Project
ENCODE: H3K4me3 ChIP-Seq of different mouse tissues during embryonic development
Navigation
Downloads
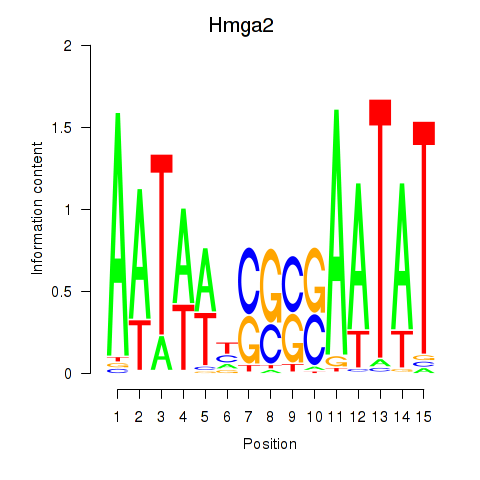
Results for Hmga2
Z-value: 3.25
Transcription factors associated with Hmga2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Hmga2
|
ENSMUSG00000056758.8 | high mobility group AT-hook 2 |
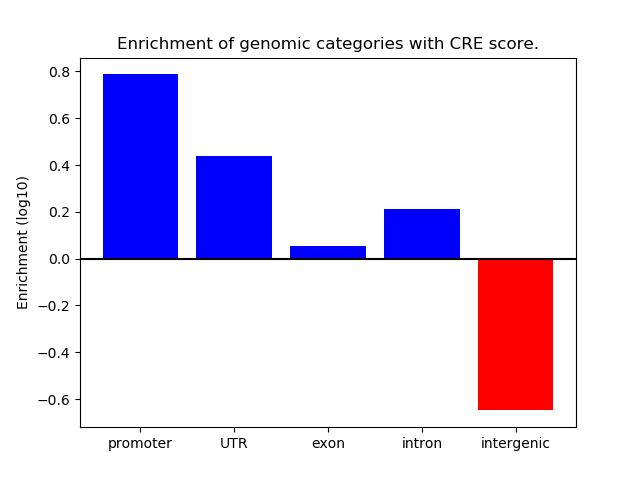
Correlations of motif activity and signal intensity at CREs associated with the motif's TFs:
This plot shows correlation between observed signal intensity of a CRE associated with the transcription factor across all samples and activity of the motif.
For each TF, only the top 5 correlated CREs are shown.
| CRE | Gene | Distance | Association probability | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|---|---|
| chr10_120474317_120474690 | Hmga2 | 1789 | 0.363089 | -0.62 | 1.6e-07 | Click! |
| chr10_120477592_120477743 | Hmga2 | 1198 | 0.496402 | -0.55 | 5.7e-06 | Click! |
| chr10_120473161_120473721 | Hmga2 | 2851 | 0.268641 | -0.37 | 3.8e-03 | Click! |
| chr10_120478339_120478490 | Hmga2 | 1945 | 0.345705 | -0.32 | 1.4e-02 | Click! |
| chr10_120475064_120476270 | Hmga2 | 625 | 0.739650 | -0.27 | 3.9e-02 | Click! |
Activity of the Hmga2 motif across conditions
Conditions sorted by the z-value of the Hmga2 motif activity
Move your cursor over a bar to see sample name and corresponding Z-value.
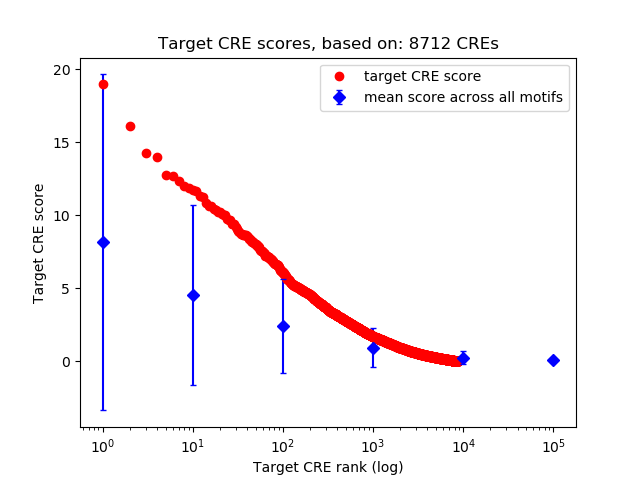
Top target CREs of the motif:
| Cis Regulatory Element (CRE) | Target Score | Top associated gene | Gene Info | Distance of CRE to TSS | CRE/Gene association probability |
|---|---|---|---|---|---|
| chr1_42700192_42700666 | 19.03 |
Pou3f3 |
POU domain, class 3, transcription factor 3 |
4661 |
0.15 |
| chr1_66386919_66387899 | 16.14 |
Map2 |
microtubule-associated protein 2 |
398 |
0.87 |
| chr12_88724589_88725423 | 14.24 |
Nrxn3 |
neurexin III |
3 |
0.98 |
| chr6_134886811_134888239 | 14.01 |
Gpr19 |
G protein-coupled receptor 19 |
243 |
0.87 |
| chrX_61116034_61117613 | 12.79 |
Cdr1os |
cerebellar degeneration related antigen 1, opposite strand |
425 |
0.47 |
| chr3_88213113_88214199 | 12.71 |
Gm3764 |
predicted gene 3764 |
829 |
0.3 |
| chr4_22484307_22484937 | 12.34 |
Pou3f2 |
POU domain, class 3, transcription factor 2 |
3744 |
0.2 |
| chr7_92234907_92236280 | 12.00 |
Dlg2 |
discs large MAGUK scaffold protein 2 |
466 |
0.88 |
| chr3_68573207_68574269 | 11.87 |
Schip1 |
schwannomin interacting protein 1 |
1493 |
0.45 |
| chr6_55678280_55679200 | 11.74 |
Neurod6 |
neurogenic differentiation 6 |
2523 |
0.32 |
| chr13_83715222_83716973 | 11.66 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
5284 |
0.15 |
| chr4_22485878_22486449 | 11.35 |
Pou3f2 |
POU domain, class 3, transcription factor 2 |
2203 |
0.26 |
| chr6_55680133_55680881 | 11.25 |
Neurod6 |
neurogenic differentiation 6 |
756 |
0.69 |
| chr7_51621596_51622924 | 10.86 |
Slc17a6 |
solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 |
39 |
0.98 |
| chr2_136715590_136716138 | 10.66 |
Snap25 |
synaptosomal-associated protein 25 |
2386 |
0.31 |
| chr14_108909943_108910203 | 10.66 |
Slitrk1 |
SLIT and NTRK-like family, member 1 |
4085 |
0.37 |
| chr14_39471112_39471496 | 10.43 |
Nrg3 |
neuregulin 3 |
1362 |
0.61 |
| chr13_83727321_83728283 | 10.34 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
304 |
0.83 |
| chr3_34652462_34653573 | 10.23 |
Sox2 |
SRY (sex determining region Y)-box 2 |
2612 |
0.16 |
| chr13_83717521_83718816 | 10.20 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
3213 |
0.17 |
| chr10_29143400_29144848 | 10.13 |
Soga3 |
SOGA family member 3 |
65 |
0.5 |
| chr18_23038497_23038648 | 10.06 |
Nol4 |
nucleolar protein 4 |
84 |
0.99 |
| chr2_18042311_18043883 | 10.04 |
Skida1 |
SKI/DACH domain containing 1 |
1475 |
0.25 |
| chr9_96731522_96733329 | 9.79 |
Zbtb38 |
zinc finger and BTB domain containing 38 |
244 |
0.91 |
| chr9_41585694_41587243 | 9.71 |
Mir100hg |
Mir100 Mirlet7a-2 Mir125b-1 cluster host gene |
1301 |
0.29 |
| chr10_90830503_90831025 | 9.67 |
Anks1b |
ankyrin repeat and sterile alpha motif domain containing 1B |
921 |
0.56 |
| chrX_75577044_75577513 | 9.42 |
Rab39b |
RAB39B, member RAS oncogene family |
953 |
0.4 |
| chr15_39392966_39393260 | 9.40 |
Rims2 |
regulating synaptic membrane exocytosis 2 |
1313 |
0.57 |
| chr9_91227777_91228605 | 9.37 |
Gm29602 |
predicted gene 29602 |
12176 |
0.18 |
| chr4_9270530_9270692 | 9.17 |
Clvs1 |
clavesin 1 |
844 |
0.67 |
| chr1_6734529_6735444 | 9.05 |
St18 |
suppression of tumorigenicity 18 |
116 |
0.98 |
| chr12_72234504_72235243 | 8.95 |
Rtn1 |
reticulon 1 |
866 |
0.66 |
| chr13_83742301_83742867 | 8.89 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
3721 |
0.15 |
| chr2_6881874_6882908 | 8.75 |
Gm13389 |
predicted gene 13389 |
1879 |
0.3 |
| chr14_75963713_75963971 | 8.71 |
Gm25517 |
predicted gene, 25517 |
8777 |
0.18 |
| chr6_80020534_80021026 | 8.66 |
Lrrtm4 |
leucine rich repeat transmembrane neuronal 4 |
1137 |
0.48 |
| chr3_17793835_17795104 | 8.65 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
427 |
0.75 |
| chr1_23764777_23765091 | 8.65 |
B3gat2 |
beta-1,3-glucuronyltransferase 2 (glucuronosyltransferase S) |
2923 |
0.36 |
| chr1_9601164_9602408 | 8.64 |
Vxn |
vexin |
587 |
0.67 |
| chr2_178143444_178143670 | 8.56 |
Phactr3 |
phosphatase and actin regulator 3 |
1624 |
0.46 |
| chr1_81077232_81078427 | 8.55 |
Nyap2 |
neuronal tyrosine-phophorylated phosphoinositide 3-kinase adaptor 2 |
246 |
0.96 |
| chr6_39874717_39875333 | 8.39 |
Tmem178b |
transmembrane protein 178B |
1954 |
0.27 |
| chr13_110277692_110277997 | 8.36 |
Rab3c |
RAB3C, member RAS oncogene family |
2306 |
0.36 |
| chr15_85667333_85667919 | 8.27 |
Lncppara |
long noncoding RNA near Ppara |
14010 |
0.14 |
| chr17_52602081_52603198 | 8.26 |
Gm27217 |
predicted gene 27217 |
21 |
0.54 |
| chr10_49785211_49786117 | 8.21 |
Grik2 |
glutamate receptor, ionotropic, kainate 2 (beta 2) |
2255 |
0.25 |
| chr1_99772154_99773556 | 8.16 |
Cntnap5b |
contactin associated protein-like 5B |
90 |
0.98 |
| chr10_111247804_111248910 | 8.10 |
Osbpl8 |
oxysterol binding protein-like 8 |
289 |
0.91 |
| chr2_65848718_65849208 | 8.06 |
Csrnp3 |
cysteine-serine-rich nuclear protein 3 |
3108 |
0.27 |
| chrX_165326400_165326667 | 8.05 |
Glra2 |
glycine receptor, alpha 2 subunit |
860 |
0.76 |
| chr5_111421306_111422790 | 7.95 |
Gm43119 |
predicted gene 43119 |
1541 |
0.35 |
| chr9_52145956_52147029 | 7.93 |
Zc3h12c |
zinc finger CCCH type containing 12C |
21619 |
0.17 |
| chr2_105678552_105679922 | 7.93 |
Pax6 |
paired box 6 |
630 |
0.68 |
| chr10_70596836_70597226 | 7.84 |
Phyhipl |
phytanoyl-CoA hydroxylase interacting protein-like |
2051 |
0.4 |
| chr18_23036665_23037864 | 7.74 |
Nol4 |
nucleolar protein 4 |
1392 |
0.59 |
| chr7_79505833_79506958 | 7.62 |
Mir9-3 |
microRNA 9-3 |
1131 |
0.28 |
| chrX_88112223_88112880 | 7.58 |
Il1rapl1 |
interleukin 1 receptor accessory protein-like 1 |
3094 |
0.35 |
| chrX_152643367_152644550 | 7.56 |
Shroom2 |
shroom family member 2 |
34 |
0.98 |
| chr16_77646908_77647363 | 7.55 |
C130023A14Rik |
RIKEN cDNA C130023A14 gene |
795 |
0.35 |
| chr3_17796058_17796354 | 7.50 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
462 |
0.52 |
| chr15_100873048_100873688 | 7.44 |
Scn8a |
sodium channel, voltage-gated, type VIII, alpha |
1948 |
0.32 |
| chrX_110818058_110818665 | 7.39 |
Pou3f4 |
POU domain, class 3, transcription factor 4 |
4081 |
0.27 |
| chr16_77536184_77536608 | 7.36 |
Gm36963 |
predicted gene, 36963 |
3486 |
0.16 |
| chr7_62416059_62417205 | 7.24 |
Mkrn3 |
makorin, ring finger protein, 3 |
3507 |
0.2 |
| chr1_194622071_194623282 | 7.22 |
Plxna2 |
plexin A2 |
2851 |
0.26 |
| chr6_96113911_96115198 | 7.22 |
Tafa1 |
TAFA chemokine like family member 1 |
95 |
0.98 |
| chr2_181768465_181769553 | 7.20 |
Myt1 |
myelin transcription factor 1 |
1497 |
0.33 |
| chr17_91090702_91091377 | 7.15 |
Nrxn1 |
neurexin I |
1694 |
0.28 |
| chr1_157243137_157243316 | 7.13 |
Rasal2 |
RAS protein activator like 2 |
1264 |
0.52 |
| chr4_97582473_97584218 | 7.12 |
E130114P18Rik |
RIKEN cDNA E130114P18 gene |
1251 |
0.53 |
| chr6_37297951_37298469 | 7.08 |
Dgki |
diacylglycerol kinase, iota |
1427 |
0.53 |
| chr8_12400578_12402091 | 7.03 |
Gm25239 |
predicted gene, 25239 |
4931 |
0.15 |
| chr6_136169672_136170219 | 7.01 |
Grin2b |
glutamate receptor, ionotropic, NMDA2B (epsilon 2) |
1944 |
0.34 |
| chr5_20228356_20229007 | 7.01 |
Magi2 |
membrane associated guanylate kinase, WW and PDZ domain containing 2 |
495 |
0.83 |
| chr7_62366760_62367184 | 6.93 |
Magel2 |
melanoma antigen, family L, 2 |
10038 |
0.18 |
| chr8_14382368_14383445 | 6.87 |
Dlgap2 |
DLG associated protein 2 |
910 |
0.66 |
| chr3_34653590_34654523 | 6.85 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
1980 |
0.2 |
| chr6_103513736_103514218 | 6.78 |
Chl1 |
cell adhesion molecule L1-like |
2647 |
0.25 |
| chr7_128689996_128690220 | 6.76 |
Gm43580 |
predicted gene 43580 |
1573 |
0.19 |
| chr8_112570000_112570856 | 6.71 |
Cntnap4 |
contactin associated protein-like 4 |
373 |
0.76 |
| chr12_46812593_46812909 | 6.70 |
Gm48542 |
predicted gene, 48542 |
1243 |
0.48 |
| chr1_25228097_25229399 | 6.69 |
Adgrb3 |
adhesion G protein-coupled receptor B3 |
78 |
0.96 |
| chr13_99510773_99511921 | 6.67 |
Map1b |
microtubule-associated protein 1B |
5171 |
0.17 |
| chr3_34651759_34652037 | 6.67 |
Sox2 |
SRY (sex determining region Y)-box 2 |
1493 |
0.26 |
| chr2_65620767_65621991 | 6.61 |
Scn2a |
sodium channel, voltage-gated, type II, alpha |
568 |
0.82 |
| chr5_144547334_144548656 | 6.57 |
Nptx2 |
neuronal pentraxin 2 |
2093 |
0.41 |
| chr6_28827182_28827949 | 6.57 |
Lrrc4 |
leucine rich repeat containing 4 |
2780 |
0.27 |
| chr4_91372863_91373225 | 6.57 |
Elavl2 |
ELAV like RNA binding protein 1 |
14 |
0.83 |
| chr12_71048832_71049275 | 6.50 |
Arid4a |
AT rich interactive domain 4A (RBP1-like) |
712 |
0.65 |
| chr5_9723538_9723989 | 6.46 |
Grm3 |
glutamate receptor, metabotropic 3 |
1407 |
0.48 |
| chr16_77503379_77503945 | 6.43 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
3278 |
0.16 |
| chr14_122481093_122481670 | 6.31 |
Zic2 |
zinc finger protein of the cerebellum 2 |
3281 |
0.14 |
| chr3_5222045_5223052 | 6.26 |
Zfhx4 |
zinc finger homeodomain 4 |
1043 |
0.46 |
| chr2_118779656_118780774 | 6.23 |
Disp2 |
dispatched RND tramsporter family member 2 |
456 |
0.71 |
| chr4_25797578_25797990 | 6.20 |
Fut9 |
fucosyltransferase 9 |
2071 |
0.32 |
| chr14_93887737_93887976 | 6.20 |
Pcdh9 |
protocadherin 9 |
876 |
0.72 |
| chr4_110290101_110291006 | 6.20 |
Elavl4 |
ELAV like RNA binding protein 4 |
281 |
0.95 |
| chr4_33926104_33927188 | 6.14 |
Cnr1 |
cannabinoid receptor 1 (brain) |
444 |
0.88 |
| chr2_97468266_97469202 | 6.14 |
Lrrc4c |
leucine rich repeat containing 4C |
645 |
0.83 |
| chr18_42897926_42898199 | 6.08 |
Ppp2r2b |
protein phosphatase 2, regulatory subunit B, beta |
753 |
0.73 |
| chr1_66322822_66322976 | 6.07 |
Map2 |
microtubule-associated protein 2 |
797 |
0.63 |
| chr1_66175344_66176454 | 6.03 |
Map2 |
microtubule-associated protein 2 |
349 |
0.92 |
| chr6_83171941_83172736 | 6.00 |
Gm15624 |
predicted gene 15624 |
216 |
0.83 |
| chr7_36700944_36701609 | 5.96 |
Tshz3 |
teashirt zinc finger family member 3 |
3059 |
0.18 |
| chr17_51584124_51584467 | 5.94 |
Gm31143 |
predicted gene, 31143 |
49437 |
0.15 |
| chr7_51624664_51625766 | 5.91 |
Slc17a6 |
solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 |
67 |
0.98 |
| chr18_34249368_34249586 | 5.84 |
Apc |
APC, WNT signaling pathway regulator |
1779 |
0.35 |
| chr9_75683375_75684591 | 5.78 |
Scg3 |
secretogranin III |
8 |
0.97 |
| chr16_97167039_97167850 | 5.69 |
Dscam |
DS cell adhesion molecule |
3308 |
0.37 |
| chr15_30459904_30460844 | 5.68 |
Ctnnd2 |
catenin (cadherin associated protein), delta 2 |
2586 |
0.34 |
| chr12_61527334_61527568 | 5.62 |
Lrfn5 |
leucine rich repeat and fibronectin type III domain containing 5 |
3503 |
0.24 |
| chr9_91359026_91359334 | 5.61 |
Zic4 |
zinc finger protein of the cerebellum 4 |
3233 |
0.14 |
| chrX_152642489_152643012 | 5.61 |
Shroom2 |
shroom family member 2 |
1174 |
0.54 |
| chr14_122478089_122479067 | 5.59 |
Zic2 |
zinc finger protein of the cerebellum 2 |
478 |
0.68 |
| chr15_89532557_89533956 | 5.59 |
Shank3 |
SH3 and multiple ankyrin repeat domains 3 |
366 |
0.78 |
| chr13_78193022_78193812 | 5.58 |
Nr2f1 |
nuclear receptor subfamily 2, group F, member 1 |
2956 |
0.18 |
| chr15_64917271_64918226 | 5.57 |
Adcy8 |
adenylate cyclase 8 |
4523 |
0.29 |
| chr7_79536839_79536990 | 5.57 |
Gm35040 |
predicted gene, 35040 |
871 |
0.39 |
| chr9_79829898_79830916 | 5.47 |
Filip1 |
filamin A interacting protein 1 |
591 |
0.7 |
| chr1_138724629_138724941 | 5.44 |
Gm8790 |
predicted gene 8790 |
34940 |
0.14 |
| chr1_17146083_17146352 | 5.42 |
Gdap1 |
ganglioside-induced differentiation-associated-protein 1 |
749 |
0.64 |
| chr12_89815214_89815490 | 5.35 |
Nrxn3 |
neurexin III |
2869 |
0.41 |
| chr10_109008310_109009456 | 5.35 |
Syt1 |
synaptotagmin I |
217 |
0.96 |
| chr5_126768981_126769769 | 5.34 |
Gm33347 |
predicted gene, 33347 |
42091 |
0.14 |
| chr4_9270926_9271667 | 5.32 |
Clvs1 |
clavesin 1 |
159 |
0.96 |
| chr3_67892003_67892637 | 5.31 |
Iqschfp |
Iqcj and Schip1 fusion protein |
88 |
0.51 |
| chr5_114279370_114279652 | 5.31 |
Foxn4 |
forkhead box N4 |
5704 |
0.15 |
| chr11_105590925_105591452 | 5.28 |
Tanc2 |
tetratricopeptide repeat, ankyrin repeat and coiled-coil containing 2 |
1197 |
0.51 |
| chr13_83660956_83661532 | 5.26 |
Mef2c |
myocyte enhancer factor 2C |
8356 |
0.18 |
| chr18_69349496_69350227 | 5.26 |
Tcf4 |
transcription factor 4 |
917 |
0.69 |
| chr1_32175544_32175738 | 5.25 |
Khdrbs2 |
KH domain containing, RNA binding, signal transduction associated 2 |
2754 |
0.36 |
| chr8_54958887_54959474 | 5.24 |
Gm45263 |
predicted gene 45263 |
639 |
0.67 |
| chr8_109337659_109338724 | 5.21 |
Gm1943 |
predicted gene 1943 |
2673 |
0.35 |
| chr7_60170570_60171087 | 5.17 |
Gm7367 |
predicted pseudogene 7367 |
15219 |
0.13 |
| chr3_17792584_17792950 | 5.16 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
1153 |
0.46 |
| chr15_25713605_25714406 | 5.16 |
Myo10 |
myosin X |
134 |
0.97 |
| chr2_65848318_65848473 | 5.16 |
Csrnp3 |
cysteine-serine-rich nuclear protein 3 |
2540 |
0.31 |
| chr3_88208985_88210116 | 5.14 |
Gm3764 |
predicted gene 3764 |
78 |
0.92 |
| chr12_81779620_81779953 | 5.13 |
Map3k9 |
mitogen-activated protein kinase kinase kinase 9 |
1088 |
0.41 |
| chr11_77933689_77934630 | 5.13 |
Sez6 |
seizure related gene 6 |
3120 |
0.17 |
| chr12_81778878_81779163 | 5.11 |
Map3k9 |
mitogen-activated protein kinase kinase kinase 9 |
1854 |
0.28 |
| chrX_23283125_23283785 | 5.11 |
Klhl13 |
kelch-like 13 |
1374 |
0.57 |
| chr18_25750468_25751272 | 5.09 |
Celf4 |
CUGBP, Elav-like family member 4 |
1822 |
0.41 |
| chr4_91380440_91381612 | 5.08 |
Elavl2 |
ELAV like RNA binding protein 1 |
4530 |
0.22 |
| chr8_49461633_49461934 | 5.08 |
4930555F03Rik |
RIKEN cDNA 4930555F03 gene |
1540 |
0.37 |
| chr1_76637121_76637710 | 5.07 |
Mir6343 |
microRNA 6343 |
127855 |
0.06 |
| chr13_83739310_83740387 | 5.06 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
985 |
0.29 |
| chr10_87485570_87486803 | 5.03 |
Ascl1 |
achaete-scute family bHLH transcription factor 1 |
7474 |
0.2 |
| chr8_122676327_122676714 | 5.03 |
Cbfa2t3 |
CBFA2/RUNX1 translocation partner 3 |
1552 |
0.24 |
| chr16_77235848_77236417 | 5.02 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
185 |
0.96 |
| chr14_93701256_93701943 | 5.02 |
Gm48967 |
predicted gene, 48967 |
82753 |
0.1 |
| chr4_13752540_13753163 | 5.02 |
Runx1t1 |
RUNX1 translocation partner 1 |
1554 |
0.54 |
| chr15_20450538_20451616 | 5.01 |
Cdh12 |
cadherin 12 |
1812 |
0.3 |
| chr7_60001655_60001979 | 4.98 |
Snurf |
SNRPN upstream reading frame |
3232 |
0.09 |
| chr9_47533054_47533827 | 4.97 |
Cadm1 |
cell adhesion molecule 1 |
3067 |
0.25 |
| chr16_77416103_77416788 | 4.97 |
Gm38071 |
predicted gene, 38071 |
179 |
0.91 |
| chr13_84905458_84906287 | 4.95 |
Gm4059 |
predicted gene 4059 |
68445 |
0.12 |
| chr5_38151138_38153189 | 4.93 |
Nsg1 |
neuron specific gene family member 1 |
6868 |
0.16 |
| chr11_29376424_29376575 | 4.90 |
Ccdc88a |
coiled coil domain containing 88A |
2151 |
0.25 |
| chr17_91085493_91086001 | 4.89 |
Gm47307 |
predicted gene, 47307 |
2659 |
0.21 |
| chrX_88113433_88114223 | 4.88 |
Il1rapl1 |
interleukin 1 receptor accessory protein-like 1 |
1817 |
0.46 |
| chr3_110139298_110139994 | 4.88 |
Ntng1 |
netrin G1 |
3358 |
0.33 |
| chr3_34104464_34105670 | 4.87 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
609 |
0.68 |
| chr14_76420544_76421824 | 4.86 |
Tsc22d1 |
TSC22 domain family, member 1 |
2356 |
0.39 |
| chr2_124092543_124092695 | 4.86 |
Sema6d |
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D |
2650 |
0.37 |
| chr18_72347538_72348154 | 4.86 |
Dcc |
deleted in colorectal carcinoma |
3171 |
0.38 |
| chr13_83729448_83730058 | 4.84 |
Gm26803 |
predicted gene, 26803 |
171 |
0.91 |
| chr1_154725377_154725528 | 4.82 |
Cacna1e |
calcium channel, voltage-dependent, R type, alpha 1E subunit |
468 |
0.89 |
| chr3_88216674_88217001 | 4.81 |
Gm25641 |
predicted gene, 25641 |
1141 |
0.2 |
| chr3_17783692_17784517 | 4.81 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
5817 |
0.2 |
| chr1_172485277_172486996 | 4.79 |
Igsf9 |
immunoglobulin superfamily, member 9 |
3822 |
0.12 |
| chr6_107531806_107531957 | 4.79 |
Lrrn1 |
leucine rich repeat protein 1, neuronal |
2113 |
0.37 |
| chr4_13754294_13755025 | 4.78 |
Runx1t1 |
RUNX1 translocation partner 1 |
3362 |
0.37 |
| chr16_7039399_7040361 | 4.75 |
Rbfox1 |
RNA binding protein, fox-1 homolog (C. elegans) 1 |
29966 |
0.27 |
| chr13_110277341_110277607 | 4.75 |
Rab3c |
RAB3C, member RAS oncogene family |
2676 |
0.33 |
| chr2_70557968_70558533 | 4.75 |
Gad1 |
glutamate decarboxylase 1 |
3792 |
0.17 |
| chr6_13837266_13838183 | 4.75 |
Gpr85 |
G protein-coupled receptor 85 |
483 |
0.82 |
| chr11_34315414_34316667 | 4.75 |
Insyn2b |
inhibitory synaptic factor family member 2B |
1218 |
0.45 |
| chr7_140080531_140082545 | 4.74 |
Caly |
calcyon neuron-specific vesicular protein |
689 |
0.48 |
| chr3_146769678_146770812 | 4.74 |
Prkacb |
protein kinase, cAMP dependent, catalytic, beta |
16 |
0.98 |
| chr9_82831054_82831404 | 4.73 |
Irak1bp1 |
interleukin-1 receptor-associated kinase 1 binding protein 1 |
1423 |
0.53 |
| chr1_115688569_115688799 | 4.72 |
Cntnap5a |
contactin associated protein-like 5A |
3928 |
0.28 |
| chr3_17784763_17784914 | 4.71 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
5083 |
0.21 |
| chr18_80982763_80983698 | 4.70 |
Sall3 |
spalt like transcription factor 3 |
3306 |
0.17 |
| chr9_41587250_41587725 | 4.69 |
Mir100hg |
Mir100 Mirlet7a-2 Mir125b-1 cluster host gene |
282 |
0.84 |
| chrX_23282702_23282891 | 4.68 |
Klhl13 |
kelch-like 13 |
2033 |
0.45 |
| chr4_99373530_99374221 | 4.68 |
Gm37556 |
predicted gene, 37556 |
11837 |
0.17 |
| chr16_77597699_77598173 | 4.67 |
Mir99a |
microRNA 99a |
1000 |
0.31 |
| chr16_81202167_81203211 | 4.66 |
Ncam2 |
neural cell adhesion molecule 2 |
1932 |
0.44 |
| chr1_25827305_25827886 | 4.64 |
Adgrb3 |
adhesion G protein-coupled receptor B3 |
835 |
0.43 |
| chr2_158375202_158376961 | 4.64 |
Snhg11 |
small nucleolar RNA host gene 11 |
319 |
0.74 |
| chr1_154725630_154727200 | 4.64 |
Cacna1e |
calcium channel, voltage-dependent, R type, alpha 1E subunit |
59 |
0.99 |
| chr8_90958597_90958859 | 4.63 |
Chd9 |
chromodomain helicase DNA binding protein 9 |
3293 |
0.23 |
| chr1_109984209_109985108 | 4.63 |
Cdh7 |
cadherin 7, type 2 |
921 |
0.74 |
| chr12_88727306_88727457 | 4.61 |
Nrxn3 |
neurexin III |
1700 |
0.42 |
| chr2_78873863_78874409 | 4.60 |
Ube2e3 |
ubiquitin-conjugating enzyme E2E 3 |
4458 |
0.27 |
| chr13_8208366_8208661 | 4.60 |
Adarb2 |
adenosine deaminase, RNA-specific, B2 |
5591 |
0.19 |
| chr3_84953332_84953942 | 4.59 |
Fbxw7 |
F-box and WD-40 domain protein 7 |
1491 |
0.53 |
| chr13_84063384_84064052 | 4.58 |
Gm17750 |
predicted gene, 17750 |
1054 |
0.58 |
| chr8_109249737_109249899 | 4.54 |
D030068K23Rik |
RIKEN cDNA D030068K23 gene |
48 |
0.99 |
Network of associatons between targets according to the STRING database.

Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.1 | 12.2 | GO:0099558 | maintenance of synapse structure(GO:0099558) |
| 3.7 | 11.0 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 3.6 | 7.1 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 3.5 | 10.4 | GO:0021986 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 2.7 | 10.9 | GO:2000821 | regulation of grooming behavior(GO:2000821) |
| 2.7 | 2.7 | GO:0090494 | catecholamine uptake(GO:0090493) dopamine uptake(GO:0090494) |
| 2.5 | 27.0 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 2.3 | 9.2 | GO:0072051 | juxtaglomerular apparatus development(GO:0072051) |
| 2.1 | 19.0 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 2.0 | 8.2 | GO:0050923 | regulation of negative chemotaxis(GO:0050923) |
| 2.0 | 10.1 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 1.9 | 7.7 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 1.8 | 12.9 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 1.8 | 16.6 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 1.8 | 7.3 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 1.8 | 10.8 | GO:0090273 | regulation of somatostatin secretion(GO:0090273) |
| 1.8 | 7.0 | GO:0033602 | negative regulation of dopamine secretion(GO:0033602) |
| 1.6 | 4.7 | GO:0044337 | canonical Wnt signaling pathway involved in positive regulation of apoptotic process(GO:0044337) |
| 1.5 | 4.5 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 1.5 | 4.5 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 1.4 | 7.1 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 1.4 | 7.1 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 1.4 | 11.1 | GO:0045217 | cell-cell junction maintenance(GO:0045217) |
| 1.4 | 1.4 | GO:0097114 | NMDA glutamate receptor clustering(GO:0097114) |
| 1.4 | 2.7 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 1.3 | 5.3 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 1.3 | 3.9 | GO:1901843 | positive regulation of high voltage-gated calcium channel activity(GO:1901843) |
| 1.3 | 9.1 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 1.3 | 7.8 | GO:0048852 | diencephalon morphogenesis(GO:0048852) |
| 1.3 | 10.4 | GO:0071625 | vocalization behavior(GO:0071625) |
| 1.3 | 3.9 | GO:1902071 | regulation of hypoxia-inducible factor-1alpha signaling pathway(GO:1902071) |
| 1.3 | 7.7 | GO:0097090 | presynaptic membrane organization(GO:0097090) |
| 1.2 | 14.6 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 1.2 | 3.6 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 1.2 | 3.6 | GO:0046959 | habituation(GO:0046959) |
| 1.2 | 20.4 | GO:0001504 | neurotransmitter uptake(GO:0001504) |
| 1.2 | 2.4 | GO:0097113 | AMPA glutamate receptor clustering(GO:0097113) glutamate receptor clustering(GO:0097688) |
| 1.2 | 6.0 | GO:0097151 | positive regulation of inhibitory postsynaptic potential(GO:0097151) |
| 1.2 | 2.3 | GO:0021773 | striatal medium spiny neuron differentiation(GO:0021773) |
| 1.1 | 4.5 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 1.1 | 3.4 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 1.1 | 9.0 | GO:0021830 | interneuron migration from the subpallium to the cortex(GO:0021830) |
| 1.1 | 3.2 | GO:1903223 | positive regulation of oxidative stress-induced neuron death(GO:1903223) |
| 1.1 | 3.2 | GO:0060060 | post-embryonic retina morphogenesis in camera-type eye(GO:0060060) |
| 1.1 | 7.5 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 1.1 | 3.2 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 1.0 | 1.0 | GO:0061642 | chemoattraction of axon(GO:0061642) |
| 1.0 | 4.1 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 1.0 | 13.2 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 1.0 | 3.0 | GO:0048677 | axon extension involved in regeneration(GO:0048677) sprouting of injured axon(GO:0048682) |
| 1.0 | 16.9 | GO:0071475 | cellular hyperosmotic salinity response(GO:0071475) |
| 1.0 | 4.9 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 1.0 | 1.9 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 0.9 | 1.9 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.9 | 1.9 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.9 | 1.9 | GO:0071205 | protein localization to juxtaparanode region of axon(GO:0071205) |
| 0.9 | 1.8 | GO:0048696 | regulation of collateral sprouting in absence of injury(GO:0048696) |
| 0.9 | 4.5 | GO:0034047 | regulation of protein phosphatase type 2A activity(GO:0034047) |
| 0.9 | 2.7 | GO:0033693 | neurofilament bundle assembly(GO:0033693) |
| 0.9 | 0.9 | GO:0021586 | pons maturation(GO:0021586) |
| 0.9 | 2.7 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.8 | 43.8 | GO:0021954 | central nervous system neuron development(GO:0021954) |
| 0.8 | 0.8 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.8 | 2.4 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.8 | 4.6 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.8 | 2.3 | GO:1904395 | positive regulation of postsynaptic membrane organization(GO:1901628) regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904393) positive regulation of skeletal muscle acetylcholine-gated channel clustering(GO:1904395) |
| 0.8 | 6.8 | GO:0034643 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.7 | 3.7 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.7 | 0.7 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.7 | 2.2 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.7 | 2.9 | GO:0098597 | observational learning(GO:0098597) |
| 0.7 | 4.3 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.7 | 2.8 | GO:2001016 | positive regulation of skeletal muscle cell differentiation(GO:2001016) |
| 0.7 | 1.4 | GO:0072069 | distal convoluted tubule development(GO:0072025) DCT cell differentiation(GO:0072069) metanephric distal convoluted tubule development(GO:0072221) metanephric distal tubule development(GO:0072235) metanephric DCT cell differentiation(GO:0072240) |
| 0.7 | 1.3 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 0.7 | 3.3 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.7 | 2.0 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.6 | 1.3 | GO:0061346 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) planar cell polarity pathway involved in heart morphogenesis(GO:0061346) |
| 0.6 | 2.5 | GO:1902004 | positive regulation of beta-amyloid formation(GO:1902004) positive regulation of amyloid precursor protein catabolic process(GO:1902993) |
| 0.6 | 5.1 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.6 | 8.6 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.6 | 9.8 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.6 | 1.2 | GO:2000598 | regulation of cyclin catabolic process(GO:2000598) negative regulation of cyclin catabolic process(GO:2000599) |
| 0.6 | 1.2 | GO:0001661 | conditioned taste aversion(GO:0001661) |
| 0.6 | 1.8 | GO:0032687 | negative regulation of interferon-alpha production(GO:0032687) |
| 0.6 | 1.7 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.6 | 1.1 | GO:0021912 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) |
| 0.6 | 4.4 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.5 | 1.6 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.5 | 1.1 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.5 | 3.2 | GO:0021942 | radial glia guided migration of Purkinje cell(GO:0021942) |
| 0.5 | 1.5 | GO:0070948 | regulation of neutrophil mediated cytotoxicity(GO:0070948) regulation of neutrophil mediated killing of symbiont cell(GO:0070949) |
| 0.5 | 2.0 | GO:0042271 | susceptibility to natural killer cell mediated cytotoxicity(GO:0042271) |
| 0.5 | 0.9 | GO:0051665 | membrane raft localization(GO:0051665) |
| 0.5 | 1.4 | GO:0010046 | response to mycotoxin(GO:0010046) |
| 0.5 | 2.3 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.5 | 4.5 | GO:0033523 | histone H2B ubiquitination(GO:0033523) |
| 0.4 | 2.2 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.4 | 1.3 | GO:0072194 | kidney smooth muscle tissue development(GO:0072194) |
| 0.4 | 1.7 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.4 | 3.9 | GO:0007413 | axonal fasciculation(GO:0007413) |
| 0.4 | 5.1 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.4 | 1.7 | GO:0014719 | skeletal muscle satellite cell activation(GO:0014719) |
| 0.4 | 1.3 | GO:0046881 | positive regulation of follicle-stimulating hormone secretion(GO:0046881) |
| 0.4 | 0.4 | GO:0097476 | motor neuron migration(GO:0097475) spinal cord motor neuron migration(GO:0097476) |
| 0.4 | 5.0 | GO:0007616 | long-term memory(GO:0007616) |
| 0.4 | 1.7 | GO:1902261 | positive regulation of delayed rectifier potassium channel activity(GO:1902261) positive regulation of voltage-gated potassium channel activity(GO:1903818) |
| 0.4 | 0.4 | GO:1903261 | regulation of serine phosphorylation of STAT3 protein(GO:1903261) |
| 0.4 | 1.2 | GO:0006420 | arginyl-tRNA aminoacylation(GO:0006420) |
| 0.4 | 2.5 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.4 | 2.5 | GO:2001054 | negative regulation of mesenchymal cell apoptotic process(GO:2001054) |
| 0.4 | 2.1 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 0.4 | 1.2 | GO:0001983 | baroreceptor response to increased systemic arterial blood pressure(GO:0001983) |
| 0.4 | 4.1 | GO:0032291 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.4 | 1.6 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 0.4 | 1.6 | GO:1904058 | positive regulation of sensory perception of pain(GO:1904058) |
| 0.4 | 0.4 | GO:0021550 | medulla oblongata development(GO:0021550) |
| 0.4 | 1.6 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.4 | 1.2 | GO:0035609 | C-terminal protein deglutamylation(GO:0035609) |
| 0.4 | 0.8 | GO:0042939 | glutathione transport(GO:0034635) tripeptide transport(GO:0042939) |
| 0.4 | 3.9 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.4 | 5.4 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.4 | 1.2 | GO:0072017 | distal tubule development(GO:0072017) |
| 0.4 | 1.2 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.4 | 2.7 | GO:0060080 | inhibitory postsynaptic potential(GO:0060080) |
| 0.4 | 0.8 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.4 | 2.6 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.4 | 2.6 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.4 | 0.4 | GO:1990791 | dorsal root ganglion development(GO:1990791) |
| 0.4 | 0.7 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.4 | 1.1 | GO:1904252 | negative regulation of bile acid biosynthetic process(GO:0070858) negative regulation of bile acid metabolic process(GO:1904252) |
| 0.4 | 1.5 | GO:0055059 | asymmetric neuroblast division(GO:0055059) |
| 0.4 | 1.5 | GO:0035247 | peptidyl-arginine omega-N-methylation(GO:0035247) |
| 0.4 | 2.5 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.4 | 1.1 | GO:0002121 | inter-male aggressive behavior(GO:0002121) |
| 0.4 | 1.4 | GO:0046878 | positive regulation of saliva secretion(GO:0046878) |
| 0.4 | 1.1 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.4 | 1.1 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.3 | 2.0 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.3 | 1.7 | GO:0048681 | negative regulation of axon regeneration(GO:0048681) |
| 0.3 | 3.0 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.3 | 1.0 | GO:0042891 | antibiotic transport(GO:0042891) |
| 0.3 | 0.7 | GO:0032497 | detection of lipopolysaccharide(GO:0032497) |
| 0.3 | 1.3 | GO:0051454 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.3 | 0.7 | GO:2000474 | regulation of opioid receptor signaling pathway(GO:2000474) |
| 0.3 | 2.0 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.3 | 0.7 | GO:0060166 | olfactory pit development(GO:0060166) |
| 0.3 | 1.0 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.3 | 1.3 | GO:0030091 | protein repair(GO:0030091) |
| 0.3 | 1.6 | GO:0061590 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.3 | 1.0 | GO:0033122 | negative regulation of purine nucleotide catabolic process(GO:0033122) |
| 0.3 | 1.3 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.3 | 0.6 | GO:0030917 | midbrain-hindbrain boundary development(GO:0030917) |
| 0.3 | 0.6 | GO:2000173 | negative regulation of branching morphogenesis of a nerve(GO:2000173) |
| 0.3 | 1.2 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.3 | 3.1 | GO:0035640 | exploration behavior(GO:0035640) |
| 0.3 | 2.4 | GO:0072501 | cellular divalent inorganic anion homeostasis(GO:0072501) |
| 0.3 | 1.8 | GO:0021604 | cranial nerve structural organization(GO:0021604) |
| 0.3 | 1.5 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.3 | 0.9 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.3 | 0.3 | GO:0042668 | auditory receptor cell fate determination(GO:0042668) |
| 0.3 | 0.9 | GO:0021892 | cerebral cortex GABAergic interneuron differentiation(GO:0021892) |
| 0.3 | 1.2 | GO:0072697 | protein localization to cell cortex(GO:0072697) |
| 0.3 | 21.3 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.3 | 0.6 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) |
| 0.3 | 0.8 | GO:0070649 | formin-nucleated actin cable assembly(GO:0070649) |
| 0.3 | 0.6 | GO:0097252 | oligodendrocyte apoptotic process(GO:0097252) |
| 0.3 | 12.8 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.3 | 1.4 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.3 | 22.1 | GO:0007612 | learning(GO:0007612) |
| 0.3 | 5.7 | GO:0090659 | adult walking behavior(GO:0007628) walking behavior(GO:0090659) |
| 0.3 | 1.4 | GO:1903715 | regulation of aerobic respiration(GO:1903715) |
| 0.3 | 4.3 | GO:1902668 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.3 | 1.1 | GO:0002669 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.3 | 0.3 | GO:0035905 | ascending aorta development(GO:0035905) ascending aorta morphogenesis(GO:0035910) |
| 0.3 | 3.7 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) |
| 0.3 | 0.3 | GO:1902809 | regulation of skeletal muscle fiber differentiation(GO:1902809) |
| 0.3 | 1.8 | GO:0033327 | Leydig cell differentiation(GO:0033327) |
| 0.3 | 1.3 | GO:0034638 | phosphatidylcholine catabolic process(GO:0034638) |
| 0.3 | 0.5 | GO:2001170 | negative regulation of ATP biosynthetic process(GO:2001170) |
| 0.3 | 1.0 | GO:0051031 | tRNA transport(GO:0051031) |
| 0.3 | 0.5 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.2 | 1.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.2 | 1.2 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.2 | 0.5 | GO:0002087 | regulation of respiratory gaseous exchange by neurological system process(GO:0002087) |
| 0.2 | 0.7 | GO:0001975 | response to amphetamine(GO:0001975) |
| 0.2 | 3.7 | GO:0008210 | estrogen metabolic process(GO:0008210) |
| 0.2 | 0.5 | GO:0035935 | androgen secretion(GO:0035935) regulation of androgen secretion(GO:2000834) positive regulation of androgen secretion(GO:2000836) |
| 0.2 | 1.2 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.2 | 1.0 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.2 | 1.7 | GO:0006268 | DNA unwinding involved in DNA replication(GO:0006268) |
| 0.2 | 0.7 | GO:1903587 | regulation of blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:1903587) |
| 0.2 | 0.2 | GO:0072048 | pattern specification involved in kidney development(GO:0061004) renal system pattern specification(GO:0072048) nephrogenic mesenchyme development(GO:0072076) |
| 0.2 | 1.6 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.2 | 0.7 | GO:2000402 | negative regulation of lymphocyte migration(GO:2000402) |
| 0.2 | 1.4 | GO:0001964 | startle response(GO:0001964) |
| 0.2 | 1.4 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.2 | 0.7 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.2 | 0.2 | GO:1902263 | apoptotic process involved in embryonic digit morphogenesis(GO:1902263) |
| 0.2 | 0.4 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.2 | 3.1 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.2 | 2.2 | GO:0060576 | intestinal epithelial cell development(GO:0060576) |
| 0.2 | 0.7 | GO:0097503 | sialylation(GO:0097503) |
| 0.2 | 1.3 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.2 | 0.6 | GO:0042126 | nitrate metabolic process(GO:0042126) |
| 0.2 | 0.9 | GO:0010528 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.2 | 0.6 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.2 | 0.6 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.2 | 0.6 | GO:1902953 | positive regulation of ER to Golgi vesicle-mediated transport(GO:1902953) |
| 0.2 | 1.5 | GO:0035562 | negative regulation of chromatin binding(GO:0035562) |
| 0.2 | 0.6 | GO:0010513 | positive regulation of phosphatidylinositol biosynthetic process(GO:0010513) |
| 0.2 | 0.2 | GO:0019676 | ammonia assimilation cycle(GO:0019676) |
| 0.2 | 5.2 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.2 | 2.1 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.2 | 0.2 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.2 | 0.2 | GO:0021847 | ventricular zone neuroblast division(GO:0021847) |
| 0.2 | 9.4 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.2 | 0.8 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.2 | 1.0 | GO:0090611 | ubiquitin-independent protein catabolic process via the multivesicular body sorting pathway(GO:0090611) |
| 0.2 | 0.4 | GO:0061304 | retinal blood vessel morphogenesis(GO:0061304) |
| 0.2 | 0.4 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.2 | 1.0 | GO:0045607 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.2 | 0.4 | GO:0019614 | catechol-containing compound catabolic process(GO:0019614) catecholamine catabolic process(GO:0042424) |
| 0.2 | 1.2 | GO:0040016 | embryonic cleavage(GO:0040016) |
| 0.2 | 2.2 | GO:0036065 | fucosylation(GO:0036065) |
| 0.2 | 0.8 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.2 | 7.5 | GO:0007270 | neuron-neuron synaptic transmission(GO:0007270) |
| 0.2 | 0.8 | GO:0008228 | opsonization(GO:0008228) |
| 0.2 | 0.8 | GO:0098908 | regulation of neuronal action potential(GO:0098908) |
| 0.2 | 0.2 | GO:0072179 | nephric duct formation(GO:0072179) |
| 0.2 | 1.1 | GO:0031987 | locomotion involved in locomotory behavior(GO:0031987) |
| 0.2 | 2.3 | GO:0002089 | lens morphogenesis in camera-type eye(GO:0002089) |
| 0.2 | 0.4 | GO:0021877 | forebrain neuron fate commitment(GO:0021877) |
| 0.2 | 0.2 | GO:0045176 | apical protein localization(GO:0045176) |
| 0.2 | 0.4 | GO:0045875 | negative regulation of sister chromatid cohesion(GO:0045875) |
| 0.2 | 0.6 | GO:0014028 | notochord formation(GO:0014028) |
| 0.2 | 1.5 | GO:0045075 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.2 | 0.4 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.2 | 3.7 | GO:0030901 | midbrain development(GO:0030901) |
| 0.2 | 0.5 | GO:0072695 | negative regulation of DNA recombination at telomere(GO:0048239) regulation of DNA recombination at telomere(GO:0072695) |
| 0.2 | 0.4 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.2 | 4.1 | GO:0090004 | positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.2 | 0.7 | GO:0070813 | hydrogen sulfide metabolic process(GO:0070813) |
| 0.2 | 7.2 | GO:0050905 | neuromuscular process(GO:0050905) |
| 0.2 | 0.2 | GO:1900368 | regulation of RNA interference(GO:1900368) |
| 0.2 | 0.7 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.2 | 0.5 | GO:1902774 | late endosome to lysosome transport(GO:1902774) |
| 0.2 | 1.2 | GO:0051231 | spindle elongation(GO:0051231) |
| 0.2 | 0.2 | GO:0097154 | GABAergic neuron differentiation(GO:0097154) |
| 0.2 | 0.3 | GO:0048865 | stem cell fate commitment(GO:0048865) |
| 0.2 | 0.8 | GO:0042428 | serotonin metabolic process(GO:0042428) |
| 0.2 | 0.5 | GO:0021794 | thalamus development(GO:0021794) |
| 0.2 | 0.5 | GO:0032066 | nucleolus to nucleoplasm transport(GO:0032066) |
| 0.2 | 0.3 | GO:0071608 | macrophage inflammatory protein-1 alpha production(GO:0071608) |
| 0.2 | 0.6 | GO:0070120 | ciliary neurotrophic factor-mediated signaling pathway(GO:0070120) |
| 0.2 | 0.5 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.2 | 0.6 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.2 | 0.5 | GO:0060689 | cell differentiation involved in salivary gland development(GO:0060689) |
| 0.2 | 0.5 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.2 | 0.5 | GO:0090038 | negative regulation of protein kinase C signaling(GO:0090038) |
| 0.2 | 0.6 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.2 | 0.5 | GO:0072385 | minus-end-directed organelle transport along microtubule(GO:0072385) |
| 0.2 | 0.3 | GO:0071787 | endoplasmic reticulum tubular network assembly(GO:0071787) |
| 0.2 | 3.1 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.2 | 0.5 | GO:0060836 | lymphatic endothelial cell differentiation(GO:0060836) |
| 0.2 | 0.5 | GO:0050655 | dermatan sulfate metabolic process(GO:0030205) dermatan sulfate proteoglycan metabolic process(GO:0050655) |
| 0.2 | 1.4 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.2 | 0.9 | GO:0061003 | positive regulation of dendritic spine morphogenesis(GO:0061003) |
| 0.2 | 2.6 | GO:0008053 | mitochondrial fusion(GO:0008053) |
| 0.2 | 0.2 | GO:0097051 | establishment of protein localization to endoplasmic reticulum membrane(GO:0097051) |
| 0.2 | 0.3 | GO:0046726 | positive regulation by virus of viral protein levels in host cell(GO:0046726) |
| 0.2 | 0.9 | GO:0010457 | centriole-centriole cohesion(GO:0010457) |
| 0.2 | 1.2 | GO:0060384 | innervation(GO:0060384) |
| 0.1 | 0.3 | GO:0071280 | cellular response to copper ion(GO:0071280) |
| 0.1 | 0.3 | GO:0034729 | histone H3-K79 methylation(GO:0034729) |
| 0.1 | 0.6 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.1 | 0.4 | GO:0043328 | late endosome to vacuole transport via multivesicular body sorting pathway(GO:0032511) protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.1 | 0.9 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.1 | 0.9 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.1 | 0.6 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) |
| 0.1 | 0.4 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 0.1 | 4.7 | GO:0050806 | positive regulation of synaptic transmission(GO:0050806) |
| 0.1 | 0.3 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.1 | 0.9 | GO:0008063 | Toll signaling pathway(GO:0008063) |
| 0.1 | 1.2 | GO:0033262 | regulation of nuclear cell cycle DNA replication(GO:0033262) |
| 0.1 | 0.1 | GO:0036257 | multivesicular body organization(GO:0036257) |
| 0.1 | 0.6 | GO:0097039 | protein linear polyubiquitination(GO:0097039) |
| 0.1 | 0.4 | GO:1990414 | replication-born double-strand break repair via sister chromatid exchange(GO:1990414) |
| 0.1 | 1.3 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.1 | 0.6 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.1 | 0.1 | GO:0014012 | peripheral nervous system axon regeneration(GO:0014012) |
| 0.1 | 1.6 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.1 | 1.1 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 0.3 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.1 | 0.4 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.1 | 0.1 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.1 | 0.4 | GO:0060178 | regulation of exocyst localization(GO:0060178) |
| 0.1 | 0.4 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.1 | 0.1 | GO:1903935 | response to sodium arsenite(GO:1903935) |
| 0.1 | 1.3 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 0.1 | GO:0021692 | cerebellar Purkinje cell layer morphogenesis(GO:0021692) |
| 0.1 | 0.8 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.1 | 0.5 | GO:0070671 | response to interleukin-12(GO:0070671) |
| 0.1 | 0.5 | GO:0032747 | positive regulation of interleukin-23 production(GO:0032747) |
| 0.1 | 0.4 | GO:0072429 | response to intra-S DNA damage checkpoint signaling(GO:0072429) |
| 0.1 | 0.3 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.1 | 2.6 | GO:0042462 | eye photoreceptor cell development(GO:0042462) |
| 0.1 | 0.4 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.1 | 0.5 | GO:0038031 | non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.1 | 0.4 | GO:2000325 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.1 | 0.4 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.1 | 0.7 | GO:0031620 | regulation of fever generation(GO:0031620) |
| 0.1 | 2.1 | GO:0050650 | chondroitin sulfate proteoglycan biosynthetic process(GO:0050650) |
| 0.1 | 0.2 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) |
| 0.1 | 2.9 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 0.2 | GO:0090135 | actin filament branching(GO:0090135) |
| 0.1 | 1.1 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.1 | 0.4 | GO:0006553 | lysine metabolic process(GO:0006553) |
| 0.1 | 0.5 | GO:1904707 | positive regulation of vascular smooth muscle cell proliferation(GO:1904707) |
| 0.1 | 0.1 | GO:0060535 | trachea cartilage morphogenesis(GO:0060535) |
| 0.1 | 0.4 | GO:0043653 | mitochondrial fragmentation involved in apoptotic process(GO:0043653) |
| 0.1 | 0.5 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.1 | 0.2 | GO:0003419 | growth plate cartilage chondrocyte proliferation(GO:0003419) |
| 0.1 | 0.9 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 7.7 | GO:0050770 | regulation of axonogenesis(GO:0050770) |
| 0.1 | 0.2 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.1 | 0.1 | GO:0071316 | cellular response to nicotine(GO:0071316) |
| 0.1 | 0.2 | GO:0007619 | courtship behavior(GO:0007619) |
| 0.1 | 1.2 | GO:0021766 | hippocampus development(GO:0021766) |
| 0.1 | 0.4 | GO:0015824 | proline transport(GO:0015824) |
| 0.1 | 1.7 | GO:0032098 | regulation of appetite(GO:0032098) |
| 0.1 | 0.1 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.1 | 0.5 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.1 | 0.3 | GO:0044828 | negative regulation by host of viral genome replication(GO:0044828) |
| 0.1 | 0.7 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.1 | 0.6 | GO:0071242 | cellular response to ammonium ion(GO:0071242) |
| 0.1 | 0.2 | GO:2001023 | regulation of response to drug(GO:2001023) |
| 0.1 | 0.2 | GO:0006990 | positive regulation of transcription from RNA polymerase II promoter involved in unfolded protein response(GO:0006990) |
| 0.1 | 0.3 | GO:0098904 | regulation of AV node cell action potential(GO:0098904) |
| 0.1 | 0.4 | GO:0048007 | antigen processing and presentation of lipid antigen via MHC class Ib(GO:0048003) antigen processing and presentation, exogenous lipid antigen via MHC class Ib(GO:0048007) |
| 0.1 | 0.4 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.1 | 0.3 | GO:1901841 | regulation of high voltage-gated calcium channel activity(GO:1901841) |
| 0.1 | 0.3 | GO:0035928 | RNA import into mitochondrion(GO:0035927) rRNA import into mitochondrion(GO:0035928) |
| 0.1 | 0.3 | GO:1903998 | regulation of eating behavior(GO:1903998) |
| 0.1 | 0.3 | GO:1903847 | regulation of aorta morphogenesis(GO:1903847) positive regulation of aorta morphogenesis(GO:1903849) |
| 0.1 | 0.2 | GO:0007213 | G-protein coupled acetylcholine receptor signaling pathway(GO:0007213) |
| 0.1 | 0.5 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.1 | 0.2 | GO:2000523 | regulation of T cell costimulation(GO:2000523) positive regulation of T cell costimulation(GO:2000525) |
| 0.1 | 0.7 | GO:0006012 | galactose metabolic process(GO:0006012) |
| 0.1 | 0.5 | GO:0099515 | actin filament-based transport(GO:0099515) |
| 0.1 | 0.8 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.1 | 0.4 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.1 | 0.4 | GO:0035280 | miRNA loading onto RISC involved in gene silencing by miRNA(GO:0035280) |
| 0.1 | 0.1 | GO:0051464 | positive regulation of cortisol secretion(GO:0051464) |
| 0.1 | 0.3 | GO:1900045 | negative regulation of protein K63-linked ubiquitination(GO:1900045) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.1 | 0.5 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 0.3 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.1 | 0.6 | GO:2000369 | regulation of clathrin-mediated endocytosis(GO:2000369) |
| 0.1 | 0.2 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.1 | 0.3 | GO:0046439 | cysteine catabolic process(GO:0009093) L-cysteine catabolic process(GO:0019448) L-cysteine metabolic process(GO:0046439) |
| 0.1 | 0.2 | GO:0060509 | Type I pneumocyte differentiation(GO:0060509) |
| 0.1 | 0.7 | GO:0015813 | L-glutamate transport(GO:0015813) |
| 0.1 | 0.5 | GO:0071732 | cellular response to nitric oxide(GO:0071732) |
| 0.1 | 0.4 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.1 | 0.8 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 0.3 | GO:0015740 | C4-dicarboxylate transport(GO:0015740) |
| 0.1 | 0.1 | GO:0061428 | negative regulation of transcription from RNA polymerase II promoter in response to hypoxia(GO:0061428) |
| 0.1 | 0.4 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.1 | 0.6 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.1 | 0.8 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 0.3 | GO:1903358 | regulation of Golgi organization(GO:1903358) |
| 0.1 | 0.2 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.1 | 0.1 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.1 | 0.2 | GO:0006624 | vacuolar protein processing(GO:0006624) |
| 0.1 | 0.3 | GO:1902570 | protein localization to nucleolus(GO:1902570) |
| 0.1 | 0.6 | GO:0071044 | histone mRNA catabolic process(GO:0071044) |
| 0.1 | 1.8 | GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) |
| 0.1 | 0.1 | GO:0048242 | epinephrine secretion(GO:0048242) |
| 0.1 | 0.3 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 0.5 | GO:2000060 | positive regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000060) |
| 0.1 | 0.2 | GO:0035964 | COPI-coated vesicle budding(GO:0035964) |
| 0.1 | 0.2 | GO:0042940 | D-amino acid transport(GO:0042940) |
| 0.1 | 0.1 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.1 | 0.2 | GO:0006344 | maintenance of chromatin silencing(GO:0006344) |
| 0.1 | 0.1 | GO:1904706 | negative regulation of vascular smooth muscle cell proliferation(GO:1904706) |
| 0.1 | 0.4 | GO:0071440 | regulation of histone H3-K14 acetylation(GO:0071440) |
| 0.1 | 0.2 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 0.1 | GO:0002371 | dendritic cell cytokine production(GO:0002371) |
| 0.1 | 0.1 | GO:0010360 | negative regulation of anion channel activity(GO:0010360) |
| 0.1 | 0.1 | GO:0098915 | membrane repolarization during ventricular cardiac muscle cell action potential(GO:0098915) |
| 0.1 | 0.4 | GO:0015816 | glycine transport(GO:0015816) |
| 0.1 | 0.3 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 0.6 | GO:0006878 | cellular copper ion homeostasis(GO:0006878) |
| 0.1 | 0.7 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.1 | 0.4 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) |
| 0.1 | 0.5 | GO:0006563 | L-serine metabolic process(GO:0006563) |
| 0.1 | 0.1 | GO:0060916 | mesenchymal cell proliferation involved in lung development(GO:0060916) |
| 0.1 | 0.1 | GO:1903279 | regulation of calcium:sodium antiporter activity(GO:1903279) |
| 0.1 | 0.4 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 0.2 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.1 | GO:0045085 | negative regulation of interleukin-2 biosynthetic process(GO:0045085) |
| 0.1 | 0.5 | GO:1900102 | negative regulation of endoplasmic reticulum unfolded protein response(GO:1900102) |
| 0.1 | 0.6 | GO:0006337 | nucleosome disassembly(GO:0006337) protein-DNA complex disassembly(GO:0032986) |
| 0.1 | 0.1 | GO:0032741 | positive regulation of interleukin-18 production(GO:0032741) |
| 0.1 | 0.1 | GO:0051660 | establishment of centrosome localization(GO:0051660) |
| 0.1 | 0.4 | GO:0048227 | plasma membrane to endosome transport(GO:0048227) |
| 0.1 | 0.2 | GO:0060579 | ventral spinal cord interneuron fate commitment(GO:0060579) cell fate commitment involved in pattern specification(GO:0060581) |
| 0.1 | 0.3 | GO:0070345 | negative regulation of fat cell proliferation(GO:0070345) |
| 0.1 | 0.3 | GO:1901341 | positive regulation of store-operated calcium channel activity(GO:1901341) |
| 0.1 | 1.0 | GO:0044705 | mating behavior(GO:0007617) multi-organism reproductive behavior(GO:0044705) |
| 0.1 | 0.1 | GO:1900377 | negative regulation of melanin biosynthetic process(GO:0048022) negative regulation of secondary metabolite biosynthetic process(GO:1900377) |
| 0.1 | 0.2 | GO:1902969 | mitotic DNA replication(GO:1902969) |
| 0.1 | 0.1 | GO:0051964 | negative regulation of synapse assembly(GO:0051964) |
| 0.1 | 0.2 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.1 | 0.1 | GO:0090230 | regulation of centromere complex assembly(GO:0090230) |
| 0.1 | 0.1 | GO:2000017 | positive regulation of determination of dorsal identity(GO:2000017) |
| 0.1 | 2.8 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.1 | 0.2 | GO:0042732 | D-xylose metabolic process(GO:0042732) |
| 0.1 | 0.2 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.1 | 0.1 | GO:0072201 | negative regulation of mesenchymal cell proliferation(GO:0072201) |
| 0.1 | 0.3 | GO:0015919 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.1 | 0.1 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.1 | 0.2 | GO:0060452 | positive regulation of cardiac muscle contraction(GO:0060452) |
| 0.1 | 0.4 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.1 | 0.1 | GO:0009996 | negative regulation of cell fate specification(GO:0009996) |
| 0.1 | 0.2 | GO:0006172 | ADP biosynthetic process(GO:0006172) |
| 0.1 | 0.7 | GO:0045948 | positive regulation of translational initiation(GO:0045948) |
| 0.1 | 0.2 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.1 | 0.8 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 0.1 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.1 | 0.1 | GO:0044336 | canonical Wnt signaling pathway involved in negative regulation of apoptotic process(GO:0044336) |
| 0.1 | 0.2 | GO:0043097 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine nucleoside salvage(GO:0043097) |
| 0.1 | 0.1 | GO:0010871 | negative regulation of receptor biosynthetic process(GO:0010871) |
| 0.1 | 0.1 | GO:0014873 | response to muscle activity involved in regulation of muscle adaptation(GO:0014873) |
| 0.1 | 0.4 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.1 | 0.1 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) |
| 0.1 | 0.4 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.1 | 0.3 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 0.1 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) |
| 0.1 | 0.2 | GO:0003415 | chondrocyte hypertrophy(GO:0003415) |
| 0.1 | 0.2 | GO:0060023 | soft palate development(GO:0060023) |
| 0.1 | 0.5 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 0.4 | GO:0007202 | activation of phospholipase C activity(GO:0007202) |
| 0.1 | 0.3 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.1 | 0.1 | GO:0061101 | neuroendocrine cell differentiation(GO:0061101) |
| 0.1 | 0.6 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.1 | 0.2 | GO:0001827 | inner cell mass cell fate commitment(GO:0001827) |
| 0.1 | 0.1 | GO:1902455 | negative regulation of stem cell population maintenance(GO:1902455) |
| 0.1 | 0.1 | GO:0008105 | asymmetric protein localization(GO:0008105) |
| 0.1 | 0.1 | GO:1904177 | regulation of adipose tissue development(GO:1904177) |
| 0.1 | 0.1 | GO:1900157 | regulation of bone mineralization involved in bone maturation(GO:1900157) |
| 0.1 | 0.1 | GO:0016557 | peroxisome membrane biogenesis(GO:0016557) |
| 0.0 | 0.1 | GO:0070601 | centromeric sister chromatid cohesion(GO:0070601) |
| 0.0 | 0.1 | GO:1902512 | positive regulation of apoptotic DNA fragmentation(GO:1902512) positive regulation of DNA catabolic process(GO:1903626) |
| 0.0 | 0.1 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.0 | 0.1 | GO:0045080 | positive regulation of chemokine biosynthetic process(GO:0045080) |
| 0.0 | 0.1 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.0 | 0.1 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.0 | 0.0 | GO:0050713 | negative regulation of interleukin-1 beta secretion(GO:0050713) |
| 0.0 | 0.3 | GO:0006264 | mitochondrial DNA replication(GO:0006264) |
| 0.0 | 0.1 | GO:0044065 | regulation of respiratory system process(GO:0044065) |
| 0.0 | 0.2 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.0 | 0.1 | GO:0009629 | response to gravity(GO:0009629) |
| 0.0 | 0.1 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.0 | 0.2 | GO:0050917 | sensory perception of umami taste(GO:0050917) |
| 0.0 | 0.6 | GO:0051205 | protein insertion into membrane(GO:0051205) |
| 0.0 | 0.0 | GO:0090403 | oxidative stress-induced premature senescence(GO:0090403) |
| 0.0 | 0.1 | GO:0046104 | thymidine metabolic process(GO:0046104) |
| 0.0 | 0.0 | GO:0070427 | nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070427) |
| 0.0 | 0.2 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.0 | 0.6 | GO:0043982 | histone H4-K5 acetylation(GO:0043981) histone H4-K8 acetylation(GO:0043982) |
| 0.0 | 0.1 | GO:0006235 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) dTTP metabolic process(GO:0046075) |
| 0.0 | 0.2 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.0 | 0.1 | GO:0002756 | MyD88-independent toll-like receptor signaling pathway(GO:0002756) |
| 0.0 | 0.3 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.0 | 0.2 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 0.0 | 0.0 | GO:0021938 | smoothened signaling pathway involved in regulation of cerebellar granule cell precursor cell proliferation(GO:0021938) |
| 0.0 | 0.2 | GO:0042737 | drug catabolic process(GO:0042737) |
| 0.0 | 0.2 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 0.0 | 0.1 | GO:0002692 | negative regulation of cellular extravasation(GO:0002692) |
| 0.0 | 0.0 | GO:0002461 | tolerance induction dependent upon immune response(GO:0002461) |
| 0.0 | 0.4 | GO:0007342 | fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.0 | 0.0 | GO:0030908 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.0 | 0.5 | GO:0048714 | positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.0 | 0.0 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 0.0 | 0.1 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.0 | 0.0 | GO:0031498 | chromatin disassembly(GO:0031498) |
| 0.0 | 0.1 | GO:2000553 | positive regulation of T-helper 2 cell cytokine production(GO:2000553) |
| 0.0 | 0.1 | GO:0072718 | response to cisplatin(GO:0072718) |
| 0.0 | 0.1 | GO:0010825 | positive regulation of centrosome duplication(GO:0010825) |
| 0.0 | 0.1 | GO:0070966 | nuclear-transcribed mRNA catabolic process, no-go decay(GO:0070966) |
| 0.0 | 0.2 | GO:2000348 | regulation of CD40 signaling pathway(GO:2000348) |
| 0.0 | 0.2 | GO:0002591 | positive regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002591) |
| 0.0 | 0.2 | GO:0045542 | positive regulation of cholesterol biosynthetic process(GO:0045542) positive regulation of cholesterol metabolic process(GO:0090205) |
| 0.0 | 0.2 | GO:0046606 | negative regulation of centrosome duplication(GO:0010826) negative regulation of centrosome cycle(GO:0046606) |
| 0.0 | 0.1 | GO:0070245 | positive regulation of thymocyte apoptotic process(GO:0070245) |
| 0.0 | 0.3 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.0 | 0.2 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.0 | 0.2 | GO:0006501 | C-terminal protein lipidation(GO:0006501) |
| 0.0 | 0.3 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.0 | 0.3 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.0 | 0.2 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.0 | 0.1 | GO:1902459 | positive regulation of stem cell population maintenance(GO:1902459) |
| 0.0 | 0.1 | GO:0035511 | oxidative DNA demethylation(GO:0035511) |
| 0.0 | 0.1 | GO:2000587 | regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000586) negative regulation of platelet-derived growth factor receptor-beta signaling pathway(GO:2000587) |
| 0.0 | 0.1 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.0 | 0.0 | GO:0061668 | mitochondrial ribosome assembly(GO:0061668) |
| 0.0 | 0.0 | GO:0072603 | interleukin-5 secretion(GO:0072603) interleukin-13 secretion(GO:0072611) regulation of interleukin-5 secretion(GO:2000662) positive regulation of interleukin-5 secretion(GO:2000664) regulation of interleukin-13 secretion(GO:2000665) positive regulation of interleukin-13 secretion(GO:2000667) |
| 0.0 | 0.0 | GO:0002043 | blood vessel endothelial cell proliferation involved in sprouting angiogenesis(GO:0002043) |
| 0.0 | 0.1 | GO:0006551 | leucine metabolic process(GO:0006551) |
| 0.0 | 0.3 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.0 | 0.3 | GO:0060211 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) |
| 0.0 | 0.0 | GO:0021523 | somatic motor neuron differentiation(GO:0021523) |
| 0.0 | 0.0 | GO:0042414 | epinephrine metabolic process(GO:0042414) |
| 0.0 | 0.0 | GO:0003348 | cardiac endothelial cell differentiation(GO:0003348) |
| 0.0 | 0.1 | GO:0032304 | negative regulation of icosanoid secretion(GO:0032304) |
| 0.0 | 0.1 | GO:0051388 | positive regulation of neurotrophin TRK receptor signaling pathway(GO:0051388) |
| 0.0 | 0.1 | GO:1902902 | negative regulation of autophagosome assembly(GO:1902902) |
| 0.0 | 0.0 | GO:0019856 | 'de novo' pyrimidine nucleobase biosynthetic process(GO:0006207) pyrimidine nucleobase biosynthetic process(GO:0019856) |
| 0.0 | 0.1 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) regulation of transforming growth factor-beta secretion(GO:2001201) |
| 0.0 | 3.0 | GO:0007338 | single fertilization(GO:0007338) |
| 0.0 | 0.2 | GO:0046146 | tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.0 | 0.0 | GO:0002434 | immune complex clearance(GO:0002434) |
| 0.0 | 0.0 | GO:0010725 | regulation of primitive erythrocyte differentiation(GO:0010725) |
| 0.0 | 0.3 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 0.1 | GO:0030263 | apoptotic chromosome condensation(GO:0030263) |
| 0.0 | 0.8 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 0.3 | GO:1904355 | positive regulation of telomere capping(GO:1904355) |
| 0.0 | 0.0 | GO:0000715 | nucleotide-excision repair, DNA damage recognition(GO:0000715) |
| 0.0 | 0.0 | GO:2001269 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:2001269) |
| 0.0 | 0.3 | GO:1902751 | positive regulation of cell cycle G2/M phase transition(GO:1902751) |
| 0.0 | 0.0 | GO:1904714 | regulation of chaperone-mediated autophagy(GO:1904714) |
| 0.0 | 0.1 | GO:0060124 | positive regulation of growth hormone secretion(GO:0060124) |
| 0.0 | 0.0 | GO:0090365 | regulation of mRNA modification(GO:0090365) |
| 0.0 | 0.2 | GO:0002759 | regulation of antimicrobial humoral response(GO:0002759) |
| 0.0 | 0.0 | GO:0071651 | positive regulation of chemokine (C-C motif) ligand 5 production(GO:0071651) |
| 0.0 | 0.0 | GO:0060591 | chondroblast differentiation(GO:0060591) |
| 0.0 | 0.6 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.0 | 0.2 | GO:0030049 | muscle filament sliding(GO:0030049) |
| 0.0 | 0.1 | GO:0071257 | cellular response to electrical stimulus(GO:0071257) |
| 0.0 | 0.1 | GO:0051410 | detoxification of nitrogen compound(GO:0051410) cellular detoxification of nitrogen compound(GO:0070458) |
| 0.0 | 0.0 | GO:0032025 | response to cobalt ion(GO:0032025) |
| 0.0 | 0.2 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 0.0 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 0.0 | 0.1 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.0 | 0.0 | GO:0031587 | positive regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0031587) |
| 0.0 | 0.5 | GO:0006221 | pyrimidine nucleotide biosynthetic process(GO:0006221) |
| 0.0 | 0.0 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.0 | 0.1 | GO:0006296 | nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) nucleotide-excision repair, DNA incision, 5'-to lesion(GO:0006296) |
| 0.0 | 0.1 | GO:0051596 | methylglyoxal catabolic process to D-lactate via S-lactoyl-glutathione(GO:0019243) methylglyoxal catabolic process(GO:0051596) methylglyoxal catabolic process to lactate(GO:0061727) |
| 0.0 | 0.0 | GO:0003139 | secondary heart field specification(GO:0003139) |
| 0.0 | 0.1 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 0.0 | 0.0 | GO:2000909 | regulation of cholesterol import(GO:0060620) regulation of sterol import(GO:2000909) |
| 0.0 | 0.1 | GO:2000015 | regulation of determination of dorsal identity(GO:2000015) |
| 0.0 | 0.0 | GO:0014832 | urinary bladder smooth muscle contraction(GO:0014832) |
| 0.0 | 0.1 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.0 | 0.1 | GO:0050955 | thermoception(GO:0050955) |
| 0.0 | 0.1 | GO:0007135 | meiosis II(GO:0007135) |
| 0.0 | 0.2 | GO:0035561 | regulation of chromatin binding(GO:0035561) |
| 0.0 | 0.1 | GO:0090073 | positive regulation of protein homodimerization activity(GO:0090073) |
| 0.0 | 0.2 | GO:0051601 | exocyst localization(GO:0051601) |
| 0.0 | 0.1 | GO:0035428 | hexose transmembrane transport(GO:0035428) |
| 0.0 | 0.3 | GO:0001731 | formation of translation preinitiation complex(GO:0001731) |
| 0.0 | 1.6 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 0.1 | GO:0045792 | negative regulation of cell size(GO:0045792) |
| 0.0 | 0.1 | GO:0033240 | positive regulation of cellular amine metabolic process(GO:0033240) |
| 0.0 | 0.1 | GO:0070375 | ERK5 cascade(GO:0070375) |
| 0.0 | 0.2 | GO:0060856 | establishment of blood-brain barrier(GO:0060856) |
| 0.0 | 0.5 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.0 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.0 | 0.1 | GO:0032020 | ISG15-protein conjugation(GO:0032020) |
| 0.0 | 0.0 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 0.0 | 0.1 | GO:0018344 | protein geranylgeranylation(GO:0018344) |
| 0.0 | 1.2 | GO:0030038 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.0 | 0.0 | GO:0010288 | response to lead ion(GO:0010288) |
| 0.0 | 0.1 | GO:0006562 | proline catabolic process(GO:0006562) |
| 0.0 | 0.1 | GO:0048675 | axon extension(GO:0048675) |
| 0.0 | 0.1 | GO:0046642 | negative regulation of alpha-beta T cell proliferation(GO:0046642) |
| 0.0 | 0.1 | GO:0015705 | iodide transport(GO:0015705) |
| 0.0 | 0.1 | GO:0030730 | sequestering of triglyceride(GO:0030730) |
| 0.0 | 0.1 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.0 | 0.0 | GO:0010182 | carbohydrate mediated signaling(GO:0009756) hexose mediated signaling(GO:0009757) sugar mediated signaling pathway(GO:0010182) glucose mediated signaling pathway(GO:0010255) |
| 0.0 | 0.2 | GO:0042461 | photoreceptor cell development(GO:0042461) |
| 0.0 | 0.1 | GO:0019086 | late viral transcription(GO:0019086) |
| 0.0 | 0.1 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.0 | 0.1 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.0 | 0.1 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 0.0 | 0.1 | GO:1902035 | positive regulation of hematopoietic stem cell proliferation(GO:1902035) |
| 0.0 | 0.2 | GO:0039702 | viral budding via host ESCRT complex(GO:0039702) |
| 0.0 | 1.1 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 0.0 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.0 | 0.0 | GO:1904431 | positive regulation of t-circle formation(GO:1904431) |
| 0.0 | 0.0 | GO:0016561 | protein import into peroxisome matrix, translocation(GO:0016561) |
| 0.0 | 0.0 | GO:0032196 | transposition(GO:0032196) |
| 0.0 | 0.1 | GO:1903966 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.0 | 0.0 | GO:0034331 | cell junction maintenance(GO:0034331) |
| 0.0 | 0.0 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) |
| 0.0 | 0.0 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.0 | 0.1 | GO:0060672 | epithelial cell differentiation involved in embryonic placenta development(GO:0060671) epithelial cell morphogenesis involved in placental branching(GO:0060672) |
| 0.0 | 0.0 | GO:0033615 | mitochondrial proton-transporting ATP synthase complex assembly(GO:0033615) |
| 0.0 | 0.1 | GO:0034653 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
| 0.0 | 0.1 | GO:0060292 | long term synaptic depression(GO:0060292) |
| 0.0 | 0.3 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.1 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.0 | 0.1 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.0 | 0.0 | GO:0018916 | nitrobenzene metabolic process(GO:0018916) |
| 0.0 | 0.1 | GO:0097500 | receptor localization to nonmotile primary cilium(GO:0097500) |
| 0.0 | 0.0 | GO:1901339 | regulation of store-operated calcium channel activity(GO:1901339) |
| 0.0 | 0.0 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.0 | 0.1 | GO:0006361 | transcription initiation from RNA polymerase I promoter(GO:0006361) |
| 0.0 | 0.0 | GO:0097167 | circadian regulation of translation(GO:0097167) |
| 0.0 | 0.1 | GO:0032401 | establishment of melanosome localization(GO:0032401) |
| 0.0 | 0.1 | GO:0045540 | regulation of cholesterol biosynthetic process(GO:0045540) |
| 0.0 | 0.0 | GO:0043988 | histone H3-S28 phosphorylation(GO:0043988) |
| 0.0 | 0.3 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.1 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.0 | 0.3 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.0 | GO:0007418 | ventral midline development(GO:0007418) |
| 0.0 | 0.0 | GO:0051791 | medium-chain fatty acid metabolic process(GO:0051791) |
| 0.0 | 0.0 | GO:0072034 | renal vesicle induction(GO:0072034) |
| 0.0 | 0.1 | GO:0000389 | mRNA 3'-splice site recognition(GO:0000389) |
| 0.0 | 0.1 | GO:1902018 | negative regulation of cilium assembly(GO:1902018) |
| 0.0 | 0.1 | GO:0097035 | regulation of membrane lipid distribution(GO:0097035) |
| 0.0 | 0.0 | GO:0033227 | dsRNA transport(GO:0033227) |
| 0.0 | 0.0 | GO:0071462 | cellular response to water stimulus(GO:0071462) |
| 0.0 | 0.4 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.0 | GO:0018364 | peptidyl-glutamine methylation(GO:0018364) |
| 0.0 | 0.0 | GO:0042699 | follicle-stimulating hormone signaling pathway(GO:0042699) |
| 0.0 | 0.7 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 0.2 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.0 | 0.1 | GO:0050435 | beta-amyloid metabolic process(GO:0050435) |
| 0.0 | 0.0 | GO:0099612 | protein localization to paranode region of axon(GO:0002175) protein localization to axon(GO:0099612) |
| 0.0 | 0.0 | GO:0045026 | plasma membrane fusion(GO:0045026) |
| 0.0 | 0.0 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.0 | 0.1 | GO:0006220 | pyrimidine nucleotide metabolic process(GO:0006220) |
| 0.0 | 0.0 | GO:0051182 | coenzyme transport(GO:0051182) |
| 0.0 | 0.0 | GO:0006498 | N-terminal protein lipidation(GO:0006498) |
| 0.0 | 0.0 | GO:0071873 | response to norepinephrine(GO:0071873) |
| 0.0 | 0.1 | GO:0010820 | positive regulation of T cell chemotaxis(GO:0010820) |
| 0.0 | 0.0 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.0 | 0.0 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.0 | 0.0 | GO:0090360 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.0 | 0.0 | GO:0010624 | regulation of Schwann cell proliferation(GO:0010624) negative regulation of Schwann cell proliferation(GO:0010626) |
| 0.0 | 0.1 | GO:0035330 | regulation of hippo signaling(GO:0035330) |
| 0.0 | 0.0 | GO:0016480 | negative regulation of transcription from RNA polymerase III promoter(GO:0016480) |
| 0.0 | 0.0 | GO:0071895 | odontoblast differentiation(GO:0071895) |
| 0.0 | 0.0 | GO:0007320 | insemination(GO:0007320) |
| 0.0 | 0.0 | GO:0031936 | negative regulation of chromatin silencing(GO:0031936) |
| 0.0 | 0.0 | GO:0021819 | layer formation in cerebral cortex(GO:0021819) |
| 0.0 | 0.1 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.0 | 0.0 | GO:1901889 | negative regulation of cell junction assembly(GO:1901889) |
| 0.0 | 0.1 | GO:1900116 | sequestering of extracellular ligand from receptor(GO:0035581) extracellular regulation of signal transduction(GO:1900115) extracellular negative regulation of signal transduction(GO:1900116) |
| 0.0 | 0.0 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.0 | 0.0 | GO:0045653 | negative regulation of megakaryocyte differentiation(GO:0045653) |
| 0.0 | 0.0 | GO:0015870 | acetylcholine transport(GO:0015870) acetate ester transport(GO:1901374) |
| 0.0 | 0.0 | GO:2000152 | regulation of ubiquitin-specific protease activity(GO:2000152) |
| 0.0 | 0.0 | GO:0003072 | angiotensin mediated vasoconstriction involved in regulation of systemic arterial blood pressure(GO:0001998) regulation of blood vessel size by renin-angiotensin(GO:0002034) renal control of peripheral vascular resistance involved in regulation of systemic arterial blood pressure(GO:0003072) |
| 0.0 | 0.1 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.0 | 0.0 | GO:0010640 | regulation of platelet-derived growth factor receptor signaling pathway(GO:0010640) |
| 0.0 | 0.0 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.0 | 0.0 | GO:0010888 | negative regulation of lipid storage(GO:0010888) |
| 0.0 | 0.0 | GO:0070681 | glutaminyl-tRNAGln biosynthesis via transamidation(GO:0070681) |
| 0.0 | 0.0 | GO:0042135 | neurotransmitter catabolic process(GO:0042135) |
| 0.0 | 0.0 | GO:0006824 | cobalt ion transport(GO:0006824) |
| 0.0 | 0.0 | GO:0002426 | immunoglobulin production in mucosal tissue(GO:0002426) |
| 0.0 | 0.0 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.0 | 0.0 | GO:0033684 | regulation of luteinizing hormone secretion(GO:0033684) |
| 0.0 | 0.1 | GO:0007271 | synaptic transmission, cholinergic(GO:0007271) |
| 0.0 | 0.0 | GO:0018202 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) peptidyl-histidine modification(GO:0018202) |
| 0.0 | 0.0 | GO:0034720 | histone H3-K4 demethylation(GO:0034720) |
| 0.0 | 0.1 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.0 | 0.0 | GO:0007625 | grooming behavior(GO:0007625) |
| 0.0 | 0.4 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.0 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.0 | GO:0060399 | positive regulation of growth hormone receptor signaling pathway(GO:0060399) |
| 0.0 | 0.0 | GO:0032988 | ribonucleoprotein complex disassembly(GO:0032988) |
| 0.0 | 0.0 | GO:0070213 | protein auto-ADP-ribosylation(GO:0070213) |
| 0.0 | 0.0 | GO:0097694 | establishment of RNA localization to telomere(GO:0097694) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.1 | 16.8 | GO:0043083 | synaptic cleft(GO:0043083) |
| 1.9 | 7.6 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 1.7 | 17.0 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 1.6 | 13.1 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 1.6 | 7.9 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 1.5 | 3.0 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 1.3 | 3.9 | GO:0072534 | perineuronal net(GO:0072534) |
| 1.2 | 20.2 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 1.2 | 61.4 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 1.1 | 34.2 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.9 | 13.0 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.9 | 6.3 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.9 | 2.7 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 0.9 | 2.6 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.9 | 4.3 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.8 | 5.9 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.8 | 4.0 | GO:0097433 | dense body(GO:0097433) |
| 0.8 | 5.3 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.7 | 23.2 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.7 | 5.0 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.7 | 2.1 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.7 | 6.9 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.7 | 5.4 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.6 | 3.2 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.6 | 8.1 | GO:0043196 | varicosity(GO:0043196) |
| 0.6 | 0.6 | GO:0042585 | germinal vesicle(GO:0042585) |
| 0.6 | 2.4 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.6 | 8.1 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.6 | 2.2 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.5 | 2.6 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.5 | 4.2 | GO:0030673 | axolemma(GO:0030673) |
| 0.5 | 4.6 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.5 | 1.5 | GO:0016461 | unconventional myosin complex(GO:0016461) |
| 0.5 | 11.1 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.5 | 1.4 | GO:0070110 | ciliary neurotrophic factor receptor complex(GO:0070110) |
| 0.4 | 4.5 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.4 | 64.3 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.4 | 2.0 | GO:0031315 | extrinsic component of mitochondrial outer membrane(GO:0031315) |
| 0.4 | 1.1 | GO:0097149 | centralspindlin complex(GO:0097149) |
| 0.4 | 11.6 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.4 | 0.7 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.4 | 0.7 | GO:0032777 | Piccolo NuA4 histone acetyltransferase complex(GO:0032777) |
| 0.4 | 2.1 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.3 | 0.7 | GO:0044316 | cone cell pedicle(GO:0044316) |
| 0.3 | 1.7 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.3 | 0.6 | GO:0033648 | host intracellular organelle(GO:0033647) host intracellular membrane-bounded organelle(GO:0033648) |
| 0.3 | 9.8 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.3 | 0.9 | GO:0031417 | NatC complex(GO:0031417) |
| 0.3 | 1.5 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.3 | 0.9 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.3 | 2.6 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.3 | 4.0 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.3 | 0.3 | GO:0035867 | alphav-beta3 integrin-IGF-1-IGF1R complex(GO:0035867) |
| 0.3 | 1.3 | GO:0071547 | piP-body(GO:0071547) |
| 0.3 | 0.3 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.2 | 1.0 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.2 | 0.7 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.2 | 0.5 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.2 | 0.9 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.2 | 1.1 | GO:0070695 | FHF complex(GO:0070695) |
| 0.2 | 0.7 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.2 | 3.6 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.2 | 0.8 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.2 | 0.6 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.2 | 0.4 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.2 | 0.4 | GO:0032437 | cuticular plate(GO:0032437) |
| 0.2 | 1.0 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.2 | 1.2 | GO:0005883 | neurofilament(GO:0005883) |
| 0.2 | 4.0 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.2 | 12.8 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.2 | 18.1 | GO:0030427 | site of polarized growth(GO:0030427) |
| 0.2 | 1.8 | GO:0034709 | methylosome(GO:0034709) |
| 0.2 | 0.5 | GO:1990696 | USH2 complex(GO:1990696) |
| 0.2 | 0.7 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.2 | 0.3 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.2 | 0.5 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.2 | 3.4 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.2 | 0.2 | GO:0000811 | GINS complex(GO:0000811) |
| 0.2 | 2.0 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.2 | 0.6 | GO:0033268 | node of Ranvier(GO:0033268) |
| 0.2 | 1.1 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.2 | 0.5 | GO:0005608 | laminin-3 complex(GO:0005608) |
| 0.2 | 1.5 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 0.4 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.1 | 0.4 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.1 | 1.1 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.1 | 4.4 | GO:0030175 | filopodium(GO:0030175) |
| 0.1 | 13.5 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.1 | 0.4 | GO:0020016 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.1 | 0.1 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.1 | 7.7 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 1.6 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 2.8 | GO:0016235 | aggresome(GO:0016235) |
| 0.1 | 0.8 | GO:0070688 | MLL5-L complex(GO:0070688) |
| 0.1 | 0.3 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.1 | 0.8 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 0.2 | GO:0005827 | polar microtubule(GO:0005827) |
| 0.1 | 0.3 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.1 | 0.5 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 0.1 | 0.1 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.1 | 0.4 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.1 | 0.2 | GO:0036194 | muscle cell projection(GO:0036194) muscle cell projection membrane(GO:0036195) |
| 0.1 | 4.5 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.1 | 0.1 | GO:0036488 | CHOP-C/EBP complex(GO:0036488) |
| 0.1 | 1.2 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.1 | 4.4 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.1 | 0.7 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.1 | 0.4 | GO:0072687 | meiotic spindle(GO:0072687) |
| 0.1 | 12.0 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 0.3 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.1 | 0.8 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.1 | 1.1 | GO:0043205 | fibril(GO:0043205) |
| 0.1 | 0.2 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.1 | 0.4 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.1 | 0.3 | GO:0033503 | HULC complex(GO:0033503) |
| 0.1 | 0.7 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 0.3 | GO:0071204 | histone pre-mRNA 3'end processing complex(GO:0071204) |
| 0.1 | 2.1 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 1.0 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.1 | 4.5 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.1 | 0.1 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.1 | 2.6 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 0.1 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.1 | 3.0 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.1 | 1.6 | GO:0043679 | axon terminus(GO:0043679) |
| 0.1 | 0.5 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 3.3 | GO:0043204 | perikaryon(GO:0043204) |
| 0.1 | 0.3 | GO:0089701 | U2AF(GO:0089701) |
| 0.1 | 0.9 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.1 | 0.6 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.1 | 0.4 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.1 | 0.2 | GO:0030891 | VCB complex(GO:0030891) |
| 0.1 | 0.2 | GO:0016222 | procollagen-proline 4-dioxygenase complex(GO:0016222) |
| 0.1 | 1.1 | GO:0030904 | retromer complex(GO:0030904) |
| 0.1 | 0.2 | GO:0000791 | euchromatin(GO:0000791) |
| 0.1 | 0.2 | GO:0097470 | ribbon synapse(GO:0097470) |
| 0.1 | 0.2 | GO:0043203 | axon hillock(GO:0043203) |
| 0.1 | 0.4 | GO:0042587 | glycogen granule(GO:0042587) |
| 0.1 | 1.5 | GO:0005844 | polysome(GO:0005844) |
| 0.1 | 0.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 1.4 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.1 | 0.1 | GO:0008275 | gamma-tubulin small complex(GO:0008275) |
| 0.1 | 0.1 | GO:0031261 | DNA replication preinitiation complex(GO:0031261) |
| 0.1 | 0.2 | GO:1990393 | 3M complex(GO:1990393) |
| 0.1 | 0.2 | GO:0031313 | extrinsic component of endosome membrane(GO:0031313) |
| 0.1 | 0.4 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.1 | 0.7 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.1 | 0.5 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 0.5 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.1 | 0.2 | GO:0001674 | female germ cell nucleus(GO:0001674) |
| 0.0 | 0.5 | GO:0033276 | transcription factor TFTC complex(GO:0033276) |
| 0.0 | 1.1 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.0 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.0 | 0.1 | GO:0002193 | MAML1-RBP-Jkappa- ICN1 complex(GO:0002193) |
| 0.0 | 0.2 | GO:0046696 | lipopolysaccharide receptor complex(GO:0046696) |
| 0.0 | 0.1 | GO:0010009 | cytoplasmic side of endosome membrane(GO:0010009) |
| 0.0 | 0.2 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 0.3 | GO:0044306 | neuron projection terminus(GO:0044306) |
| 0.0 | 0.5 | GO:0016327 | apicolateral plasma membrane(GO:0016327) |
| 0.0 | 0.1 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.2 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.0 | 0.3 | GO:0071541 | eukaryotic translation initiation factor 3 complex, eIF3m(GO:0071541) |
| 0.0 | 0.7 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.5 | GO:0097225 | sperm midpiece(GO:0097225) |
| 0.0 | 0.4 | GO:0000782 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) |
| 0.0 | 0.1 | GO:0043159 | acrosomal matrix(GO:0043159) |
| 0.0 | 0.0 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.3 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 0.2 | GO:0097539 | ciliary transition fiber(GO:0097539) |
| 0.0 | 0.1 | GO:0036396 | MIS complex(GO:0036396) |
| 0.0 | 0.1 | GO:0042406 | extrinsic component of endoplasmic reticulum membrane(GO:0042406) |
| 0.0 | 0.1 | GO:0016942 | insulin-like growth factor binding protein complex(GO:0016942) growth factor complex(GO:0036454) insulin-like growth factor ternary complex(GO:0042567) |
| 0.0 | 0.1 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.0 | 0.2 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.0 | 1.3 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.0 | 0.3 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.0 | 0.1 | GO:0002081 | outer acrosomal membrane(GO:0002081) |
| 0.0 | 0.2 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.0 | 0.3 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.0 | 0.3 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 0.0 | GO:0097422 | tubular endosome(GO:0097422) |
| 0.0 | 0.2 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.1 | GO:0017101 | aminoacyl-tRNA synthetase multienzyme complex(GO:0017101) |
| 0.0 | 0.2 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) |
| 0.0 | 0.5 | GO:0030286 | dynein complex(GO:0030286) |
| 0.0 | 0.1 | GO:0005638 | lamin filament(GO:0005638) |
| 0.0 | 0.2 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.0 | 0.0 | GO:0032589 | neuron projection membrane(GO:0032589) |
| 0.0 | 0.1 | GO:0070939 | Dsl1p complex(GO:0070939) |
| 0.0 | 0.0 | GO:0043511 | inhibin complex(GO:0043511) |
| 0.0 | 0.1 | GO:1990075 | periciliary membrane compartment(GO:1990075) |
| 0.0 | 0.2 | GO:0001741 | XY body(GO:0001741) |
| 0.0 | 0.0 | GO:0044393 | microspike(GO:0044393) |
| 0.0 | 0.5 | GO:0000314 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 0.5 | GO:0044439 | microbody part(GO:0044438) peroxisomal part(GO:0044439) |
| 0.0 | 0.0 | GO:0071953 | elastic fiber(GO:0071953) |
| 0.0 | 0.0 | GO:0071008 | U2-type post-mRNA release spliceosomal complex(GO:0071008) |
| 0.0 | 0.0 | GO:0031618 | nuclear pericentric heterochromatin(GO:0031618) |
| 0.0 | 0.0 | GO:0000814 | ESCRT II complex(GO:0000814) |
| 0.0 | 0.1 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.0 | 0.1 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.0 | GO:0031306 | intrinsic component of mitochondrial outer membrane(GO:0031306) |
| 0.0 | 0.0 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 2.2 | GO:0000790 | nuclear chromatin(GO:0000790) |
| 0.0 | 0.0 | GO:0030956 | glutamyl-tRNA(Gln) amidotransferase complex(GO:0030956) |
| 0.0 | 0.0 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.1 | GO:0000800 | lateral element(GO:0000800) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 12.2 | 36.7 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 2.6 | 18.3 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 2.6 | 5.2 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 2.3 | 6.9 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 2.2 | 4.3 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 2.0 | 11.9 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 1.8 | 21.6 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) |
| 1.6 | 8.2 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 1.6 | 6.5 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 1.6 | 6.4 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 1.5 | 4.6 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 1.5 | 4.5 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 1.5 | 11.8 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 1.4 | 4.3 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 1.4 | 9.9 | GO:0003680 | AT DNA binding(GO:0003680) |
| 1.4 | 4.1 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 1.3 | 11.6 | GO:0001091 | RNA polymerase II basal transcription factor binding(GO:0001091) |
| 1.2 | 6.1 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 1.2 | 4.7 | GO:0015375 | glycine:sodium symporter activity(GO:0015375) |
| 1.1 | 3.4 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 1.1 | 3.2 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 1.0 | 3.1 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 1.0 | 4.9 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 1.0 | 3.9 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.9 | 0.9 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 0.9 | 6.4 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.9 | 9.9 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.9 | 6.3 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.9 | 2.6 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.8 | 4.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 0.8 | 4.8 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 0.8 | 4.7 | GO:0001640 | adenylate cyclase inhibiting G-protein coupled glutamate receptor activity(GO:0001640) |
| 0.8 | 3.0 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.7 | 14.9 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.7 | 3.7 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.7 | 2.9 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.7 | 2.1 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.7 | 4.9 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.7 | 16.6 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.6 | 13.1 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.6 | 14.9 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.6 | 10.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.6 | 3.6 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.6 | 12.6 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.6 | 2.8 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.6 | 11.3 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.6 | 1.7 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) |
| 0.6 | 2.2 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.6 | 6.1 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.5 | 1.1 | GO:0098988 | G-protein coupled glutamate receptor activity(GO:0098988) |
| 0.5 | 1.1 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.5 | 3.1 | GO:0015198 | oligopeptide transporter activity(GO:0015198) |
| 0.5 | 1.5 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.5 | 4.0 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.5 | 3.9 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.5 | 4.3 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.5 | 7.0 | GO:0070273 | phosphatidylinositol-4-phosphate binding(GO:0070273) |
| 0.5 | 1.4 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) |
| 0.5 | 2.8 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.5 | 1.8 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.4 | 5.3 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.4 | 1.7 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.4 | 0.8 | GO:0097001 | ceramide binding(GO:0097001) |
| 0.4 | 1.2 | GO:0004814 | arginine-tRNA ligase activity(GO:0004814) |
| 0.4 | 1.2 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.4 | 1.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.4 | 1.2 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.4 | 1.5 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.4 | 6.8 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.4 | 0.8 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.4 | 5.6 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.4 | 2.9 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.4 | 3.9 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.4 | 1.8 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.4 | 1.4 | GO:0005167 | neurotrophin TRK receptor binding(GO:0005167) |
| 0.3 | 3.5 | GO:0008066 | glutamate receptor activity(GO:0008066) |
| 0.3 | 1.4 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.3 | 1.0 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.3 | 2.0 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.3 | 2.4 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.3 | 1.0 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.3 | 0.7 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 0.3 | 1.3 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.3 | 1.3 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
| 0.3 | 2.8 | GO:0046625 | sphingolipid binding(GO:0046625) |
| 0.3 | 1.2 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.3 | 0.3 | GO:0048763 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.3 | 3.8 | GO:0008601 | protein phosphatase type 2A regulator activity(GO:0008601) |
| 0.3 | 1.2 | GO:0052832 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.3 | 0.6 | GO:0010484 | H3 histone acetyltransferase activity(GO:0010484) |
| 0.3 | 0.8 | GO:0008309 | double-stranded DNA exodeoxyribonuclease activity(GO:0008309) |
| 0.3 | 1.3 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.3 | 4.4 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.3 | 1.0 | GO:0008486 | diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) |
| 0.3 | 1.6 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.3 | 0.3 | GO:0004676 | 3-phosphoinositide-dependent protein kinase activity(GO:0004676) |
| 0.3 | 0.8 | GO:0001875 | lipopolysaccharide receptor activity(GO:0001875) |
| 0.3 | 4.1 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.3 | 0.8 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.3 | 1.0 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.2 | 3.2 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.2 | 3.9 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.2 | 2.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.2 | 0.7 | GO:0008900 | hydrogen:potassium-exchanging ATPase activity(GO:0008900) |
| 0.2 | 4.5 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.2 | 0.9 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.2 | 1.8 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.2 | 5.5 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.2 | 6.2 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.2 | 4.4 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.2 | 3.2 | GO:0034236 | protein kinase A catalytic subunit binding(GO:0034236) |
| 0.2 | 5.3 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.2 | 0.6 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.2 | 0.8 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.2 | 3.6 | GO:0005112 | Notch binding(GO:0005112) |
| 0.2 | 0.2 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.2 | 1.3 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.2 | 3.5 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.2 | 0.6 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.2 | 1.0 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.2 | 0.2 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.2 | 1.2 | GO:0043008 | ATP-dependent protein binding(GO:0043008) |
| 0.2 | 2.8 | GO:0031402 | sodium ion binding(GO:0031402) |
| 0.2 | 0.6 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.2 | 0.6 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.2 | 0.8 | GO:0004897 | ciliary neurotrophic factor receptor activity(GO:0004897) |
| 0.2 | 1.9 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.2 | 0.7 | GO:0000213 | tRNA-intron endonuclease activity(GO:0000213) |
| 0.2 | 0.9 | GO:0004075 | biotin carboxylase activity(GO:0004075) |
| 0.2 | 0.7 | GO:0004528 | phosphodiesterase I activity(GO:0004528) |
| 0.2 | 2.2 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.2 | 1.8 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.2 | 2.0 | GO:0001223 | transcription coactivator binding(GO:0001223) |
| 0.2 | 2.8 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.2 | 0.7 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.2 | 2.1 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.2 | 1.2 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.2 | 1.0 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.2 | 0.3 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
| 0.2 | 2.0 | GO:0031005 | filamin binding(GO:0031005) |
| 0.2 | 1.9 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.2 | 3.2 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.2 | 0.3 | GO:0034191 | apolipoprotein A-I receptor binding(GO:0034191) |
| 0.2 | 11.2 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.2 | 0.5 | GO:0019153 | protein-disulfide reductase (glutathione) activity(GO:0019153) |
| 0.2 | 0.5 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.2 | 1.0 | GO:0003997 | acyl-CoA oxidase activity(GO:0003997) |
| 0.2 | 0.2 | GO:0046976 | histone methyltransferase activity (H3-K27 specific)(GO:0046976) |
| 0.2 | 0.3 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.2 | 0.5 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.1 | 0.4 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.1 | 0.1 | GO:0043125 | ErbB-3 class receptor binding(GO:0043125) |
| 0.1 | 0.1 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.1 | 0.4 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.1 | 1.0 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.1 | 0.4 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 0.4 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.1 | 1.0 | GO:0090079 | translation activator activity(GO:0008494) translation regulator activity, nucleic acid binding(GO:0090079) |
| 0.1 | 0.8 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.1 | 0.4 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.1 | 0.9 | GO:0043731 | 3-(3-hydroxyphenyl)propionate hydroxylase activity(GO:0008688) 4-chlorobenzaldehyde oxidase activity(GO:0018471) 3,5-xylenol methylhydroxylase activity(GO:0018630) phenylacetate hydroxylase activity(GO:0018631) 4-nitrophenol 4-monooxygenase activity(GO:0018632) dimethyl sulfide monooxygenase activity(GO:0018633) alpha-pinene monooxygenase [NADH] activity(GO:0018634) 1-hydroxy-2-naphthoate hydroxylase activity(GO:0018637) toluene 4-monooxygenase activity(GO:0018638) xylene monooxygenase activity(GO:0018639) dibenzothiophene monooxygenase activity(GO:0018640) 6-hydroxy-3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018641) chlorophenol 4-monooxygenase activity(GO:0018642) carbon disulfide oxygenase activity(GO:0018643) toluene 2-monooxygenase activity(GO:0018644) 1-hydroxy-2-oxolimonene 1,2-monooxygenase activity(GO:0018646) phenanthrene 1,2-monooxygenase activity(GO:0018647) tetrahydrofuran hydroxylase activity(GO:0018649) styrene monooxygenase activity(GO:0018650) toluene-4-sulfonate monooxygenase activity(GO:0018651) toluene-sulfonate methyl-monooxygenase activity(GO:0018652) 3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018653) 2-hydroxy-phenylacetate hydroxylase activity(GO:0018654) 2-oxo-delta3-4,5,5-trimethylcyclopentenylacetyl-CoA 1,2-monooxygenase activity(GO:0018655) phenanthrene 3,4-monooxygenase activity(GO:0018656) toluene 3-monooxygenase activity(GO:0018657) 4-hydroxyphenylacetate,NADH:oxygen oxidoreductase (3-hydroxylating) activity(GO:0018660) limonene monooxygenase activity(GO:0019113) 2-methylnaphthalene hydroxylase activity(GO:0034526) 1-methylnaphthalene hydroxylase activity(GO:0034534) bisphenol A hydroxylase A activity(GO:0034560) salicylate 5-hydroxylase activity(GO:0034785) isobutylamine N-hydroxylase activity(GO:0034791) branched-chain dodecylbenzene sulfonate monooxygenase activity(GO:0034802) 3-HSA hydroxylase activity(GO:0034819) 4-hydroxypyridine-3-hydroxylase activity(GO:0034894) 2-octaprenyl-3-methyl-6-methoxy-1,4-benzoquinol hydroxylase activity(GO:0043719) 6-hydroxynicotinate 3-monooxygenase activity(GO:0043731) thalianol hydroxylase activity(GO:0080014) |
| 0.1 | 0.3 | GO:0035851 | Krueppel-associated box domain binding(GO:0035851) |
| 0.1 | 0.4 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.1 | 0.7 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.1 | 0.4 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.1 | 0.7 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.1 | 1.1 | GO:0017166 | vinculin binding(GO:0017166) |
| 0.1 | 0.3 | GO:0001069 | regulatory region RNA binding(GO:0001069) |
| 0.1 | 0.8 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.1 | 0.6 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.3 | GO:1990932 | 5.8S rRNA binding(GO:1990932) |
| 0.1 | 2.0 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.1 | 0.1 | GO:0034903 | mono-butyltin dioxygenase activity(GO:0018586) tri-n-butyltin dioxygenase activity(GO:0018588) di-n-butyltin dioxygenase activity(GO:0018589) methylsilanetriol hydroxylase activity(GO:0018590) methyl tertiary butyl ether 3-monooxygenase activity(GO:0018591) 4-nitrocatechol 4-monooxygenase activity(GO:0018592) 4-chlorophenoxyacetate monooxygenase activity(GO:0018593) tert-butanol 2-monooxygenase activity(GO:0018594) alpha-pinene monooxygenase activity(GO:0018595) dimethylsilanediol hydroxylase activity(GO:0018596) ammonia monooxygenase activity(GO:0018597) hydroxymethylsilanetriol oxidase activity(GO:0018598) 2-hydroxyisobutyrate 3-monooxygenase activity(GO:0018599) alpha-pinene dehydrogenase activity(GO:0018600) bisphenol A hydroxylase B activity(GO:0034559) 2,2-bis(4-hydroxyphenyl)-1-propanol hydroxylase activity(GO:0034562) 9-fluorenone-3,4-dioxygenase activity(GO:0034786) anthracene 9,10-dioxygenase activity(GO:0034816) 2-(methylthio)benzothiazole monooxygenase activity(GO:0034857) 2-hydroxybenzothiazole monooxygenase activity(GO:0034858) benzothiazole monooxygenase activity(GO:0034859) 2,6-dihydroxybenzothiazole monooxygenase activity(GO:0034862) pinacolone 5-monooxygenase activity(GO:0034870) thioacetamide S-oxygenase activity(GO:0034873) thioacetamide S-oxide S-oxygenase activity(GO:0034874) endosulfan monooxygenase I activity(GO:0034888) N-nitrodimethylamine hydroxylase activity(GO:0034893) 4-(1-ethyl-1,4-dimethyl-pentyl)phenol monoxygenase activity(GO:0034897) endosulfan ether monooxygenase activity(GO:0034903) pyrene 4,5-monooxygenase activity(GO:0034925) pyrene 1,2-monooxygenase activity(GO:0034927) 1-hydroxypyrene 6,7-monooxygenase activity(GO:0034928) 1-hydroxypyrene 7,8-monooxygenase activity(GO:0034929) phenylboronic acid monooxygenase activity(GO:0034950) spheroidene monooxygenase activity(GO:0043823) |
| 0.1 | 0.7 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.1 | 0.2 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 2.0 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 0.2 | GO:1990825 | sequence-specific mRNA binding(GO:1990825) |
| 0.1 | 0.6 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 0.5 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.1 | 1.2 | GO:0022821 | potassium ion antiporter activity(GO:0022821) |
| 0.1 | 2.9 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 0.3 | GO:0019961 | interferon binding(GO:0019961) |
| 0.1 | 1.7 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.1 | 0.9 | GO:0005452 | inorganic anion exchanger activity(GO:0005452) |
| 0.1 | 0.9 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 0.3 | GO:0008732 | threonine aldolase activity(GO:0004793) L-allo-threonine aldolase activity(GO:0008732) |
| 0.1 | 0.6 | GO:0015185 | gamma-aminobutyric acid transmembrane transporter activity(GO:0015185) |
| 0.1 | 0.3 | GO:0089720 | caspase binding(GO:0089720) |
| 0.1 | 2.8 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 1.3 | GO:0045236 | CXCR chemokine receptor binding(GO:0045236) |
| 0.1 | 0.4 | GO:0048256 | flap endonuclease activity(GO:0048256) |
| 0.1 | 0.3 | GO:0004103 | choline kinase activity(GO:0004103) |
| 0.1 | 0.3 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.1 | 1.1 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.1 | 0.2 | GO:0050656 | 3'-phosphoadenosine 5'-phosphosulfate binding(GO:0050656) |
| 0.1 | 0.7 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 0.5 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 0.4 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.1 | 0.7 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.1 | 0.8 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.1 | 0.6 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.1 | 0.4 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 0.1 | 0.4 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.1 | 1.2 | GO:0043274 | phospholipase binding(GO:0043274) |
| 0.1 | 2.3 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.1 | 0.4 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.1 | 0.3 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.1 | 0.4 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.1 | 2.1 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 0.7 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 0.1 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.4 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.1 | 0.4 | GO:0102344 | 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.1 | 0.2 | GO:0008312 | 7S RNA binding(GO:0008312) |
| 0.1 | 0.9 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 0.3 | GO:0015556 | succinate transmembrane transporter activity(GO:0015141) C4-dicarboxylate transmembrane transporter activity(GO:0015556) |
| 0.1 | 0.5 | GO:0016937 | short-branched-chain-acyl-CoA dehydrogenase activity(GO:0016937) |
| 0.1 | 0.3 | GO:0071074 | eukaryotic initiation factor eIF2 binding(GO:0071074) |
| 0.1 | 0.6 | GO:0098505 | G-rich strand telomeric DNA binding(GO:0098505) |
| 0.1 | 5.5 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 0.4 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.1 | 0.2 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.1 | 0.9 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.1 | 1.9 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.1 | 1.0 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.5 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.1 | 0.1 | GO:1902282 | voltage-gated potassium channel activity involved in ventricular cardiac muscle cell action potential repolarization(GO:1902282) |
| 0.1 | 0.2 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.1 | 0.9 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 0.3 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.1 | 0.4 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.1 | 0.2 | GO:0008184 | glycogen phosphorylase activity(GO:0008184) |
| 0.1 | 1.6 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 0.6 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.1 | 2.7 | GO:0080131 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) heparan sulfate 2-O-sulfotransferase activity(GO:0004394) HNK-1 sulfotransferase activity(GO:0016232) heparan sulfate 6-O-sulfotransferase activity(GO:0017095) trans-9R,10R-dihydrodiolphenanthrene sulfotransferase activity(GO:0018721) 1-phenanthrol sulfotransferase activity(GO:0018722) 3-phenanthrol sulfotransferase activity(GO:0018723) 4-phenanthrol sulfotransferase activity(GO:0018724) trans-3,4-dihydrodiolphenanthrene sulfotransferase activity(GO:0018725) 9-phenanthrol sulfotransferase activity(GO:0018726) 2-phenanthrol sulfotransferase activity(GO:0018727) phenanthrol sulfotransferase activity(GO:0019111) 1-hydroxypyrene sulfotransferase activity(GO:0034930) proteoglycan sulfotransferase activity(GO:0050698) cholesterol sulfotransferase activity(GO:0051922) hydroxyjasmonate sulfotransferase activity(GO:0080131) |
| 0.1 | 0.5 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.1 | 0.8 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 1.4 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.1 | 0.3 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.1 | 0.8 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.1 | 0.1 | GO:0031748 | D1 dopamine receptor binding(GO:0031748) |
| 0.1 | 0.3 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.1 | 0.6 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.1 | 0.2 | GO:0016854 | racemase and epimerase activity(GO:0016854) |
| 0.1 | 0.4 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.1 | 0.1 | GO:0004661 | protein geranylgeranyltransferase activity(GO:0004661) |
| 0.1 | 0.1 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 1.2 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.1 | 0.4 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.1 | 0.2 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.1 | 0.1 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.1 | 0.7 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.1 | 1.4 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.1 | 1.1 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.1 | GO:0015222 | serotonin transmembrane transporter activity(GO:0015222) |
| 0.1 | 0.1 | GO:0070697 | activin receptor binding(GO:0070697) |
| 0.1 | 0.1 | GO:0050693 | LBD domain binding(GO:0050693) |
| 0.1 | 0.2 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.1 | 0.1 | GO:0015111 | iodide transmembrane transporter activity(GO:0015111) |
| 0.1 | 0.3 | GO:0015180 | L-alanine transmembrane transporter activity(GO:0015180) alanine transmembrane transporter activity(GO:0022858) |
| 0.1 | 11.5 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.1 | 0.8 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.1 | 0.5 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 0.3 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.1 | 0.2 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.1 | 0.4 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 0.1 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) |
| 0.1 | 0.2 | GO:0032190 | acrosin binding(GO:0032190) |
| 0.1 | 0.3 | GO:0047760 | butyrate-CoA ligase activity(GO:0047760) |
| 0.1 | 0.6 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.0 | 1.5 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.2 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.0 | 0.2 | GO:0030306 | ADP-ribosylation factor binding(GO:0030306) |
| 0.0 | 0.1 | GO:0008147 | structural constituent of bone(GO:0008147) |
| 0.0 | 6.0 | GO:0003729 | mRNA binding(GO:0003729) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.5 | GO:0048038 | quinone binding(GO:0048038) |
| 0.0 | 0.2 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.0 | 0.1 | GO:0019788 | NEDD8 transferase activity(GO:0019788) |
| 0.0 | 0.1 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.0 | 0.1 | GO:0030943 | mitochondrion targeting sequence binding(GO:0030943) |
| 0.0 | 0.1 | GO:0048045 | 4-hydroxybenzoate octaprenyltransferase activity(GO:0008412) protoheme IX farnesyltransferase activity(GO:0008495) (S)-2,3-di-O-geranylgeranylglyceryl phosphate synthase activity(GO:0043888) cadaverine aminopropyltransferase activity(GO:0043918) agmatine aminopropyltransferase activity(GO:0043919) 1,4-dihydroxy-2-naphthoate octaprenyltransferase activity(GO:0046428) trans-pentaprenyltranstransferase activity(GO:0048045) ATP dimethylallyltransferase activity(GO:0052622) ADP dimethylallyltransferase activity(GO:0052623) |
| 0.0 | 0.3 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 0.0 | 1.6 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.0 | 0.7 | GO:0001205 | transcriptional activator activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001205) |
| 0.0 | 0.2 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.0 | 0.2 | GO:0000386 | second spliceosomal transesterification activity(GO:0000386) |
| 0.0 | 3.6 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.3 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.0 | 0.1 | GO:0035514 | DNA demethylase activity(GO:0035514) |
| 0.0 | 0.3 | GO:0050681 | androgen receptor binding(GO:0050681) |
| 0.0 | 0.3 | GO:0031996 | thioesterase binding(GO:0031996) |
| 0.0 | 0.3 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.0 | 0.2 | GO:0004331 | fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.0 | 0.0 | GO:0004673 | protein histidine kinase activity(GO:0004673) |
| 0.0 | 0.8 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.0 | 0.1 | GO:0004461 | lactose synthase activity(GO:0004461) |
| 0.0 | 0.2 | GO:0016681 | ubiquinol-cytochrome-c reductase activity(GO:0008121) oxidoreductase activity, acting on diphenols and related substances as donors, cytochrome as acceptor(GO:0016681) |
| 0.0 | 0.1 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.0 | 0.1 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 0.0 | 0.2 | GO:0038132 | neuregulin binding(GO:0038132) |
| 0.0 | 0.1 | GO:0008107 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.0 | 3.3 | GO:0017137 | Rab GTPase binding(GO:0017137) |
| 0.0 | 0.1 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.0 | 0.0 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 0.1 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.0 | 0.8 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.1 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.0 | 0.1 | GO:0015216 | adenine nucleotide transmembrane transporter activity(GO:0000295) purine ribonucleotide transmembrane transporter activity(GO:0005346) purine nucleotide transmembrane transporter activity(GO:0015216) |
| 0.0 | 0.3 | GO:0004176 | ATP-dependent peptidase activity(GO:0004176) |
| 0.0 | 0.3 | GO:0016717 | oxidoreductase activity, acting on paired donors, with oxidation of a pair of donors resulting in the reduction of molecular oxygen to two molecules of water(GO:0016717) |
| 0.0 | 0.5 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 0.1 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.0 | 0.2 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 0.1 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.0 | 0.8 | GO:0097472 | cyclin-dependent protein serine/threonine kinase activity(GO:0004693) cyclin-dependent protein kinase activity(GO:0097472) |
| 0.0 | 0.1 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.0 | 0.5 | GO:0046966 | thyroid hormone receptor binding(GO:0046966) |
| 0.0 | 0.4 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.0 | 0.0 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.0 | 0.4 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.0 | 0.2 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.1 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.0 | 0.5 | GO:0003954 | NADH dehydrogenase activity(GO:0003954) |
| 0.0 | 0.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 0.1 | GO:0019776 | Atg8 ligase activity(GO:0019776) |
| 0.0 | 0.2 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.0 | 0.2 | GO:0032451 | demethylase activity(GO:0032451) |
| 0.0 | 0.1 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.0 | 0.3 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.0 | 0.6 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.1 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.0 | 0.0 | GO:0052872 | tocotrienol omega-hydroxylase activity(GO:0052872) |
| 0.0 | 1.0 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 0.1 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 0.0 | 0.1 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.0 | 0.1 | GO:0016774 | phosphotransferase activity, carboxyl group as acceptor(GO:0016774) |
| 0.0 | 0.1 | GO:0071209 | U7 snRNA binding(GO:0071209) |
| 0.0 | 0.2 | GO:0032794 | GTPase activating protein binding(GO:0032794) |
| 0.0 | 0.2 | GO:0031559 | oxidosqualene cyclase activity(GO:0031559) |
| 0.0 | 0.1 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.0 | 0.1 | GO:0003916 | DNA topoisomerase activity(GO:0003916) DNA topoisomerase type II (ATP-hydrolyzing) activity(GO:0003918) DNA topoisomerase II activity(GO:0061505) |
| 0.0 | 0.0 | GO:0035240 | dopamine binding(GO:0035240) |
| 0.0 | 11.2 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
| 0.0 | 0.0 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) |
| 0.0 | 0.1 | GO:0001025 | RNA polymerase III transcription factor binding(GO:0001025) |
| 0.0 | 0.0 | GO:0032357 | oxidized DNA binding(GO:0032356) oxidized purine DNA binding(GO:0032357) |
| 0.0 | 0.0 | GO:0005119 | smoothened binding(GO:0005119) |
| 0.0 | 0.5 | GO:0005048 | signal sequence binding(GO:0005048) |
| 0.0 | 0.1 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.0 | 0.1 | GO:0004616 | phosphogluconate dehydrogenase (decarboxylating) activity(GO:0004616) |
| 0.0 | 0.4 | GO:0061733 | peptide-lysine-N-acetyltransferase activity(GO:0061733) |
| 0.0 | 0.1 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.0 | 0.0 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.0 | 0.0 | GO:0001016 | RNA polymerase III regulatory region DNA binding(GO:0001016) |
| 0.0 | 0.0 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.0 | 0.0 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.0 | 0.1 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 1.5 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.0 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.0 | 0.0 | GO:0015142 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.0 | 0.0 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 0.2 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.0 | GO:0004027 | alcohol sulfotransferase activity(GO:0004027) |
| 0.0 | 0.0 | GO:0050567 | glutaminyl-tRNA synthase (glutamine-hydrolyzing) activity(GO:0050567) |
| 0.0 | 0.0 | GO:0042296 | ISG15 transferase activity(GO:0042296) |
| 0.0 | 0.0 | GO:0036122 | BMP binding(GO:0036122) |
| 0.0 | 0.0 | GO:0070653 | high-density lipoprotein particle receptor binding(GO:0070653) |
| 0.0 | 0.0 | GO:0051429 | corticotropin-releasing hormone receptor binding(GO:0051429) |
| 0.0 | 0.0 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.0 | 0.0 | GO:0008260 | 3-oxoacid CoA-transferase activity(GO:0008260) |
| 0.0 | 0.0 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.0 | 0.1 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 19.2 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.6 | 25.5 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.4 | 2.1 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.4 | 12.1 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.4 | 9.2 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.3 | 4.8 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.3 | 4.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.3 | 0.9 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.3 | 3.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.3 | 0.3 | PID IGF1 PATHWAY | IGF1 pathway |
| 0.2 | 3.4 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.2 | 5.6 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.2 | 3.0 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.2 | 6.7 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 8.5 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.1 | 2.2 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.1 | 0.3 | PID INTEGRIN4 PATHWAY | Alpha6 beta4 integrin-ligand interactions |
| 0.1 | 0.2 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.1 | 4.7 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.1 | 0.9 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 1.4 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 1.8 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 0.5 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 2.4 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 1.2 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 0.2 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.1 | 0.3 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.1 | 2.2 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.1 | 4.2 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 0.8 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.1 | 3.4 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 1.6 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.1 | 1.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 0.3 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.1 | 1.0 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.1 | 1.1 | PID ARF 3PATHWAY | Arf1 pathway |
| 0.1 | 0.8 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 1.7 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 0.8 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.0 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 0.4 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 0.3 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 0.3 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 0.2 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.0 | 0.3 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.0 | 0.0 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.0 | 0.1 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.2 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.0 | 3.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.2 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 0.1 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.0 | 0.1 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 0.3 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.0 | 0.2 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 0.0 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 12.3 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 1.1 | 22.0 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 1.0 | 7.6 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.9 | 0.9 | REACTOME SIGNAL REGULATORY PROTEIN SIRP FAMILY INTERACTIONS | Genes involved in Signal regulatory protein (SIRP) family interactions |
| 0.9 | 13.0 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.8 | 0.8 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.8 | 15.9 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.7 | 16.3 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.7 | 7.8 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.6 | 4.7 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.5 | 8.1 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.5 | 8.5 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.5 | 5.1 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.5 | 7.6 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.5 | 14.3 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.4 | 19.6 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.4 | 8.2 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.4 | 0.7 | REACTOME VIRAL MESSENGER RNA SYNTHESIS | Genes involved in Viral Messenger RNA Synthesis |
| 0.3 | 4.9 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.3 | 1.0 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.3 | 4.5 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.3 | 4.5 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.3 | 3.4 | REACTOME TRAFFICKING OF GLUR2 CONTAINING AMPA RECEPTORS | Genes involved in Trafficking of GluR2-containing AMPA receptors |
| 0.3 | 11.7 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.2 | 4.6 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.2 | 0.7 | REACTOME BILE ACID AND BILE SALT METABOLISM | Genes involved in Bile acid and bile salt metabolism |
| 0.2 | 2.6 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.2 | 4.3 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.2 | 1.5 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.2 | 1.3 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.2 | 2.1 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 0.2 | 1.6 | REACTOME BASE FREE SUGAR PHOSPHATE REMOVAL VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Base-free sugar-phosphate removal via the single-nucleotide replacement pathway |
| 0.2 | 2.7 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.2 | 1.7 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 2.1 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.1 | 1.5 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 1.1 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 5.2 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.1 | 0.6 | REACTOME REGULATION OF INSULIN SECRETION BY GLUCAGON LIKE PEPTIDE1 | Genes involved in Regulation of Insulin Secretion by Glucagon-like Peptide-1 |
| 0.1 | 2.3 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.5 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 1.8 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.1 | 1.7 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 1.2 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 0.1 | REACTOME GRB2 SOS PROVIDES LINKAGE TO MAPK SIGNALING FOR INTERGRINS | Genes involved in GRB2:SOS provides linkage to MAPK signaling for Intergrins |
| 0.1 | 1.7 | REACTOME RORA ACTIVATES CIRCADIAN EXPRESSION | Genes involved in RORA Activates Circadian Expression |
| 0.1 | 0.8 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.1 | 1.2 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 0.3 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.1 | 1.0 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.1 | 0.1 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.1 | 1.4 | REACTOME CD28 DEPENDENT PI3K AKT SIGNALING | Genes involved in CD28 dependent PI3K/Akt signaling |
| 0.1 | 3.2 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.1 | 2.0 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.1 | 0.7 | REACTOME SIGNALING BY NOTCH1 | Genes involved in Signaling by NOTCH1 |
| 0.1 | 1.2 | REACTOME REGULATORY RNA PATHWAYS | Genes involved in Regulatory RNA pathways |
| 0.1 | 0.9 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 0.1 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 1.3 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 0.2 | REACTOME P53 INDEPENDENT G1 S DNA DAMAGE CHECKPOINT | Genes involved in p53-Independent G1/S DNA damage checkpoint |
| 0.1 | 0.6 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.1 | 0.1 | REACTOME NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Neurotransmitter Release Cycle |
| 0.0 | 7.7 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.3 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.8 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.0 | 0.5 | REACTOME NCAM SIGNALING FOR NEURITE OUT GROWTH | Genes involved in NCAM signaling for neurite out-growth |
| 0.0 | 0.5 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.0 | 0.1 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.4 | REACTOME P75NTR SIGNALS VIA NFKB | Genes involved in p75NTR signals via NF-kB |
| 0.0 | 0.1 | REACTOME GPVI MEDIATED ACTIVATION CASCADE | Genes involved in GPVI-mediated activation cascade |
| 0.0 | 0.2 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.0 | 0.5 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.0 | 0.3 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.0 | 0.9 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.0 | REACTOME NFKB ACTIVATION THROUGH FADD RIP1 PATHWAY MEDIATED BY CASPASE 8 AND10 | Genes involved in NF-kB activation through FADD/RIP-1 pathway mediated by caspase-8 and -10 |
| 0.0 | 0.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.2 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.0 | 0.1 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.0 | 0.1 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 1.1 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.1 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.0 | REACTOME PLATELET ADHESION TO EXPOSED COLLAGEN | Genes involved in Platelet Adhesion to exposed collagen |
| 0.0 | 0.0 | REACTOME RESOLUTION OF AP SITES VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Resolution of AP sites via the single-nucleotide replacement pathway |
| 0.0 | 0.0 | REACTOME SEMA3A PAK DEPENDENT AXON REPULSION | Genes involved in Sema3A PAK dependent Axon repulsion |
| 0.0 | 0.1 | REACTOME OPIOID SIGNALLING | Genes involved in Opioid Signalling |
| 0.0 | 0.0 | REACTOME ACTIVATED TAK1 MEDIATES P38 MAPK ACTIVATION | Genes involved in activated TAK1 mediates p38 MAPK activation |
| 0.0 | 0.1 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.1 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.0 | 0.1 | REACTOME METABOLISM OF STEROID HORMONES AND VITAMINS A AND D | Genes involved in Metabolism of steroid hormones and vitamins A and D |
| 0.0 | 0.0 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |
| 0.0 | 0.1 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.0 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.2 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 0.0 | REACTOME ABORTIVE ELONGATION OF HIV1 TRANSCRIPT IN THE ABSENCE OF TAT | Genes involved in Abortive elongation of HIV-1 transcript in the absence of Tat |
| 0.0 | 0.2 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |