(GO:0030917) |
| 0.1 | 0.2 | GO:0060065 | uterus development(GO:0060065) |
| 0.1 | 0.7 | GO:0006837 | serotonin transport(GO:0006837) |
| 0.1 | 0.5 | GO:0061029 | eyelid development in camera-type eye(GO:0061029) |
| 0.1 | 1.5 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.1 | 0.1 | GO:0003321 | positive regulation of blood pressure by epinephrine-norepinephrine(GO:0003321) |
| 0.1 | 0.5 | GO:0032455 | nerve growth factor processing(GO:0032455) |
| 0.1 | 0.4 | GO:0048251 | elastic fiber assembly(GO:0048251) |
| 0.1 | 0.5 | GO:0018101 | protein citrullination(GO:0018101) |
| 0.1 | 0.2 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) pyramidal neuron development(GO:0021860) |
| 0.1 | 0.2 | GO:0032413 | negative regulation of ion transmembrane transporter activity(GO:0032413) |
| 0.1 | 0.6 | GO:0002068 | glandular epithelial cell development(GO:0002068) |
| 0.1 | 0.1 | GO:0021800 | cerebral cortex tangential migration(GO:0021800) |
| 0.1 | 0.2 | GO:0010700 | negative regulation of norepinephrine secretion(GO:0010700) |
| 0.1 | 0.5 | GO:0051927 | obsolete negative regulation of calcium ion transport via voltage-gated calcium channel activity(GO:0051927) |
| 0.1 | 0.5 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.1 | 2.1 | GO:0006182 | cGMP biosynthetic process(GO:0006182) |
| 0.1 | 0.3 | GO:0030538 | embryonic genitalia morphogenesis(GO:0030538) |
| 0.1 | 1.6 | GO:0045104 | intermediate filament cytoskeleton organization(GO:0045104) |
| 0.1 | 0.5 | GO:0007619 | courtship behavior(GO:0007619) |
| 0.1 | 0.2 | GO:0061430 | bone trabecula formation(GO:0060346) bone trabecula morphogenesis(GO:0061430) |
| 0.1 | 1.1 | GO:0009650 | UV protection(GO:0009650) |
| 0.1 | 0.4 | GO:0001757 | somite specification(GO:0001757) |
| 0.1 | 0.4 | GO:0022617 | extracellular matrix disassembly(GO:0022617) |
| 0.1 | 0.3 | GO:0045007 | depurination(GO:0045007) |
| 0.1 | 0.4 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.1 | 0.4 | GO:0060992 | response to fungicide(GO:0060992) |
| 0.1 | 0.1 | GO:0030252 | growth hormone secretion(GO:0030252) |
| 0.1 | 0.2 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.1 | 1.5 | GO:0051350 | negative regulation of adenylate cyclase activity(GO:0007194) negative regulation of cyclase activity(GO:0031280) negative regulation of lyase activity(GO:0051350) |
| 0.1 | 0.3 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.1 | 1.1 | GO:0006688 | glycosphingolipid biosynthetic process(GO:0006688) |
| 0.1 | 0.1 | GO:0042637 | catagen(GO:0042637) |
| 0.1 | 0.1 | GO:0060527 | prostate glandular acinus morphogenesis(GO:0060526) prostate epithelial cord arborization involved in prostate glandular acinus morphogenesis(GO:0060527) |
| 0.1 | 0.5 | GO:0046184 | aldehyde biosynthetic process(GO:0046184) |
| 0.1 | 0.4 | GO:0035283 | central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.1 | 0.3 | GO:0043137 | DNA replication, Okazaki fragment processing(GO:0033567) DNA replication, removal of RNA primer(GO:0043137) |
| 0.1 | 0.5 | GO:0002829 | negative regulation of type 2 immune response(GO:0002829) |
| 0.1 | 1.5 | GO:0034199 | activation of protein kinase A activity(GO:0034199) |
| 0.1 | 0.6 | GO:0015904 | tetracycline transport(GO:0015904) |
| 0.1 | 2.6 | GO:0071804 | cellular potassium ion transport(GO:0071804) potassium ion transmembrane transport(GO:0071805) |
| 0.1 | 0.4 | GO:0090026 | regulation of monocyte chemotaxis(GO:0090025) positive regulation of monocyte chemotaxis(GO:0090026) |
| 0.1 | 0.4 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.1 | 0.1 | GO:0042268 | regulation of cytolysis(GO:0042268) |
| 0.1 | 0.3 | GO:0051895 | negative regulation of focal adhesion assembly(GO:0051895) negative regulation of cell junction assembly(GO:1901889) negative regulation of adherens junction organization(GO:1903392) |
| 0.1 | 0.7 | GO:0033599 | regulation of mammary gland epithelial cell proliferation(GO:0033599) |
| 0.1 | 0.3 | GO:0007171 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) |
| 0.1 | 0.5 | GO:0042249 | establishment of planar polarity of embryonic epithelium(GO:0042249) |
| 0.1 | 0.1 | GO:0002066 | columnar/cuboidal epithelial cell development(GO:0002066) |
| 0.1 | 0.3 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.1 | 0.3 | GO:0006562 | proline catabolic process(GO:0006562) |
| 0.1 | 0.8 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.1 | 0.3 | GO:0045077 | negative regulation of interferon-gamma biosynthetic process(GO:0045077) |
| 0.1 | 0.2 | GO:0031943 | regulation of glucocorticoid metabolic process(GO:0031943) |
| 0.1 | 0.5 | GO:0032026 | response to magnesium ion(GO:0032026) |
| 0.1 | 0.4 | GO:0048935 | peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 0.1 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.1 | 0.4 | GO:0003214 | cardiac left ventricle morphogenesis(GO:0003214) |
| 0.1 | 0.1 | GO:0060592 | mammary gland formation(GO:0060592) |
| 0.1 | 0.3 | GO:0006384 | transcription initiation from RNA polymerase III promoter(GO:0006384) |
| 0.1 | 0.2 | GO:0030800 | negative regulation of cyclic nucleotide metabolic process(GO:0030800) negative regulation of cyclic nucleotide biosynthetic process(GO:0030803) negative regulation of nucleotide biosynthetic process(GO:0030809) negative regulation of cAMP metabolic process(GO:0030815) negative regulation of cAMP biosynthetic process(GO:0030818) negative regulation of purine nucleotide biosynthetic process(GO:1900372) negative regulation of purine nucleotide metabolic process(GO:1900543) |
| 0.1 | 0.3 | GO:0061072 | iris morphogenesis(GO:0061072) |
| 0.1 | 0.1 | GO:0045989 | positive regulation of striated muscle contraction(GO:0045989) |
| 0.1 | 0.4 | GO:0031641 | regulation of myelination(GO:0031641) |
| 0.1 | 0.3 | GO:0016246 | RNA interference(GO:0016246) |
| 0.1 | 1.8 | GO:0006940 | regulation of smooth muscle contraction(GO:0006940) |
| 0.1 | 0.2 | GO:0034638 | phosphatidylcholine catabolic process(GO:0034638) |
| 0.1 | 0.1 | GO:0048485 | sympathetic nervous system development(GO:0048485) |
| 0.1 | 1.6 | GO:0030574 | collagen catabolic process(GO:0030574) |
| 0.1 | 0.2 | GO:0060456 | positive regulation of digestive system process(GO:0060456) |
| 0.1 | 0.4 | GO:0090083 | regulation of inclusion body assembly(GO:0090083) |
| 0.1 | 0.4 | GO:0007413 | axonal fasciculation(GO:0007413) |
| 0.1 | 0.5 | GO:0045070 | positive regulation of viral genome replication(GO:0045070) |
| 0.1 | 0.2 | GO:0048702 | embryonic neurocranium morphogenesis(GO:0048702) |
| 0.1 | 0.4 | GO:0008347 | glial cell migration(GO:0008347) |
| 0.1 | 0.3 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.1 | 0.5 | GO:0015802 | basic amino acid transport(GO:0015802) |
| 0.1 | 0.3 | GO:0071800 | podosome assembly(GO:0071800) |
| 0.1 | 0.2 | GO:0001945 | lymph vessel development(GO:0001945) |
| 0.1 | 0.5 | GO:0006477 | protein sulfation(GO:0006477) |
| 0.1 | 0.9 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.1 | 0.2 | GO:0021889 | olfactory bulb interneuron differentiation(GO:0021889) |
| 0.1 | 0.4 | GO:0006975 | DNA damage induced protein phosphorylation(GO:0006975) |
| 0.1 | 0.4 | GO:0015824 | proline transport(GO:0015824) |
| 0.1 | 9.4 | GO:0006813 | potassium ion transport(GO:0006813) |
| 0.1 | 0.7 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.1 | 0.1 | GO:0016999 | antibiotic metabolic process(GO:0016999) antibiotic biosynthetic process(GO:0017000) |
| 0.1 | 0.9 | GO:0007528 | neuromuscular junction development(GO:0007528) |
| 0.1 | 0.3 | GO:1903318 | negative regulation of protein processing(GO:0010955) negative regulation of protein maturation(GO:1903318) |
| 0.1 | 0.7 | GO:0046697 | decidualization(GO:0046697) |
| 0.1 | 0.3 | GO:0060231 | mesenchymal to epithelial transition(GO:0060231) |
| 0.1 | 0.1 | GO:0050703 | interleukin-1 alpha secretion(GO:0050703) |
| 0.1 | 0.1 | GO:0045683 | negative regulation of epidermal cell differentiation(GO:0045605) negative regulation of keratinocyte differentiation(GO:0045617) negative regulation of epidermis development(GO:0045683) |
| 0.1 | 0.2 | GO:0051151 | negative regulation of smooth muscle cell differentiation(GO:0051151) |
| 0.1 | 0.2 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 2.9 | GO:0031424 | keratinization(GO:0031424) |
| 0.1 | 0.1 | GO:0042640 | anagen(GO:0042640) |
| 0.1 | 0.1 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 0.1 | 0.1 | GO:0061726 | mitophagy(GO:0000422) mitochondrion disassembly(GO:0061726) |
| 0.1 | 0.1 | GO:0060267 | positive regulation of respiratory burst(GO:0060267) |
| 0.1 | 1.1 | GO:0001539 | cilium or flagellum-dependent cell motility(GO:0001539) |
| 0.1 | 0.1 | GO:0018126 | protein hydroxylation(GO:0018126) |
| 0.1 | 0.5 | GO:0008038 | neuron recognition(GO:0008038) |
| 0.1 | 0.2 | GO:0046549 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
| 0.1 | 0.9 | GO:0050766 | positive regulation of phagocytosis(GO:0050766) |
| 0.1 | 0.1 | GO:0048715 | negative regulation of oligodendrocyte differentiation(GO:0048715) |
| 0.1 | 0.3 | GO:0032429 | regulation of phospholipase A2 activity(GO:0032429) |
| 0.1 | 0.4 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.1 | 0.1 | GO:0048319 | axial mesoderm morphogenesis(GO:0048319) |
| 0.1 | 0.1 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.1 | 0.1 | GO:0021514 | ventral spinal cord interneuron differentiation(GO:0021514) regulation of transcription from RNA polymerase II promoter involved in ventral spinal cord interneuron specification(GO:0021913) ventral spinal cord interneuron fate commitment(GO:0060579) cell fate commitment involved in pattern specification(GO:0060581) |
| 0.1 | 1.8 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.1 | 0.4 | GO:0007617 | mating behavior(GO:0007617) multi-organism reproductive behavior(GO:0044705) |
| 0.1 | 0.3 | GO:0007549 | dosage compensation(GO:0007549) |
| 0.1 | 0.7 | GO:0035115 | embryonic forelimb morphogenesis(GO:0035115) |
| 0.1 | 0.4 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.1 | 0.2 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.1 | 0.2 | GO:0050995 | negative regulation of lipid catabolic process(GO:0050995) |
| 0.1 | 0.1 | GO:0090175 | Wnt signaling pathway, planar cell polarity pathway(GO:0060071) regulation of establishment of planar polarity(GO:0090175) |
| 0.1 | 0.5 | GO:0009954 | proximal/distal pattern formation(GO:0009954) |
| 0.1 | 0.2 | GO:0015868 | purine ribonucleotide transport(GO:0015868) |
| 0.1 | 0.4 | GO:0016079 | synaptic vesicle exocytosis(GO:0016079) |
| 0.1 | 0.2 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 0.6 | GO:0014003 | oligodendrocyte development(GO:0014003) |
| 0.1 | 0.2 | GO:0007616 | long-term memory(GO:0007616) |
| 0.1 | 0.1 | GO:1901374 | acetylcholine transport(GO:0015870) acetate ester transport(GO:1901374) |
| 0.1 | 0.2 | GO:0002220 | innate immune response activating cell surface receptor signaling pathway(GO:0002220) |
| 0.1 | 0.4 | GO:0030325 | adrenal gland development(GO:0030325) |
| 0.1 | 0.3 | GO:0051930 | regulation of sensory perception of pain(GO:0051930) regulation of sensory perception(GO:0051931) |
| 0.1 | 1.4 | GO:2000181 | negative regulation of angiogenesis(GO:0016525) negative regulation of blood vessel morphogenesis(GO:2000181) |
| 0.1 | 0.6 | GO:0021987 | cerebral cortex development(GO:0021987) |
| 0.1 | 0.3 | GO:0060314 | regulation of ryanodine-sensitive calcium-release channel activity(GO:0060314) |
| 0.1 | 7.3 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 0.2 | GO:0043302 | positive regulation of mast cell activation involved in immune response(GO:0033008) positive regulation of leukocyte degranulation(GO:0043302) positive regulation of mast cell degranulation(GO:0043306) |
| 0.1 | 0.1 | GO:0060013 | righting reflex(GO:0060013) |
| 0.1 | 0.3 | GO:0015732 | prostaglandin transport(GO:0015732) |
| 0.1 | 0.6 | GO:0007530 | sex determination(GO:0007530) |
| 0.1 | 0.3 | GO:0002070 | epithelial cell maturation(GO:0002070) |
| 0.1 | 0.1 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 1.0 | GO:0033344 | cholesterol efflux(GO:0033344) |
| 0.1 | 0.2 | GO:0035176 | social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.1 | 0.6 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.1 | 0.3 | GO:0070373 | negative regulation of ERK1 and ERK2 cascade(GO:0070373) |
| 0.1 | 0.2 | GO:0006188 | IMP biosynthetic process(GO:0006188) IMP metabolic process(GO:0046040) |
| 0.1 | 0.1 | GO:0001893 | maternal placenta development(GO:0001893) |
| 0.1 | 0.1 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.1 | 0.2 | GO:0021854 | hypothalamus development(GO:0021854) |
| 0.1 | 0.1 | GO:1903960 | negative regulation of plasma membrane long-chain fatty acid transport(GO:0010748) negative regulation of anion transmembrane transport(GO:1903960) negative regulation of fatty acid transport(GO:2000192) |
| 0.1 | 0.5 | GO:0007016 | cytoskeletal anchoring at plasma membrane(GO:0007016) |
| 0.1 | 0.2 | GO:2000189 | regulation of cholesterol homeostasis(GO:2000188) positive regulation of cholesterol homeostasis(GO:2000189) |
| 0.1 | 0.2 | GO:0090205 | positive regulation of cholesterol metabolic process(GO:0090205) |
| 0.1 | 0.2 | GO:0008588 | release of cytoplasmic sequestered NF-kappaB(GO:0008588) |
| 0.1 | 0.5 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.1 | 0.2 | GO:0016264 | gap junction assembly(GO:0016264) |
| 0.1 | 0.4 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.1 | 0.1 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 0.1 | 0.1 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.1 | 0.2 | GO:0006104 | succinyl-CoA metabolic process(GO:0006104) |
| 0.1 | 0.1 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.1 | 0.2 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.1 | 0.2 | GO:0000052 | citrulline metabolic process(GO:0000052) |
| 0.1 | 0.1 | GO:0044803 | multi-organism membrane organization(GO:0044803) |
| 0.1 | 0.2 | GO:0018243 | protein O-linked glycosylation via serine(GO:0018242) protein O-linked glycosylation via threonine(GO:0018243) |
| 0.1 | 0.1 | GO:0051497 | negative regulation of stress fiber assembly(GO:0051497) |
| 0.1 | 0.1 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.1 | 0.5 | GO:0018214 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.1 | 0.7 | GO:0048010 | vascular endothelial growth factor receptor signaling pathway(GO:0048010) |
| 0.1 | 0.3 | GO:0060351 | cartilage development involved in endochondral bone morphogenesis(GO:0060351) |
| 0.1 | 0.1 | GO:0031652 | regulation of heat generation(GO:0031650) positive regulation of heat generation(GO:0031652) |
| 0.1 | 0.2 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.1 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.2 | GO:0060040 | retinal bipolar neuron differentiation(GO:0060040) |
| 0.0 | 0.1 | GO:0090218 | positive regulation of lipid kinase activity(GO:0090218) |
| 0.0 | 0.3 | GO:0031274 | pseudopodium assembly(GO:0031269) regulation of pseudopodium assembly(GO:0031272) positive regulation of pseudopodium assembly(GO:0031274) |
| 0.0 | 0.0 | GO:0046502 | uroporphyrinogen III metabolic process(GO:0046502) |
| 0.0 | 1.4 | GO:0001764 | neuron migration(GO:0001764) |
| 0.0 | 0.4 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 0.3 | GO:0006657 | CDP-choline pathway(GO:0006657) |
| 0.0 | 0.1 | GO:0046618 | drug export(GO:0046618) |
| 0.0 | 0.1 | GO:0046882 | negative regulation of follicle-stimulating hormone secretion(GO:0046882) |
| 0.0 | 0.3 | GO:0045880 | positive regulation of smoothened signaling pathway(GO:0045880) |
| 0.0 | 0.2 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.0 | 0.1 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.0 | 0.0 | GO:0005984 | disaccharide metabolic process(GO:0005984) |
| 0.0 | 0.2 | GO:0070528 | protein kinase C signaling(GO:0070528) |
| 0.0 | 0.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.0 | 0.3 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.0 | 0.2 | GO:0009188 | ADP biosynthetic process(GO:0006172) purine nucleoside diphosphate biosynthetic process(GO:0009136) purine ribonucleoside diphosphate biosynthetic process(GO:0009180) ribonucleoside diphosphate biosynthetic process(GO:0009188) |
| 0.0 | 0.2 | GO:0007405 | neuroblast proliferation(GO:0007405) |
| 0.0 | 0.2 | GO:0006561 | proline biosynthetic process(GO:0006561) |
| 0.0 | 0.8 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.0 | 0.2 | GO:0001573 | ganglioside metabolic process(GO:0001573) |
| 0.0 | 0.0 | GO:0051590 | positive regulation of neurotransmitter transport(GO:0051590) |
| 0.0 | 0.4 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.0 | 0.0 | GO:0002513 | tolerance induction to self antigen(GO:0002513) |
| 0.0 | 0.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 1.4 | GO:0007588 | excretion(GO:0007588) |
| 0.0 | 0.1 | GO:0046607 | positive regulation of centrosome cycle(GO:0046607) |
| 0.0 | 0.0 | GO:0021937 | cerebellar Purkinje cell-granule cell precursor cell signaling involved in regulation of granule cell precursor cell proliferation(GO:0021937) |
| 0.0 | 0.0 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.0 | 2.9 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.1 | GO:0050807 | regulation of synapse organization(GO:0050807) |
| 0.0 | 0.1 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.0 | 0.1 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) positive regulation of protein depolymerization(GO:1901881) |
| 0.0 | 0.3 | GO:0015884 | folic acid transport(GO:0015884) |
| 0.0 | 0.2 | GO:0007622 | rhythmic behavior(GO:0007622) circadian behavior(GO:0048512) |
| 0.0 | 0.1 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.0 | 0.2 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.0 | 0.0 | GO:0090400 | stress-induced premature senescence(GO:0090400) |
| 0.0 | 0.4 | GO:0003143 | heart looping(GO:0001947) embryonic heart tube morphogenesis(GO:0003143) determination of heart left/right asymmetry(GO:0061371) |
| 0.0 | 0.1 | GO:0007210 | serotonin receptor signaling pathway(GO:0007210) |
| 0.0 | 0.1 | GO:0097479 | synaptic vesicle transport(GO:0048489) synaptic vesicle localization(GO:0097479) establishment of synaptic vesicle localization(GO:0097480) |
| 0.0 | 0.2 | GO:0001960 | negative regulation of cytokine-mediated signaling pathway(GO:0001960) |
| 0.0 | 0.2 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.0 | 0.2 | GO:0006991 | response to sterol depletion(GO:0006991) |
| 0.0 | 0.0 | GO:0010092 | specification of organ identity(GO:0010092) |
| 0.0 | 0.1 | GO:0044254 | multicellular organismal protein catabolic process(GO:0044254) protein digestion(GO:0044256) multicellular organismal macromolecule catabolic process(GO:0044266) multicellular organismal protein metabolic process(GO:0044268) |
| 0.0 | 0.2 | GO:0014819 | regulation of skeletal muscle contraction(GO:0014819) |
| 0.0 | 0.1 | GO:0030579 | ubiquitin-dependent SMAD protein catabolic process(GO:0030579) |
| 0.0 | 0.3 | GO:0006939 | smooth muscle contraction(GO:0006939) |
| 0.0 | 0.1 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.0 | 0.1 | GO:0070306 | lens fiber cell differentiation(GO:0070306) |
| 0.0 | 0.0 | GO:0014850 | response to muscle activity(GO:0014850) |
| 0.0 | 0.0 | GO:0060442 | branching involved in prostate gland morphogenesis(GO:0060442) |
| 0.0 | 0.1 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.0 | 0.1 | GO:0015864 | pyrimidine nucleoside transport(GO:0015864) |
| 0.0 | 0.1 | GO:0048630 | skeletal muscle tissue growth(GO:0048630) |
| 0.0 | 0.1 | GO:0060124 | positive regulation of growth hormone secretion(GO:0060124) |
| 0.0 | 0.3 | GO:0045995 | regulation of embryonic development(GO:0045995) |
| 0.0 | 0.1 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 0.1 | GO:0070208 | protein heterotrimerization(GO:0070208) |
| 0.0 | 0.0 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.0 | 0.1 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.0 | 0.2 | GO:0010633 | negative regulation of epithelial cell migration(GO:0010633) |
| 0.0 | 0.1 | GO:0043691 | reverse cholesterol transport(GO:0043691) |
| 0.0 | 0.1 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) maturation of LSU-rRNA(GO:0000470) |
| 0.0 | 0.1 | GO:0021517 | ventral spinal cord development(GO:0021517) |
| 0.0 | 9.0 | GO:0007409 | axonogenesis(GO:0007409) |
| 0.0 | 0.3 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.0 | GO:0070244 | negative regulation of thymocyte apoptotic process(GO:0070244) |
| 0.0 | 0.2 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.0 | GO:0033629 | negative regulation of cell adhesion mediated by integrin(GO:0033629) |
| 0.0 | 0.3 | GO:0032740 | positive regulation of interleukin-17 production(GO:0032740) |
| 0.0 | 1.0 | GO:0030178 | negative regulation of Wnt signaling pathway(GO:0030178) |
| 0.0 | 0.3 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 0.1 | GO:0002281 | macrophage activation involved in immune response(GO:0002281) |
| 0.0 | 0.2 | GO:0010744 | positive regulation of macrophage derived foam cell differentiation(GO:0010744) |
| 0.0 | 2.4 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.0 | 0.2 | GO:0001706 | endoderm formation(GO:0001706) |
| 0.0 | 0.0 | GO:0015793 | glycerol transport(GO:0015793) |
| 0.0 | 0.2 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.0 | 0.1 | GO:0009106 | lipoate metabolic process(GO:0009106) |
| 0.0 | 0.1 | GO:0010216 | maintenance of DNA methylation(GO:0010216) |
| 0.0 | 0.2 | GO:0098743 | cartilage condensation(GO:0001502) cell aggregation(GO:0098743) |
| 0.0 | 0.6 | GO:0080171 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 0.2 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.0 | 0.2 | GO:0002675 | positive regulation of acute inflammatory response(GO:0002675) |
| 0.0 | 0.2 | GO:0031345 | negative regulation of cell projection organization(GO:0031345) |
| 0.0 | 0.0 | GO:0010966 | regulation of phosphate transport(GO:0010966) |
| 0.0 | 0.2 | GO:0010862 | positive regulation of pathway-restricted SMAD protein phosphorylation(GO:0010862) |
| 0.0 | 0.3 | GO:0017158 | regulation of calcium ion-dependent exocytosis(GO:0017158) |
| 0.0 | 0.0 | GO:0021591 | ventricular system development(GO:0021591) |
| 0.0 | 0.1 | GO:0019919 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) |
| 0.0 | 0.1 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.0 | 0.0 | GO:0009296 | obsolete flagellum assembly(GO:0009296) |
| 0.0 | 0.1 | GO:0016266 | O-glycan processing(GO:0016266) |
| 0.0 | 0.2 | GO:0046500 | S-adenosylmethionine metabolic process(GO:0046500) |
| 0.0 | 0.2 | GO:0046386 | deoxyribose phosphate catabolic process(GO:0046386) |
| 0.0 | 0.2 | GO:0031114 | negative regulation of microtubule depolymerization(GO:0007026) regulation of microtubule depolymerization(GO:0031114) |
| 0.0 | 0.2 | GO:0045806 | negative regulation of endocytosis(GO:0045806) |
| 0.0 | 0.1 | GO:0002011 | morphogenesis of an epithelial sheet(GO:0002011) |
| 0.0 | 0.2 | GO:0045749 | obsolete negative regulation of S phase of mitotic cell cycle(GO:0045749) |
| 0.0 | 0.6 | GO:0016338 | calcium-independent cell-cell adhesion via plasma membrane cell-adhesion molecules(GO:0016338) |
| 0.0 | 0.1 | GO:0014721 | voluntary skeletal muscle contraction(GO:0003010) twitch skeletal muscle contraction(GO:0014721) |
| 0.0 | 0.1 | GO:0002369 | T cell cytokine production(GO:0002369) regulation of T cell cytokine production(GO:0002724) |
| 0.0 | 0.1 | GO:0046501 | protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.0 | 0.1 | GO:0000966 | RNA 5'-end processing(GO:0000966) |
| 0.0 | 0.2 | GO:0033189 | response to vitamin A(GO:0033189) |
| 0.0 | 0.0 | GO:0003309 | type B pancreatic cell differentiation(GO:0003309) |
| 0.0 | 0.1 | GO:0002283 | neutrophil activation involved in immune response(GO:0002283) neutrophil degranulation(GO:0043312) |
| 0.0 | 0.1 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.1 | GO:0046606 | negative regulation of centrosome duplication(GO:0010826) negative regulation of centrosome cycle(GO:0046606) |
| 0.0 | 0.3 | GO:0015721 | bile acid and bile salt transport(GO:0015721) |
| 0.0 | 0.3 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.0 | 0.0 | GO:0031268 | pseudopodium organization(GO:0031268) |
| 0.0 | 0.0 | GO:0032410 | negative regulation of transporter activity(GO:0032410) |
| 0.0 | 0.0 | GO:0014052 | regulation of gamma-aminobutyric acid secretion(GO:0014052) |
| 0.0 | 0.2 | GO:0051925 | obsolete regulation of calcium ion transport via voltage-gated calcium channel activity(GO:0051925) |
| 0.0 | 0.0 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.0 | 0.0 | GO:0051901 | positive regulation of mitochondrial depolarization(GO:0051901) positive regulation of membrane depolarization(GO:1904181) |
| 0.0 | 0.1 | GO:0040020 | regulation of meiotic nuclear division(GO:0040020) |
| 0.0 | 0.2 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.0 | 0.1 | GO:0070193 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.0 | 0.0 | GO:0046007 | negative regulation of activated T cell proliferation(GO:0046007) |
| 0.0 | 1.7 | GO:0006814 | sodium ion transport(GO:0006814) |
| 0.0 | 0.1 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.0 | GO:0031125 | rRNA 3'-end processing(GO:0031125) |
| 0.0 | 0.1 | GO:0043324 | eye pigment biosynthetic process(GO:0006726) eye pigment metabolic process(GO:0042441) pigment metabolic process involved in developmental pigmentation(GO:0043324) pigment metabolic process involved in pigmentation(GO:0043474) |
| 0.0 | 0.0 | GO:0002331 | pre-B cell allelic exclusion(GO:0002331) |
| 0.0 | 0.1 | GO:0043954 | cellular component maintenance(GO:0043954) |
| 0.0 | 0.0 | GO:0050806 | positive regulation of synaptic transmission(GO:0050806) |
| 0.0 | 0.1 | GO:0007494 | midgut development(GO:0007494) |
| 0.0 | 0.0 | GO:0032277 | negative regulation of gonadotropin secretion(GO:0032277) |
| 0.0 | 0.1 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.0 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) regulation of neurotransmitter transport(GO:0051588) |
| 0.0 | 0.0 | GO:0007021 | tubulin complex assembly(GO:0007021) |
| 0.0 | 0.2 | GO:0042135 | neurotransmitter catabolic process(GO:0042135) |
| 0.0 | 0.2 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.1 | GO:0015871 | choline transport(GO:0015871) |
| 0.0 | 0.1 | GO:0000019 | regulation of mitotic recombination(GO:0000019) |
| 0.0 | 0.2 | GO:0018208 | peptidyl-proline modification(GO:0018208) |
| 0.0 | 0.0 | GO:0009111 | vitamin catabolic process(GO:0009111) fat-soluble vitamin catabolic process(GO:0042363) |
| 0.0 | 0.1 | GO:0046855 | phosphorylated carbohydrate dephosphorylation(GO:0046838) inositol phosphate dephosphorylation(GO:0046855) |
| 0.0 | 0.0 | GO:0018119 | protein nitrosylation(GO:0017014) peptidyl-cysteine S-nitrosylation(GO:0018119) |
| 0.0 | 0.1 | GO:0008634 | obsolete negative regulation of survival gene product expression(GO:0008634) |
| 0.0 | 0.2 | GO:0070371 | ERK1 and ERK2 cascade(GO:0070371) |
| 0.0 | 0.0 | GO:0006855 | drug transmembrane transport(GO:0006855) |
| 0.0 | 0.1 | GO:0016255 | attachment of GPI anchor to protein(GO:0016255) |
| 0.0 | 0.0 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.0 | 0.1 | GO:0031848 | protection from non-homologous end joining at telomere(GO:0031848) telomere maintenance in response to DNA damage(GO:0043247) |
| 0.0 | 0.0 | GO:0006527 | arginine catabolic process(GO:0006527) |
| 0.0 | 0.0 | GO:0097154 | GABAergic neuron differentiation(GO:0097154) |
| 0.0 | 0.1 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.0 | 0.0 | GO:0042428 | serotonin metabolic process(GO:0042428) primary amino compound metabolic process(GO:1901160) |
| 0.0 | 0.1 | GO:0050885 | neuromuscular process controlling balance(GO:0050885) |
| 0.0 | 0.4 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 0.0 | GO:0061101 | neuroendocrine cell differentiation(GO:0061101) |
| 0.0 | 0.1 | GO:0009404 | toxin metabolic process(GO:0009404) |
| 0.0 | 0.1 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.0 | 0.0 | GO:0071635 | negative regulation of transforming growth factor beta production(GO:0071635) |
| 0.0 | 0.0 | GO:0061088 | regulation of sequestering of zinc ion(GO:0061088) |
| 0.0 | 0.2 | GO:0042274 | ribosomal small subunit biogenesis(GO:0042274) |
| 0.0 | 0.3 | GO:0042472 | inner ear morphogenesis(GO:0042472) |
| 0.0 | 0.0 | GO:0021675 | nerve development(GO:0021675) |
| 0.0 | 0.1 | GO:0007007 | inner mitochondrial membrane organization(GO:0007007) |
| 0.0 | 0.1 | GO:0009304 | tRNA transcription(GO:0009304) 5S class rRNA transcription from RNA polymerase III type 1 promoter(GO:0042791) tRNA transcription from RNA polymerase III promoter(GO:0042797) |
| 0.0 | 0.0 | GO:0032634 | interleukin-5 production(GO:0032634) regulation of interleukin-5 production(GO:0032674) |
| 0.0 | 0.0 | GO:0007181 | transforming growth factor beta receptor complex assembly(GO:0007181) |
| 0.0 | 0.2 | GO:0046329 | negative regulation of JNK cascade(GO:0046329) |
| 0.0 | 0.1 | GO:0070934 | CRD-mediated mRNA stabilization(GO:0070934) |
| 0.0 | 0.2 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 0.0 | GO:0015893 | drug transport(GO:0015893) |
| 0.0 | 0.2 | GO:1901222 | activation of NF-kappaB-inducing kinase activity(GO:0007250) NIK/NF-kappaB signaling(GO:0038061) regulation of NIK/NF-kappaB signaling(GO:1901222) positive regulation of NIK/NF-kappaB signaling(GO:1901224) |
| 0.0 | 0.1 | GO:0007215 | glutamate receptor signaling pathway(GO:0007215) |
| 0.0 | 0.0 | GO:0060766 | negative regulation of androgen receptor signaling pathway(GO:0060766) |
| 0.0 | 0.0 | GO:0032328 | alanine transport(GO:0032328) |
| 0.0 | 0.1 | GO:0046415 | urate metabolic process(GO:0046415) |
| 0.0 | 0.1 | GO:0030865 | cortical cytoskeleton organization(GO:0030865) cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.0 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.0 | 0.0 | GO:0051955 | regulation of amino acid transport(GO:0051955) |
| 0.0 | 0.0 | GO:0006531 | aspartate metabolic process(GO:0006531) aspartate catabolic process(GO:0006533) |
| 0.0 | 0.0 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.0 | 0.0 | GO:0010455 | positive regulation of cell fate commitment(GO:0010455) |
| 0.0 | 0.4 | GO:0007605 | sensory perception of sound(GO:0007605) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 2.6 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.3 | 2.0 | GO:0010369 | chromocenter(GO:0010369) |
| 0.3 | 0.9 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.3 | 1.5 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.3 | 0.9 | GO:0005900 | oncostatin-M receptor complex(GO:0005900) |
| 0.3 | 0.3 | GO:0098645 | network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) |
| 0.3 | 0.8 | GO:0043259 | laminin-10 complex(GO:0043259) |
| 0.3 | 1.1 | GO:0043218 | compact myelin(GO:0043218) |
| 0.2 | 1.0 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.2 | 1.3 | GO:0001527 | microfibril(GO:0001527) |
| 0.2 | 0.6 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.2 | 0.7 | GO:0030673 | axolemma(GO:0030673) |
| 0.2 | 0.5 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.2 | 0.5 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.2 | 0.5 | GO:0042597 | outer membrane-bounded periplasmic space(GO:0030288) periplasmic space(GO:0042597) |
| 0.2 | 1.3 | GO:0030122 | AP-2 adaptor complex(GO:0030122) clathrin coat of endocytic vesicle(GO:0030128) |
| 0.2 | 4.2 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 2.4 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 0.7 | GO:0005916 | fascia adherens(GO:0005916) |
| 0.1 | 0.4 | GO:0002141 | stereocilia coupling link(GO:0002139) stereocilia ankle link(GO:0002141) stereocilia ankle link complex(GO:0002142) |
| 0.1 | 0.5 | GO:0005606 | laminin-1 complex(GO:0005606) |
| 0.1 | 2.9 | GO:0001533 | cornified envelope(GO:0001533) |
| 0.1 | 1.1 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 0.1 | GO:0005862 | muscle thin filament tropomyosin(GO:0005862) |
| 0.1 | 0.6 | GO:0071437 | invadopodium(GO:0071437) |
| 0.1 | 0.7 | GO:0005652 | nuclear lamina(GO:0005652) |
| 0.1 | 0.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.1 | 2.5 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 0.1 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.1 | 0.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.3 | GO:0032449 | CBM complex(GO:0032449) |
| 0.1 | 0.3 | GO:0031512 | motile primary cilium(GO:0031512) |
| 0.1 | 23.6 | GO:0005578 | proteinaceous extracellular matrix(GO:0005578) |
| 0.1 | 0.5 | GO:0000805 | X chromosome(GO:0000805) |
| 0.1 | 7.5 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.1 | 1.1 | GO:0005858 | axonemal dynein complex(GO:0005858) |
| 0.1 | 0.6 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.1 | 0.2 | GO:0061202 | clathrin-sculpted gamma-aminobutyric acid transport vesicle(GO:0061200) clathrin-sculpted gamma-aminobutyric acid transport vesicle membrane(GO:0061202) |
| 0.1 | 0.2 | GO:0014704 | intercalated disc(GO:0014704) cell-cell contact zone(GO:0044291) |
| 0.1 | 0.3 | GO:0036454 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) |
| 0.1 | 0.4 | GO:0031232 | extrinsic component of external side of plasma membrane(GO:0031232) |
| 0.1 | 0.5 | GO:0032589 | neuron projection membrane(GO:0032589) |
| 0.1 | 0.3 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 1.7 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 1.4 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.1 | 1.2 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.1 | 0.6 | GO:0030118 | clathrin coat(GO:0030118) |
| 0.1 | 0.5 | GO:0043034 | costamere(GO:0043034) |
| 0.1 | 0.1 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.1 | 3.3 | GO:0030426 | growth cone(GO:0030426) |
| 0.1 | 0.6 | GO:0008328 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.1 | 1.7 | GO:0030173 | integral component of Golgi membrane(GO:0030173) |
| 0.1 | 1.4 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 0.4 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.1 | 0.6 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.1 | 0.3 | GO:0001739 | sex chromatin(GO:0001739) |
| 0.0 | 0.3 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.0 | 2.4 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 3.6 | GO:0005923 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.0 | 0.3 | GO:0090576 | RNA polymerase III transcription factor complex(GO:0090576) |
| 0.0 | 0.3 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.3 | GO:0016011 | dystroglycan complex(GO:0016011) |
| 0.0 | 0.6 | GO:0042383 | sarcolemma(GO:0042383) |
| 0.0 | 2.3 | GO:0099572 | postsynaptic density(GO:0014069) postsynaptic specialization(GO:0099572) |
| 0.0 | 0.5 | GO:0000299 | obsolete integral to membrane of membrane fraction(GO:0000299) |
| 0.0 | 1.4 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.1 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.0 | 0.1 | GO:0031143 | pseudopodium(GO:0031143) |
| 0.0 | 0.2 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.0 | 20.4 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 0.2 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) |
| 0.0 | 0.5 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 0.3 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.2 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.0 | 0.1 | GO:0044447 | axoneme part(GO:0044447) |
| 0.0 | 0.2 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.0 | 0.6 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.1 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.0 | 0.1 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.0 | 0.2 | GO:0030315 | T-tubule(GO:0030315) |
| 0.0 | 2.3 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.0 | 0.0 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.0 | 0.4 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.1 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.4 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.2 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 0.0 | GO:0071664 | catenin-TCF7L2 complex(GO:0071664) |
| 0.0 | 1.9 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 0.5 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.1 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.0 | 2.4 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.1 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.0 | 0.1 | GO:0000801 | central element(GO:0000801) |
| 0.0 | 0.2 | GO:0032156 | septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.1 | GO:0031211 | palmitoyltransferase complex(GO:0002178) serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.1 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.0 | 0.1 | GO:0042105 | alpha-beta T cell receptor complex(GO:0042105) |
| 0.0 | 0.8 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 0.1 | GO:0045323 | interleukin-1 receptor complex(GO:0045323) |
| 0.0 | 0.4 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 0.2 | GO:0031526 | brush border membrane(GO:0031526) |
| 0.0 | 1.4 | GO:0036477 | somatodendritic compartment(GO:0036477) |
| 0.0 | 14.1 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.0 | 0.6 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 0.1 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.0 | 0.9 | GO:0001726 | ruffle(GO:0001726) |
| 0.0 | 0.2 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 0.0 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.0 | 0.1 | GO:0031091 | platelet alpha granule(GO:0031091) |
| 0.0 | 0.0 | GO:0001939 | female pronucleus(GO:0001939) |
| 0.0 | 0.0 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.0 | 12.5 | GO:0005576 | extracellular region(GO:0005576) |
| 0.0 | 0.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.1 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.0 | 0.0 | GO:0001950 | obsolete plasma membrane enriched fraction(GO:0001950) |
| 0.0 | 2.8 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 0.0 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 0.1 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 0.5 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.0 | 0.1 | GO:0016580 | Sin3 complex(GO:0016580) Sin3-type complex(GO:0070822) |
| 0.0 | 0.1 | GO:0070652 | HAUS complex(GO:0070652) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.8 | 3.0 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.7 | 2.1 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.6 | 1.7 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.5 | 3.1 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.5 | 4.0 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.5 | 1.4 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.5 | 1.4 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.4 | 1.3 | GO:0004117 | calmodulin-dependent cyclic-nucleotide phosphodiesterase activity(GO:0004117) |
| 0.4 | 1.1 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.4 | 1.1 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.3 | 1.0 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.3 | 1.0 | GO:0008061 | chitin binding(GO:0008061) |
| 0.3 | 1.0 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.3 | 1.6 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.3 | 1.2 | GO:0005010 | insulin-like growth factor-activated receptor activity(GO:0005010) |
| 0.3 | 0.9 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.3 | 5.3 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) transmembrane receptor protein phosphatase activity(GO:0019198) |
| 0.3 | 0.9 | GO:0050544 | arachidonic acid binding(GO:0050544) |
| 0.3 | 1.2 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.3 | 1.5 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.3 | 1.5 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.3 | 1.1 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 2.4 | GO:0048407 | platelet-derived growth factor binding(GO:0048407) |
| 0.3 | 0.8 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.2 | 0.7 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.2 | 0.9 | GO:0031545 | procollagen-proline 4-dioxygenase activity(GO:0004656) peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.2 | 0.7 | GO:0004936 | alpha-adrenergic receptor activity(GO:0004936) |
| 0.2 | 2.6 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.2 | 0.5 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.2 | 0.7 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.2 | 0.4 | GO:0030955 | potassium ion binding(GO:0030955) |
| 0.2 | 2.0 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.2 | 0.6 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.2 | 1.1 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.2 | 1.9 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.2 | 0.4 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.2 | 0.4 | GO:0043495 | protein anchor(GO:0043495) |
| 0.2 | 0.6 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.2 | 1.7 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.2 | 1.3 | GO:0005110 | frizzled-2 binding(GO:0005110) |
| 0.2 | 0.6 | GO:0097493 | extracellular matrix constituent conferring elasticity(GO:0030023) structural molecule activity conferring elasticity(GO:0097493) |
| 0.2 | 0.5 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.2 | 4.1 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.2 | 0.5 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.2 | 0.2 | GO:0001871 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.2 | 0.5 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.2 | 0.5 | GO:0050698 | proteoglycan sulfotransferase activity(GO:0050698) |
| 0.2 | 0.5 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.2 | 1.9 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.2 | 0.3 | GO:0030172 | troponin C binding(GO:0030172) |
| 0.2 | 1.3 | GO:0043395 | heparan sulfate proteoglycan binding(GO:0043395) |
| 0.2 | 1.1 | GO:0044390 | ubiquitin conjugating enzyme binding(GO:0031624) ubiquitin-like protein conjugating enzyme binding(GO:0044390) |
| 0.2 | 1.1 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.2 | 0.6 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 0.7 | GO:0005078 | MAP-kinase scaffold activity(GO:0005078) |
| 0.1 | 10.5 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 0.6 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.1 | 0.1 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) |
| 0.1 | 0.1 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.1 | 0.7 | GO:0008517 | folic acid transporter activity(GO:0008517) |
| 0.1 | 0.8 | GO:0050998 | nitric-oxide synthase binding(GO:0050998) |
| 0.1 | 0.8 | GO:0015174 | basic amino acid transmembrane transporter activity(GO:0015174) |
| 0.1 | 2.9 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 0.5 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.1 | 0.3 | GO:0070644 | vitamin D response element binding(GO:0070644) |
| 0.1 | 0.7 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 0.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.1 | 0.8 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 0.7 | GO:0030618 | transforming growth factor beta receptor, pathway-specific cytoplasmic mediator activity(GO:0030618) |
| 0.1 | 1.0 | GO:0000982 | transcription factor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0000982) transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.1 | 1.3 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) |
| 0.1 | 1.2 | GO:0050780 | dopamine receptor binding(GO:0050780) |
| 0.1 | 0.5 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.1 | 0.9 | GO:0030506 | ankyrin binding(GO:0030506) |
| 0.1 | 0.6 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 0.5 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.1 | 0.5 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.1 | 0.5 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.1 | 0.5 | GO:0004668 | protein-arginine deiminase activity(GO:0004668) |
| 0.1 | 0.8 | GO:0004716 | receptor signaling protein tyrosine kinase activity(GO:0004716) |
| 0.1 | 1.1 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 1.7 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.1 | 1.1 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.1 | 3.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 0.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 0.1 | GO:0070696 | transmembrane receptor protein serine/threonine kinase binding(GO:0070696) |
| 0.1 | 0.4 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.1 | 0.3 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.1 | 0.5 | GO:0031435 | mitogen-activated protein kinase kinase kinase binding(GO:0031435) |
| 0.1 | 0.4 | GO:0017112 | Rab guanyl-nucleotide exchange factor activity(GO:0017112) |
| 0.1 | 0.3 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 0.5 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.1 | 0.8 | GO:0042895 | antibiotic transporter activity(GO:0042895) |
| 0.1 | 0.1 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.1 | 0.4 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.1 | 0.3 | GO:0050682 | AF-2 domain binding(GO:0050682) |
| 0.1 | 0.5 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 0.3 | GO:0008475 | procollagen-lysine 5-dioxygenase activity(GO:0008475) |
| 0.1 | 0.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.1 | 0.4 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.1 | 0.4 | GO:0048027 | mRNA 5'-UTR binding(GO:0048027) |
| 0.1 | 0.5 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.1 | 0.7 | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity(GO:0016742) |
| 0.1 | 0.3 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.1 | 1.0 | GO:0030553 | cGMP binding(GO:0030553) |
| 0.1 | 0.3 | GO:0042978 | ornithine decarboxylase activator activity(GO:0042978) |
| 0.1 | 0.5 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.1 | 0.7 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 0.1 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.1 | 0.5 | GO:0004887 | thyroid hormone receptor activity(GO:0004887) |
| 0.1 | 0.4 | GO:0031013 | troponin I binding(GO:0031013) |
| 0.1 | 1.6 | GO:0008066 | glutamate receptor activity(GO:0008066) |
| 0.1 | 0.8 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.1 | 0.3 | GO:0004980 | melanocyte-stimulating hormone receptor activity(GO:0004980) |
| 0.1 | 0.5 | GO:0035240 | dopamine binding(GO:0035240) |
| 0.1 | 1.9 | GO:0005272 | sodium channel activity(GO:0005272) |
| 0.1 | 0.4 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 0.6 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 0.9 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.1 | 0.6 | GO:0003706 | obsolete ligand-regulated transcription factor activity(GO:0003706) |
| 0.1 | 0.2 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.1 | 0.3 | GO:0015379 | potassium:chloride symporter activity(GO:0015379) |
| 0.1 | 0.5 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 9.2 | GO:0005267 | potassium channel activity(GO:0005267) |
| 0.1 | 0.5 | GO:0051635 | obsolete bacterial cell surface binding(GO:0051635) |
| 0.1 | 0.2 | GO:0004735 | pyrroline-5-carboxylate reductase activity(GO:0004735) |
| 0.1 | 0.1 | GO:0005234 | extracellular-glutamate-gated ion channel activity(GO:0005234) |
| 0.1 | 0.9 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 0.2 | GO:0070548 | L-glutamine aminotransferase activity(GO:0070548) |
| 0.1 | 0.2 | GO:0004967 | glucagon receptor activity(GO:0004967) |
| 0.1 | 0.3 | GO:0004064 | arylesterase activity(GO:0004064) |
| 0.1 | 3.7 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.1 | 1.1 | GO:0004407 | histone deacetylase activity(GO:0004407) |
| 0.1 | 0.4 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.1 | 0.3 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.1 | 0.3 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.1 | 0.3 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.1 | 0.2 | GO:0000247 | C-8 sterol isomerase activity(GO:0000247) |
| 0.1 | 0.2 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.3 | GO:0008486 | diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) |
| 0.1 | 0.1 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) |
| 0.1 | 0.8 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 0.1 | 0.2 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.1 | 0.7 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.1 | 1.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 0.4 | GO:0030346 | protein phosphatase 2B binding(GO:0030346) |
| 0.1 | 0.9 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.1 | 0.7 | GO:0001619 | obsolete lysosphingolipid and lysophosphatidic acid receptor activity(GO:0001619) |
| 0.1 | 0.3 | GO:0008131 | primary amine oxidase activity(GO:0008131) |
| 0.1 | 0.5 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 0.4 | GO:0034235 | GPI anchor binding(GO:0034235) |
| 0.1 | 0.2 | GO:0043533 | inositol 1,3,4,5 tetrakisphosphate binding(GO:0043533) |
| 0.1 | 0.3 | GO:0008296 | 3'-5'-exodeoxyribonuclease activity(GO:0008296) |
| 0.1 | 0.2 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 0.2 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.1 | 8.5 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 0.5 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.1 | 0.2 | GO:0034187 | obsolete apolipoprotein E binding(GO:0034187) |
| 0.1 | 0.8 | GO:0008373 | sialyltransferase activity(GO:0008373) |
| 0.1 | 0.2 | GO:0004321 | fatty-acyl-CoA synthase activity(GO:0004321) |
| 0.1 | 0.3 | GO:0070290 | N-acylphosphatidylethanolamine-specific phospholipase D activity(GO:0070290) |
| 0.1 | 0.2 | GO:0004103 | choline kinase activity(GO:0004103) |
| 0.1 | 0.2 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.1 | 0.2 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 0.2 | GO:0016004 | phospholipase activator activity(GO:0016004) |
| 0.1 | 0.3 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.1 | 0.2 | GO:0001012 | RNA polymerase II regulatory region sequence-specific DNA binding(GO:0000977) RNA polymerase II regulatory region DNA binding(GO:0001012) |
| 0.1 | 0.9 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.1 | 0.3 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) potassium ion antiporter activity(GO:0022821) |
| 0.1 | 0.1 | GO:0019855 | calcium channel inhibitor activity(GO:0019855) |
| 0.1 | 0.2 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.1 | 0.4 | GO:0005003 | ephrin receptor activity(GO:0005003) |
| 0.1 | 0.2 | GO:0015137 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.1 | 0.4 | GO:0004954 | prostanoid receptor activity(GO:0004954) |
| 0.0 | 0.1 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) |
| 0.0 | 0.1 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.0 | 0.1 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.0 | 0.6 | GO:0005355 | glucose transmembrane transporter activity(GO:0005355) |
| 0.0 | 0.1 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.0 | 0.4 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.7 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.3 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.2 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.2 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) |
| 0.0 | 0.1 | GO:0070576 | vitamin D 24-hydroxylase activity(GO:0070576) |
| 0.0 | 0.0 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.0 | 0.3 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.3 | GO:0033613 | activating transcription factor binding(GO:0033613) |
| 0.0 | 0.1 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.0 | 0.3 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.0 | 0.5 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.3 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.2 | GO:0046870 | cadmium ion binding(GO:0046870) |
| 0.0 | 0.1 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.0 | 0.1 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.0 | 0.7 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.5 | GO:0005243 | gap junction channel activity(GO:0005243) |
| 0.0 | 0.4 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.1 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) |
| 0.0 | 0.1 | GO:0030881 | beta-2-microglobulin binding(GO:0030881) |
| 0.0 | 0.5 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.2 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.0 | 0.0 | GO:0000976 | transcription regulatory region sequence-specific DNA binding(GO:0000976) |
| 0.0 | 0.1 | GO:0019237 | centromeric DNA binding(GO:0019237) |
| 0.0 | 0.4 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.0 | 0.2 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.1 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.0 | 0.6 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.0 | 0.1 | GO:0005346 | purine ribonucleotide transmembrane transporter activity(GO:0005346) |
| 0.0 | 0.2 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.0 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.0 | 0.3 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.0 | 0.2 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.0 | 0.2 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.0 | 0.0 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.0 | 3.3 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.2 | GO:0005041 | low-density lipoprotein receptor activity(GO:0005041) |
| 0.0 | 9.2 | GO:0030528 | obsolete transcription regulator activity(GO:0030528) |
| 0.0 | 0.2 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.0 | 0.2 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.0 | 2.0 | GO:0005089 | Rho guanyl-nucleotide exchange factor activity(GO:0005089) |
| 0.0 | 0.2 | GO:0016717 | oxidoreductase activity, acting on paired donors, with oxidation of a pair of donors resulting in the reduction of molecular oxygen to two molecules of water(GO:0016717) |
| 0.0 | 0.5 | GO:0004653 | polypeptide N-acetylgalactosaminyltransferase activity(GO:0004653) |
| 0.0 | 0.3 | GO:0003709 | obsolete RNA polymerase III transcription factor activity(GO:0003709) |
| 0.0 | 0.1 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.0 | 1.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.0 | 0.2 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0050508 | glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity(GO:0050508) |
| 0.0 | 0.1 | GO:0016934 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.0 | 0.1 | GO:0042285 | UDP-xylosyltransferase activity(GO:0035252) xylosyltransferase activity(GO:0042285) |
| 0.0 | 0.6 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.0 | 0.0 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 0.2 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.1 | GO:0043208 | glycosphingolipid binding(GO:0043208) |
| 0.0 | 0.1 | GO:0030151 | molybdenum ion binding(GO:0030151) |
| 0.0 | 0.0 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.0 | 0.1 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.0 | 0.2 | GO:0004629 | phospholipase C activity(GO:0004629) |
| 0.0 | 0.1 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.0 | 0.1 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) xenobiotic transporter activity(GO:0042910) |
| 0.0 | 0.0 | GO:0015226 | amino-acid betaine transmembrane transporter activity(GO:0015199) carnitine transmembrane transporter activity(GO:0015226) |
| 0.0 | 0.6 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.2 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.0 | 0.0 | GO:0042165 | neurotransmitter binding(GO:0042165) |
| 0.0 | 0.1 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) |
| 0.0 | 0.0 | GO:0030492 | hemoglobin binding(GO:0030492) |
| 0.0 | 0.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.4 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0099589 | G-protein coupled serotonin receptor activity(GO:0004993) serotonin receptor activity(GO:0099589) |
| 0.0 | 0.0 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.0 | 0.1 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.0 | 0.2 | GO:0005337 | nucleoside transmembrane transporter activity(GO:0005337) |
| 0.0 | 0.2 | GO:0070405 | ammonium ion binding(GO:0070405) |
| 0.0 | 0.1 | GO:0015651 | quaternary ammonium group transmembrane transporter activity(GO:0015651) |
| 0.0 | 0.1 | GO:0003854 | 3-beta-hydroxy-delta5-steroid dehydrogenase activity(GO:0003854) |
| 0.0 | 0.4 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.7 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 0.3 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.1 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.0 | 0.0 | GO:0034713 | type I transforming growth factor beta receptor binding(GO:0034713) |
| 0.0 | 0.2 | GO:0005024 | transforming growth factor beta-activated receptor activity(GO:0005024) |
| 0.0 | 0.1 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.0 | 1.2 | GO:0003705 | transcription factor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0003705) |
| 0.0 | 0.0 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.0 | 2.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.0 | GO:0031702 | type 1 angiotensin receptor binding(GO:0031702) |
| 0.0 | 0.1 | GO:0019966 | interleukin-1 binding(GO:0019966) |
| 0.0 | 0.2 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.2 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.0 | 0.2 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.0 | 0.1 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.1 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.0 | 0.9 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 0.1 | GO:0080031 | methyl indole-3-acetate esterase activity(GO:0080030) methyl salicylate esterase activity(GO:0080031) methyl jasmonate esterase activity(GO:0080032) |
| 0.0 | 0.1 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.0 | 0.1 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.0 | 0.0 | GO:0019213 | deacetylase activity(GO:0019213) |
| 0.0 | 0.1 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.0 | 0.1 | GO:0001595 | angiotensin receptor activity(GO:0001595) angiotensin type II receptor activity(GO:0004945) |
| 0.0 | 0.1 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.0 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 2.8 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.1 | GO:0004470 | malic enzyme activity(GO:0004470) |
| 0.0 | 0.0 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.0 | 0.0 | GO:0043398 | HLH domain binding(GO:0043398) |
| 0.0 | 0.2 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.9 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 0.0 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.0 | 0.0 | GO:0016149 | translation release factor activity, codon specific(GO:0016149) |
| 0.0 | 0.1 | GO:0016646 | oxidoreductase activity, acting on the CH-NH group of donors, NAD or NADP as acceptor(GO:0016646) |
| 0.0 | 0.1 | GO:0043499 | obsolete eukaryotic cell surface binding(GO:0043499) |
| 0.0 | 0.1 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.0 | GO:0019959 | C-X-C chemokine binding(GO:0019958) interleukin-8 binding(GO:0019959) |
| 0.0 | 0.0 | GO:0004052 | arachidonate 12-lipoxygenase activity(GO:0004052) |
| 0.0 | 0.1 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.0 | 0.0 | GO:0030976 | thiamine pyrophosphate binding(GO:0030976) |
| 0.0 | 0.1 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 0.8 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.1 | GO:0005344 | oxygen transporter activity(GO:0005344) |
| 0.0 | 0.1 | GO:0030742 | GTP-dependent protein binding(GO:0030742) |
| 0.0 | 0.0 | GO:0032139 | dinucleotide insertion or deletion binding(GO:0032139) |
| 0.0 | 4.9 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.8 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.9 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.3 | 4.1 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.3 | 10.2 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.2 | 0.3 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.2 | 0.3 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.1 | 2.1 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 1.7 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 3.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 0.4 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.1 | 1.3 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 2.8 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.1 | 15.6 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 3.2 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 0.9 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 1.2 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 2.9 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.1 | 1.6 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.1 | 1.1 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 0.8 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 0.9 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 0.3 | PID A6B1 A6B4 INTEGRIN PATHWAY | a6b1 and a6b4 Integrin signaling |
| 0.1 | 0.9 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.1 | 0.5 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 0.2 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 11.4 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.1 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 0.6 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.0 | 0.5 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 7.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.8 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 1.0 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.6 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.1 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 0.1 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.5 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 1.0 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.1 | PID EPO PATHWAY | EPO signaling pathway |
| 0.0 | 10.9 | NABA MATRISOME ASSOCIATED | Ensemble of genes encoding ECM-associated proteins including ECM-affilaited proteins, ECM regulators and secreted factors |
| 0.0 | 0.1 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.0 | 0.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.4 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.0 | 1.0 | NABA MATRISOME | Ensemble of genes encoding extracellular matrix and extracellular matrix-associated proteins |
| 0.0 | 0.2 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 0.1 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.0 | 0.9 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.2 | PID ENDOTHELIN PATHWAY | Endothelins |
| 0.0 | 0.1 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.8 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.0 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 0.2 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.1 | PID KIT PATHWAY | Signaling events mediated by Stem cell factor receptor (c-Kit) |
| 0.0 | 0.6 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.3 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.0 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.0 | 0.3 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.1 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.0 | 0.1 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.2 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.3 | 2.9 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.3 | 2.6 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.2 | 2.4 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.2 | 2.5 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 0.2 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.2 | 5.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.2 | 9.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.2 | 2.0 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.2 | 0.6 | REACTOME PROLACTIN RECEPTOR SIGNALING | Genes involved in Prolactin receptor signaling |
| 0.1 | 1.8 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.1 | 0.4 | REACTOME GPCR LIGAND BINDING | Genes involved in GPCR ligand binding |
| 0.1 | 5.5 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.1 | 1.8 | REACTOME GRB2 EVENTS IN ERBB2 SIGNALING | Genes involved in GRB2 events in ERBB2 signaling |
| 0.1 | 0.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.1 | 3.1 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 1.4 | REACTOME SYNTHESIS SECRETION AND DEACYLATION OF GHRELIN | Genes involved in Synthesis, Secretion, and Deacylation of Ghrelin |
| 0.1 | 2.1 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 0.9 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 0.5 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.1 | 1.1 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.1 | 1.0 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.1 | 1.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 0.6 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.1 | 0.2 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.1 | 1.0 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 1.0 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 1.4 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.1 | 1.7 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 0.2 | REACTOME ARMS MEDIATED ACTIVATION | Genes involved in ARMS-mediated activation |
| 0.1 | 2.6 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 0.7 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 1.4 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 0.2 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.1 | 0.5 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 0.8 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 0.1 | 0.1 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.1 | 0.6 | REACTOME ACTIVATION OF NMDA RECEPTOR UPON GLUTAMATE BINDING AND POSTSYNAPTIC EVENTS | Genes involved in Activation of NMDA receptor upon glutamate binding and postsynaptic events |
| 0.1 | 1.1 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.1 | 0.2 | REACTOME INCRETIN SYNTHESIS SECRETION AND INACTIVATION | Genes involved in Incretin Synthesis, Secretion, and Inactivation |
| 0.1 | 0.4 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.0 | 2.3 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.7 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.5 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.6 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.7 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 1.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.7 | REACTOME CHONDROITIN SULFATE BIOSYNTHESIS | Genes involved in Chondroitin sulfate biosynthesis |
| 0.0 | 0.0 | REACTOME OPSINS | Genes involved in Opsins |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELL ADHESION PROTEINS | Genes involved in Apoptotic cleavage of cell adhesion proteins |
| 0.0 | 0.6 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 2.8 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 1.7 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.0 | REACTOME ADP SIGNALLING THROUGH P2RY1 | Genes involved in ADP signalling through P2Y purinoceptor 1 |
| 0.0 | 0.7 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.6 | REACTOME PLC BETA MEDIATED EVENTS | Genes involved in PLC beta mediated events |
| 0.0 | 0.4 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.0 | 0.1 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.0 | 0.5 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.3 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.0 | 0.3 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 2.9 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.2 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |
| 0.0 | 0.2 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.0 | 0.3 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.3 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 0.2 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 0.2 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.0 | 0.0 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.0 | 0.3 | REACTOME REGULATION OF KIT SIGNALING | Genes involved in Regulation of KIT signaling |
| 0.0 | 0.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.3 | REACTOME OPIOID SIGNALLING | Genes involved in Opioid Signalling |
| 0.0 | 0.0 | REACTOME TRANS GOLGI NETWORK VESICLE BUDDING | Genes involved in trans-Golgi Network Vesicle Budding |
| 0.0 | 3.7 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.5 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.2 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.0 | 0.1 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.1 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.0 | 0.1 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.0 | 0.2 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.4 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.0 | 0.3 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.1 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.0 | 0.2 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.0 | 1.2 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 0.0 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.0 | 0.1 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.0 | 0.2 | REACTOME HORMONE LIGAND BINDING RECEPTORS | Genes involved in Hormone ligand-binding receptors |
| 0.0 | 0.2 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.1 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.4 | REACTOME EXTRACELLULAR MATRIX ORGANIZATION | Genes involved in Extracellular matrix organization |
| 0.0 | 0.6 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 0.1 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.1 | REACTOME ABACAVIR TRANSPORT AND METABOLISM | Genes involved in Abacavir transport and metabolism |
| 0.0 | 0.0 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.0 | 0.1 | REACTOME ACTIVATION OF KAINATE RECEPTORS UPON GLUTAMATE BINDING | Genes involved in Activation of Kainate Receptors upon glutamate binding |
| 0.0 | 0.2 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.1 | REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Highly calcium permeable postsynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.1 | REACTOME REGULATION OF INSULIN SECRETION | Genes involved in Regulation of Insulin Secretion |
| 0.0 | 0.1 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.1 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |