Project
ENCODE: H3K4me3 ChIP-Seq of different mouse tissues during embryonic development
Navigation
Downloads
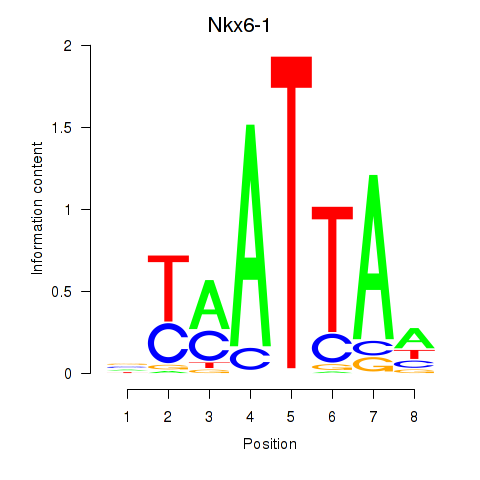
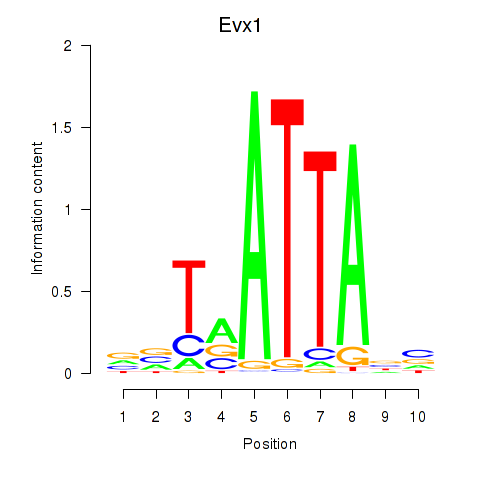
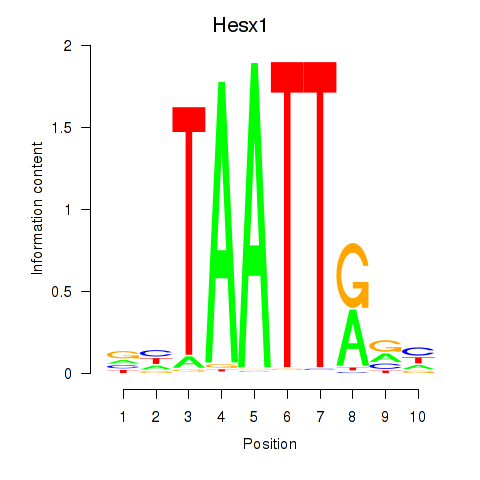
Results for Nkx6-1_Evx1_Hesx1
Z-value: 1.30
Transcription factors associated with Nkx6-1_Evx1_Hesx1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Nkx6-1
|
ENSMUSG00000035187.8 | NK6 homeobox 1 |
|
Evx1
|
ENSMUSG00000005503.8 | even-skipped homeobox 1 |
|
Hesx1
|
ENSMUSG00000040726.8 | homeobox gene expressed in ES cells |
Correlations of motif activity and signal intensity at CREs associated with the motif's TFs:
This plot shows correlation between observed signal intensity of a CRE associated with the transcription factor across all samples and activity of the motif.
For each TF, only the top 5 correlated CREs are shown.
| CRE | Gene | Distance | Association probability | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|---|---|
| chr6_52313324_52314938 | Evx1 | 579 | 0.384730 | -0.25 | 5.3e-02 | Click! |
| chr6_52305793_52306730 | Evx1 | 7237 | 0.084477 | -0.21 | 1.1e-01 | Click! |
| chr6_52309069_52309254 | Evx1 | 4337 | 0.097117 | -0.10 | 4.5e-01 | Click! |
| chr6_52306757_52306908 | Evx1 | 6666 | 0.086061 | 0.08 | 5.6e-01 | Click! |
| chr6_52309839_52312950 | Evx1 | 2104 | 0.142938 | -0.05 | 6.9e-01 | Click! |
| chr14_26994035_26994822 | Hesx1 | 12 | 0.977374 | -0.29 | 2.3e-02 | Click! |
| chr14_26991792_26991943 | Hesx1 | 2549 | 0.266698 | -0.13 | 3.4e-01 | Click! |
| chr14_27000422_27001594 | Hesx1 | 646 | 0.711316 | -0.11 | 4.1e-01 | Click! |
| chr5_101665386_101665820 | Nkx6-1 | 607 | 0.735039 | -0.49 | 8.5e-05 | Click! |
| chr5_101663336_101663763 | Nkx6-1 | 1447 | 0.412377 | -0.46 | 1.9e-04 | Click! |
| chr5_101663868_101665354 | Nkx6-1 | 385 | 0.859131 | -0.41 | 1.1e-03 | Click! |
| chr5_101647553_101648558 | Nkx6-1 | 16941 | 0.179932 | -0.27 | 4.0e-02 | Click! |
| chr5_101666359_101666510 | Nkx6-1 | 1438 | 0.410546 | 0.19 | 1.4e-01 | Click! |
Activity of the Nkx6-1_Evx1_Hesx1 motif across conditions
Conditions sorted by the z-value of the Nkx6-1_Evx1_Hesx1 motif activity
Move your cursor over a bar to see sample name and corresponding Z-value.
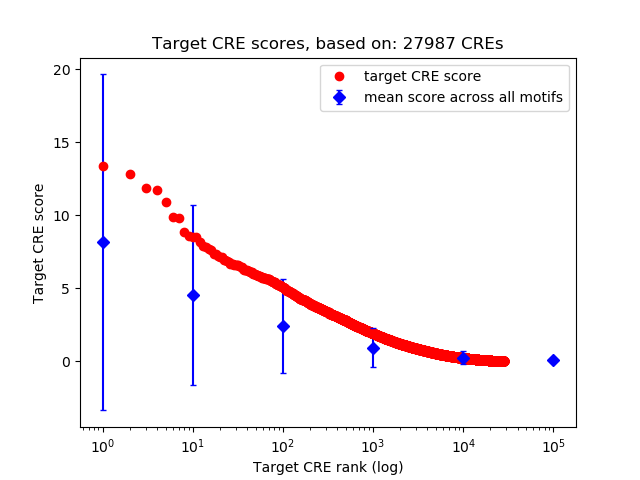
Top target CREs of the motif:
| Cis Regulatory Element (CRE) | Target Score | Top associated gene | Gene Info | Distance of CRE to TSS | CRE/Gene association probability |
|---|---|---|---|---|---|
| chr2_6881042_6881689 | 13.39 |
Gm13389 |
predicted gene 13389 |
2905 |
0.24 |
| chr12_52700044_52701597 | 12.86 |
Akap6 |
A kinase (PRKA) anchor protein 6 |
1437 |
0.46 |
| chr18_23036665_23037864 | 11.85 |
Nol4 |
nucleolar protein 4 |
1392 |
0.59 |
| chr3_4796861_4798079 | 11.72 |
1110015O18Rik |
RIKEN cDNA 1110015O18 gene |
88 |
0.98 |
| chr6_55678280_55679200 | 10.91 |
Neurod6 |
neurogenic differentiation 6 |
2523 |
0.32 |
| chr1_81077232_81078427 | 9.86 |
Nyap2 |
neuronal tyrosine-phophorylated phosphoinositide 3-kinase adaptor 2 |
246 |
0.96 |
| chr8_45508499_45509041 | 9.79 |
Sorbs2 |
sorbin and SH3 domain containing 2 |
852 |
0.61 |
| chr7_78885666_78887958 | 8.88 |
Mir7-2 |
microRNA 7-2 |
1465 |
0.28 |
| chr8_109248831_109249717 | 8.60 |
D030068K23Rik |
RIKEN cDNA D030068K23 gene |
592 |
0.83 |
| chr2_105675959_105678109 | 8.50 |
Pax6 |
paired box 6 |
905 |
0.54 |
| chr12_49387532_49388566 | 8.50 |
3110039M20Rik |
RIKEN cDNA 3110039M20 gene |
1603 |
0.26 |
| chr10_90577565_90578158 | 8.20 |
Anks1b |
ankyrin repeat and sterile alpha motif domain containing 1B |
869 |
0.72 |
| chr2_57613916_57615034 | 7.89 |
Gm13532 |
predicted gene 13532 |
14753 |
0.2 |
| chr13_83732205_83734272 | 7.84 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
672 |
0.58 |
| chr16_91320391_91321321 | 7.70 |
Gm15966 |
predicted gene 15966 |
4884 |
0.15 |
| chr12_107990188_107992301 | 7.64 |
Bcl11b |
B cell leukemia/lymphoma 11B |
12170 |
0.28 |
| chr17_51760240_51761547 | 7.35 |
C230085N15Rik |
RIKEN cDNA C230085N15 gene |
728 |
0.54 |
| chr15_25415436_25415919 | 7.33 |
Gm48957 |
predicted gene, 48957 |
614 |
0.58 |
| chr2_53501543_53502209 | 7.23 |
Gm13503 |
predicted gene 13503 |
50050 |
0.17 |
| chr2_6881874_6882908 | 7.17 |
Gm13389 |
predicted gene 13389 |
1879 |
0.3 |
| chr13_83715222_83716973 | 7.14 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
5284 |
0.15 |
| chr1_186278091_186278855 | 6.97 |
Gm37491 |
predicted gene, 37491 |
68842 |
0.11 |
| chr5_98182267_98183697 | 6.92 |
Prdm8 |
PR domain containing 8 |
2004 |
0.26 |
| chr15_30458403_30458947 | 6.85 |
Ctnnd2 |
catenin (cadherin associated protein), delta 2 |
887 |
0.67 |
| chr10_29143400_29144848 | 6.82 |
Soga3 |
SOGA family member 3 |
65 |
0.5 |
| chr4_25797578_25797990 | 6.68 |
Fut9 |
fucosyltransferase 9 |
2071 |
0.32 |
| chr8_109250884_109251908 | 6.68 |
D030068K23Rik |
RIKEN cDNA D030068K23 gene |
1530 |
0.52 |
| chr16_77596529_77597235 | 6.64 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
1980 |
0.16 |
| chr17_62659451_62660256 | 6.63 |
Gm25800 |
predicted gene, 25800 |
202733 |
0.03 |
| chr11_108606161_108607110 | 6.62 |
Cep112 |
centrosomal protein 112 |
1408 |
0.5 |
| chr18_37217058_37218378 | 6.61 |
Gm10544 |
predicted gene 10544 |
39196 |
0.08 |
| chr3_67892808_67893503 | 6.59 |
Iqschfp |
Iqcj and Schip1 fusion protein |
923 |
0.42 |
| chr7_92234907_92236280 | 6.50 |
Dlg2 |
discs large MAGUK scaffold protein 2 |
466 |
0.88 |
| chr8_34890130_34891317 | 6.49 |
Tnks |
tankyrase, TRF1-interacting ankyrin-related ADP-ribose polymerase |
572 |
0.8 |
| chr10_90578974_90579573 | 6.46 |
Anks1b |
ankyrin repeat and sterile alpha motif domain containing 1B |
2281 |
0.42 |
| chr13_73117045_73117937 | 6.30 |
Rpl31-ps2 |
ribosomal protein L31, pseudogene 2 |
115904 |
0.06 |
| chr17_90452868_90453681 | 6.28 |
Nrxn1 |
neurexin I |
1548 |
0.36 |
| chr4_35844509_35845617 | 6.28 |
Lingo2 |
leucine rich repeat and Ig domain containing 2 |
141 |
0.98 |
| chr3_79144294_79146166 | 6.24 |
Rapgef2 |
Rap guanine nucleotide exchange factor (GEF) 2 |
253 |
0.94 |
| chr12_89815214_89815490 | 6.24 |
Nrxn3 |
neurexin III |
2869 |
0.41 |
| chr8_49462071_49462635 | 6.17 |
4930555F03Rik |
RIKEN cDNA 4930555F03 gene |
970 |
0.52 |
| chr14_64592977_64593388 | 6.16 |
Mir3078 |
microRNA 3078 |
1997 |
0.25 |
| chr8_14382368_14383445 | 6.15 |
Dlgap2 |
DLG associated protein 2 |
910 |
0.66 |
| chr4_23602363_23602952 | 6.12 |
Gm25978 |
predicted gene, 25978 |
24088 |
0.22 |
| chr9_41585694_41587243 | 6.09 |
Mir100hg |
Mir100 Mirlet7a-2 Mir125b-1 cluster host gene |
1301 |
0.29 |
| chr1_66386919_66387899 | 6.04 |
Map2 |
microtubule-associated protein 2 |
398 |
0.87 |
| chr12_46816152_46816702 | 5.96 |
Nova1 |
NOVA alternative splicing regulator 1 |
533 |
0.8 |
| chr4_82915737_82916950 | 5.96 |
Frem1 |
Fras1 related extracellular matrix protein 1 |
1909 |
0.38 |
| chr4_110285249_110285423 | 5.95 |
Elavl4 |
ELAV like RNA binding protein 4 |
1280 |
0.61 |
| chr13_105247534_105248250 | 5.94 |
Rnf180 |
ring finger protein 180 |
23147 |
0.21 |
| chr9_74977325_74977617 | 5.92 |
Fam214a |
family with sequence similarity 214, member A |
1360 |
0.45 |
| chr3_66746318_66747483 | 5.90 |
Gm6555 |
predicted gene 6555 |
135450 |
0.05 |
| chr17_59279898_59280169 | 5.85 |
Gm23769 |
predicted gene, 23769 |
54190 |
0.17 |
| chr8_41054476_41055299 | 5.85 |
Mtus1 |
mitochondrial tumor suppressor 1 |
93 |
0.95 |
| chr13_44842150_44842855 | 5.84 |
Jarid2 |
jumonji, AT rich interactive domain 2 |
1719 |
0.39 |
| chr7_87586513_87587584 | 5.82 |
Grm5 |
glutamate receptor, metabotropic 5 |
2650 |
0.4 |
| chr2_65620767_65621991 | 5.75 |
Scn2a |
sodium channel, voltage-gated, type II, alpha |
568 |
0.82 |
| chr8_96455054_96456367 | 5.74 |
Gm32122 |
predicted gene, 32122 |
51848 |
0.14 |
| chr18_43686487_43688415 | 5.74 |
Jakmip2 |
janus kinase and microtubule interacting protein 2 |
174 |
0.96 |
| chr11_36676450_36677161 | 5.73 |
Tenm2 |
teneurin transmembrane protein 2 |
940 |
0.7 |
| chr18_25745414_25746450 | 5.71 |
Celf4 |
CUGBP, Elav-like family member 4 |
6760 |
0.24 |
| chr3_13946382_13947629 | 5.70 |
Ralyl |
RALY RNA binding protein-like |
594 |
0.84 |
| chr1_66324716_66324867 | 5.70 |
Map2 |
microtubule-associated protein 2 |
2689 |
0.25 |
| chr6_32584464_32585789 | 5.68 |
Plxna4 |
plexin A4 |
3066 |
0.3 |
| chr5_150262108_150262988 | 5.67 |
Fry |
FRY microtubule binding protein |
2781 |
0.26 |
| chr18_69500231_69501482 | 5.66 |
Tcf4 |
transcription factor 4 |
20 |
0.99 |
| chr12_71048832_71049275 | 5.65 |
Arid4a |
AT rich interactive domain 4A (RBP1-like) |
712 |
0.65 |
| chr10_111247804_111248910 | 5.64 |
Osbpl8 |
oxysterol binding protein-like 8 |
289 |
0.91 |
| chr16_43504464_43505047 | 5.63 |
Zbtb20 |
zinc finger and BTB domain containing 20 |
1058 |
0.61 |
| chr2_106512529_106513407 | 5.63 |
Gm14015 |
predicted gene 14015 |
10135 |
0.26 |
| chr16_77594640_77595970 | 5.62 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
403 |
0.71 |
| chr18_80984086_80984990 | 5.56 |
Sall3 |
spalt like transcription factor 3 |
1998 |
0.23 |
| chr4_54950838_54951442 | 5.51 |
Zfp462 |
zinc finger protein 462 |
3164 |
0.35 |
| chr19_58076835_58077494 | 5.49 |
Gm50287 |
predicted gene, 50287 |
18867 |
0.22 |
| chr14_55052408_55052978 | 5.49 |
Zfhx2os |
zinc finger homeobox 2, opposite strand |
1176 |
0.24 |
| chr2_181763361_181764530 | 5.48 |
Myt1 |
myelin transcription factor 1 |
613 |
0.66 |
| chr1_42259362_42260538 | 5.46 |
Gm28175 |
predicted gene 28175 |
1905 |
0.34 |
| chr11_19713213_19714451 | 5.45 |
Glns-ps1 |
glutamine synthetase pseudogene 1 |
6523 |
0.2 |
| chr2_181766837_181767244 | 5.45 |
Myt1 |
myelin transcription factor 1 |
2 |
0.97 |
| chr6_138424419_138425583 | 5.44 |
Lmo3 |
LIM domain only 3 |
386 |
0.83 |
| chrX_105390628_105392456 | 5.43 |
5330434G04Rik |
RIKEN cDNA 5330434G04 gene |
212 |
0.93 |
| chr6_143259703_143261097 | 5.38 |
D6Ertd474e |
DNA segment, Chr 6, ERATO Doi 474, expressed |
14507 |
0.2 |
| chr4_5724213_5725550 | 5.31 |
Fam110b |
family with sequence similarity 110, member B |
569 |
0.81 |
| chr1_138346039_138346510 | 5.30 |
Gm28500 |
predicted gene 28500 |
30990 |
0.17 |
| chr1_159614722_159615352 | 5.29 |
Gm10530 |
predicted gene 10530 |
288 |
0.92 |
| chr10_57784547_57786586 | 5.28 |
Fabp7 |
fatty acid binding protein 7, brain |
643 |
0.68 |
| chr8_12398370_12398923 | 5.27 |
Gm25239 |
predicted gene, 25239 |
2243 |
0.21 |
| chr19_38348426_38349048 | 5.24 |
Gm50150 |
predicted gene, 50150 |
6123 |
0.16 |
| chr13_83724063_83724214 | 5.24 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
2757 |
0.18 |
| chr2_38340293_38341511 | 5.23 |
Lhx2 |
LIM homeobox protein 2 |
190 |
0.93 |
| chr5_107497766_107498034 | 5.21 |
Btbd8 |
BTB (POZ) domain containing 8 |
121 |
0.94 |
| chr8_108536302_108537095 | 5.21 |
Gm39244 |
predicted gene, 39244 |
249 |
0.95 |
| chr11_34315414_34316667 | 5.20 |
Insyn2b |
inhibitory synaptic factor family member 2B |
1218 |
0.45 |
| chr11_43270080_43270726 | 5.17 |
Gm12146 |
predicted gene 12146 |
10619 |
0.19 |
| chr16_77236514_77236677 | 5.16 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
276 |
0.93 |
| chr13_94645221_94645665 | 5.15 |
Gm48287 |
predicted gene, 48287 |
1901 |
0.32 |
| chr2_97478080_97479309 | 5.11 |
Lrrc4c |
leucine rich repeat containing 4C |
10605 |
0.3 |
| chr1_143644977_143645827 | 5.11 |
Cdc73 |
cell division cycle 73, Paf1/RNA polymerase II complex component |
2877 |
0.24 |
| chr8_88793832_88794104 | 5.10 |
Rps6-ps2 |
ribosomal protein S6, pseudogene 2 |
12423 |
0.2 |
| chr2_72426765_72427714 | 5.09 |
Cdca7 |
cell division cycle associated 7 |
48920 |
0.13 |
| chr2_32428080_32429746 | 5.06 |
Slc25a25 |
solute carrier family 25 (mitochondrial carrier, phosphate carrier), member 25 |
1839 |
0.19 |
| chr13_83739310_83740387 | 5.01 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
985 |
0.29 |
| chr4_70530858_70531844 | 5.00 |
Megf9 |
multiple EGF-like-domains 9 |
3577 |
0.38 |
| chr13_97248475_97250229 | 5.00 |
Enc1 |
ectodermal-neural cortex 1 |
8247 |
0.17 |
| chr5_131615202_131615755 | 4.99 |
2810432F15Rik |
RIKEN cDNA 2810432F15 gene |
40 |
0.94 |
| chrX_49272929_49273965 | 4.93 |
Enox2 |
ecto-NOX disulfide-thiol exchanger 2 |
14765 |
0.24 |
| chr4_97582473_97584218 | 4.92 |
E130114P18Rik |
RIKEN cDNA E130114P18 gene |
1251 |
0.53 |
| chrX_133682515_133683917 | 4.91 |
Pcdh19 |
protocadherin 19 |
1775 |
0.49 |
| chr6_103513736_103514218 | 4.91 |
Chl1 |
cell adhesion molecule L1-like |
2647 |
0.25 |
| chr4_33926104_33927188 | 4.88 |
Cnr1 |
cannabinoid receptor 1 (brain) |
444 |
0.88 |
| chrX_110811626_110812334 | 4.84 |
Gm44593 |
predicted gene 44593 |
344 |
0.89 |
| chr7_128690432_128691249 | 4.84 |
Gm16044 |
predicted gene 16044 |
1849 |
0.17 |
| chr2_80126598_80127760 | 4.84 |
Pde1a |
phosphodiesterase 1A, calmodulin-dependent |
1655 |
0.42 |
| chr12_29529828_29531185 | 4.84 |
Gm20208 |
predicted gene, 20208 |
609 |
0.74 |
| chr3_68573207_68574269 | 4.83 |
Schip1 |
schwannomin interacting protein 1 |
1493 |
0.45 |
| chr6_138872923_138873486 | 4.81 |
Gm9038 |
predicted gene 9038 |
13080 |
0.29 |
| chr10_42579057_42580702 | 4.81 |
Nr2e1 |
nuclear receptor subfamily 2, group E, member 1 |
466 |
0.81 |
| chr1_173389412_173390669 | 4.81 |
Cadm3 |
cell adhesion molecule 3 |
22345 |
0.14 |
| chr2_52557337_52558561 | 4.78 |
Cacnb4 |
calcium channel, voltage-dependent, beta 4 subunit |
611 |
0.74 |
| chr19_24371923_24372796 | 4.77 |
Pip5k1bos |
phosphatidylinositol-4-phosphate 5-kinase, type 1 beta, opposite strand |
43650 |
0.13 |
| chr18_25749825_25750329 | 4.76 |
Celf4 |
CUGBP, Elav-like family member 4 |
2615 |
0.33 |
| chr8_88813002_88813481 | 4.74 |
Rps6-ps2 |
ribosomal protein S6, pseudogene 2 |
6850 |
0.22 |
| chrX_153501207_153502250 | 4.69 |
Ubqln2 |
ubiquilin 2 |
3501 |
0.22 |
| chr2_79456259_79457200 | 4.68 |
Neurod1 |
neurogenic differentiation 1 |
22 |
0.52 |
| chr4_24429901_24430719 | 4.68 |
Gm27243 |
predicted gene 27243 |
580 |
0.79 |
| chr8_55939901_55941088 | 4.68 |
Glra3 |
glycine receptor, alpha 3 subunit |
15 |
0.98 |
| chr13_14520339_14520490 | 4.67 |
Gm30893 |
predicted gene, 30893 |
1072 |
0.37 |
| chr2_6867243_6867952 | 4.65 |
Celf2 |
CUGBP, Elav-like family member 2 |
4375 |
0.24 |
| chr6_55680133_55680881 | 4.63 |
Neurod6 |
neurogenic differentiation 6 |
756 |
0.69 |
| chrX_23283125_23283785 | 4.62 |
Klhl13 |
kelch-like 13 |
1374 |
0.57 |
| chr13_99443316_99444666 | 4.61 |
Map1b |
microtubule-associated protein 1B |
47 |
0.98 |
| chr4_125492765_125493053 | 4.58 |
Grik3 |
glutamate receptor, ionotropic, kainate 3 |
2209 |
0.31 |
| chr10_119820571_119821220 | 4.58 |
Grip1 |
glutamate receptor interacting protein 1 |
1370 |
0.48 |
| chr15_88561170_88561806 | 4.57 |
Zdhhc25 |
zinc finger, DHHC domain containing 25 |
38814 |
0.19 |
| chr12_68997254_68997526 | 4.55 |
Gm47515 |
predicted gene, 47515 |
2420 |
0.27 |
| chr4_24429141_24429555 | 4.54 |
Gm27243 |
predicted gene 27243 |
1542 |
0.44 |
| chr18_81509510_81510030 | 4.53 |
Gm50412 |
predicted gene, 50412 |
29413 |
0.19 |
| chr1_109984209_109985108 | 4.51 |
Cdh7 |
cadherin 7, type 2 |
921 |
0.74 |
| chr3_154816919_154817899 | 4.50 |
Gm18589 |
predicted gene, 18589 |
22198 |
0.2 |
| chr1_64116857_64117480 | 4.47 |
Klf7 |
Kruppel-like factor 7 (ubiquitous) |
4314 |
0.23 |
| chr5_112574293_112574972 | 4.47 |
Sez6l |
seizure related 6 homolog like |
2236 |
0.24 |
| chr4_55928799_55929534 | 4.46 |
Gm12519 |
predicted gene 12519 |
64573 |
0.14 |
| chr8_33746530_33746945 | 4.46 |
Smim18 |
small integral membrane protein 18 |
1033 |
0.44 |
| chr15_78118149_78118442 | 4.44 |
A730060N03Rik |
RIKEN cDNA A730060N03 gene |
1411 |
0.31 |
| chr18_84082553_84082704 | 4.43 |
Tshz1 |
teashirt zinc finger family member 1 |
2447 |
0.24 |
| chr14_22037484_22037938 | 4.43 |
Gm7480 |
predicted gene 7480 |
1608 |
0.33 |
| chrX_166347339_166348040 | 4.43 |
Gpm6b |
glycoprotein m6b |
2847 |
0.32 |
| chr8_54954519_54955779 | 4.41 |
Gpm6a |
glycoprotein m6a |
306 |
0.88 |
| chr10_110454344_110454630 | 4.38 |
Nav3 |
neuron navigator 3 |
1717 |
0.42 |
| chr18_25750468_25751272 | 4.36 |
Celf4 |
CUGBP, Elav-like family member 4 |
1822 |
0.41 |
| chr1_155414821_155416000 | 4.34 |
Xpr1 |
xenotropic and polytropic retrovirus receptor 1 |
1919 |
0.42 |
| chr13_8205494_8206737 | 4.33 |
Adarb2 |
adenosine deaminase, RNA-specific, B2 |
3193 |
0.23 |
| chr12_108605770_108606876 | 4.32 |
Evl |
Ena-vasodilator stimulated phosphoprotein |
557 |
0.74 |
| chr3_8512495_8512918 | 4.30 |
Stmn2 |
stathmin-like 2 |
3120 |
0.28 |
| chr8_93812106_93812875 | 4.28 |
Gnao1 |
guanine nucleotide binding protein, alpha O |
1177 |
0.35 |
| chr18_81165961_81166641 | 4.27 |
4930594M17Rik |
RIKEN cDNA 4930594M17 gene |
69765 |
0.09 |
| chr18_31446492_31447667 | 4.26 |
Syt4 |
synaptotagmin IV |
327 |
0.87 |
| chr2_94484217_94484568 | 4.26 |
Api5 |
apoptosis inhibitor 5 |
46256 |
0.12 |
| chr8_22012864_22013642 | 4.26 |
Ccdc70 |
coiled-coil domain containing 70 |
42657 |
0.08 |
| chr9_91360032_91360505 | 4.26 |
Zic4 |
zinc finger protein of the cerebellum 4 |
2145 |
0.17 |
| chr11_46309925_46310526 | 4.25 |
Cyfip2 |
cytoplasmic FMR1 interacting protein 2 |
1995 |
0.28 |
| chr15_68931535_68932379 | 4.25 |
Khdrbs3 |
KH domain containing, RNA binding, signal transduction associated 3 |
1889 |
0.38 |
| chr3_34659833_34662467 | 4.24 |
Gm42693 |
predicted gene 42693 |
3139 |
0.15 |
| chr4_90437640_90438299 | 4.23 |
Gm12635 |
predicted gene 12635 |
14905 |
0.24 |
| chr1_168044748_168045288 | 4.22 |
Gm20711 |
predicted gene 20711 |
40879 |
0.21 |
| chr17_85394123_85395241 | 4.22 |
Rpl31-ps16 |
ribosomal protein L31, pseudogene 16 |
103951 |
0.07 |
| chr2_50902288_50902949 | 4.21 |
Gm13498 |
predicted gene 13498 |
7066 |
0.32 |
| chr3_27060001_27060284 | 4.21 |
Gm7558 |
predicted gene 7558 |
12189 |
0.17 |
| chr7_49907312_49908741 | 4.21 |
Slc6a5 |
solute carrier family 6 (neurotransmitter transporter, glycine), member 5 |
2120 |
0.4 |
| chr10_92161472_92161916 | 4.20 |
Rmst |
rhabdomyosarcoma 2 associated transcript (non-coding RNA) |
1067 |
0.55 |
| chr8_123410787_123412789 | 4.19 |
Tubb3 |
tubulin, beta 3 class III |
198 |
0.84 |
| chrX_166346941_166347276 | 4.19 |
Gpm6b |
glycoprotein m6b |
2266 |
0.36 |
| chr11_24098820_24099493 | 4.19 |
Gm12064 |
predicted gene 12064 |
16249 |
0.13 |
| chr13_83718912_83719403 | 4.18 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
2224 |
0.22 |
| chr3_84955400_84955669 | 4.17 |
Fbxw7 |
F-box and WD-40 domain protein 7 |
3388 |
0.35 |
| chr2_65932868_65933620 | 4.17 |
Csrnp3 |
cysteine-serine-rich nuclear protein 3 |
1379 |
0.47 |
| chr10_87500739_87501897 | 4.17 |
Gm48120 |
predicted gene, 48120 |
6544 |
0.19 |
| chr3_115774354_115774995 | 4.15 |
Gm9889 |
predicted gene 9889 |
59524 |
0.1 |
| chr3_68572046_68573169 | 4.14 |
Schip1 |
schwannomin interacting protein 1 |
362 |
0.89 |
| chr18_59062200_59063436 | 4.11 |
Minar2 |
membrane integral NOTCH2 associated receptor 2 |
307 |
0.94 |
| chr2_152086279_152087493 | 4.11 |
Scrt2 |
scratch family zinc finger 2 |
5357 |
0.15 |
| chr1_61472065_61473256 | 4.10 |
Gm25839 |
predicted gene, 25839 |
2219 |
0.24 |
| chr3_118430889_118431675 | 4.10 |
Gm26871 |
predicted gene, 26871 |
958 |
0.44 |
| chr1_41604694_41605241 | 4.10 |
Gm28634 |
predicted gene 28634 |
75424 |
0.12 |
| chr7_27653599_27654745 | 4.08 |
Ttc9b |
tetratricopeptide repeat domain 9B |
257 |
0.83 |
| chr1_177444257_177446079 | 4.06 |
Zbtb18 |
zinc finger and BTB domain containing 18 |
230 |
0.9 |
| chr12_88725972_88726370 | 4.06 |
Nrxn3 |
neurexin III |
490 |
0.84 |
| chr9_96733821_96734335 | 4.06 |
Zbtb38 |
zinc finger and BTB domain containing 38 |
1308 |
0.4 |
| chr3_34653590_34654523 | 4.03 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
1980 |
0.2 |
| chr1_77505286_77506951 | 4.03 |
Epha4 |
Eph receptor A4 |
8961 |
0.18 |
| chr9_69592532_69592902 | 4.03 |
Gm47203 |
predicted gene, 47203 |
7037 |
0.25 |
| chr2_105668422_105670370 | 4.03 |
Pax6 |
paired box 6 |
461 |
0.65 |
| chr5_139550965_139553757 | 4.01 |
Uncx |
UNC homeobox |
8463 |
0.18 |
| chr8_45507516_45508498 | 3.99 |
Sorbs2 |
sorbin and SH3 domain containing 2 |
89 |
0.97 |
| chr13_112289274_112289896 | 3.99 |
Ankrd55 |
ankyrin repeat domain 55 |
765 |
0.56 |
| chr16_72510590_72511319 | 3.99 |
Robo1 |
roundabout guidance receptor 1 |
52746 |
0.18 |
| chr3_34561815_34562105 | 3.97 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
1568 |
0.33 |
| chr12_29534253_29535510 | 3.97 |
Gm20208 |
predicted gene, 20208 |
10 |
0.8 |
| chr14_65423052_65425451 | 3.96 |
Pnoc |
prepronociceptin |
909 |
0.6 |
| chr14_100374663_100375528 | 3.94 |
Gm26367 |
predicted gene, 26367 |
43388 |
0.15 |
Network of associatons between targets according to the STRING database.

Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.7 | 14.1 | GO:0021912 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) |
| 4.1 | 12.3 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 2.8 | 5.7 | GO:0097477 | lateral motor column neuron migration(GO:0097477) |
| 2.7 | 13.3 | GO:2000481 | positive regulation of cAMP-dependent protein kinase activity(GO:2000481) |
| 2.6 | 10.5 | GO:0051611 | negative regulation of neurotransmitter uptake(GO:0051581) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 2.5 | 17.7 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 2.3 | 28.2 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 2.3 | 11.3 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 2.2 | 11.2 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 2.2 | 8.8 | GO:0033602 | negative regulation of dopamine secretion(GO:0033602) |
| 2.2 | 6.5 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 2.1 | 14.8 | GO:0021830 | interneuron migration from the subpallium to the cortex(GO:0021830) |
| 2.1 | 16.8 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 2.1 | 6.2 | GO:1904742 | regulation of telomeric DNA binding(GO:1904742) |
| 2.0 | 13.9 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 1.9 | 5.6 | GO:0097112 | gamma-aminobutyric acid receptor clustering(GO:0097112) |
| 1.9 | 13.0 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 1.8 | 3.7 | GO:2000809 | positive regulation of synaptic vesicle clustering(GO:2000809) |
| 1.8 | 5.5 | GO:0021553 | olfactory nerve development(GO:0021553) |
| 1.7 | 6.9 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 1.7 | 5.0 | GO:0099558 | maintenance of synapse structure(GO:0099558) |
| 1.6 | 6.3 | GO:0072051 | juxtaglomerular apparatus development(GO:0072051) |
| 1.6 | 4.7 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 1.4 | 5.7 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 1.4 | 7.0 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 1.4 | 13.9 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 1.4 | 2.7 | GO:2000821 | regulation of grooming behavior(GO:2000821) |
| 1.3 | 5.2 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 1.3 | 11.5 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 1.3 | 2.5 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 1.3 | 16.4 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 1.3 | 1.3 | GO:0061642 | chemoattraction of axon(GO:0061642) |
| 1.3 | 7.5 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 1.3 | 13.8 | GO:0034643 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 1.2 | 3.7 | GO:0021780 | oligodendrocyte cell fate specification(GO:0021778) oligodendrocyte cell fate commitment(GO:0021779) glial cell fate specification(GO:0021780) |
| 1.2 | 6.2 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 1.2 | 8.6 | GO:0098870 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 1.2 | 2.4 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 1.2 | 4.7 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 1.2 | 4.7 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 1.2 | 1.2 | GO:2000807 | regulation of synaptic vesicle clustering(GO:2000807) |
| 1.2 | 4.7 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 1.1 | 3.4 | GO:1902897 | regulation of postsynaptic density protein 95 clustering(GO:1902897) |
| 1.1 | 6.9 | GO:0098598 | vocal learning(GO:0042297) imitative learning(GO:0098596) learned vocalization behavior or vocal learning(GO:0098598) |
| 1.1 | 3.4 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 1.1 | 2.2 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 1.1 | 6.5 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 1.1 | 7.5 | GO:0071625 | vocalization behavior(GO:0071625) |
| 1.1 | 3.2 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 1.0 | 7.3 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 1.0 | 6.2 | GO:0021957 | corticospinal tract morphogenesis(GO:0021957) |
| 1.0 | 6.9 | GO:0060134 | prepulse inhibition(GO:0060134) |
| 1.0 | 3.9 | GO:0030091 | protein repair(GO:0030091) |
| 1.0 | 2.9 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 1.0 | 2.9 | GO:0006532 | aspartate biosynthetic process(GO:0006532) |
| 0.9 | 2.8 | GO:1901078 | negative regulation of relaxation of muscle(GO:1901078) |
| 0.9 | 2.8 | GO:1901843 | positive regulation of high voltage-gated calcium channel activity(GO:1901843) |
| 0.9 | 4.7 | GO:0014054 | positive regulation of gamma-aminobutyric acid secretion(GO:0014054) |
| 0.9 | 15.7 | GO:0071475 | cellular hyperosmotic salinity response(GO:0071475) |
| 0.9 | 0.9 | GO:0097475 | motor neuron migration(GO:0097475) spinal cord motor neuron migration(GO:0097476) |
| 0.9 | 2.7 | GO:0021882 | regulation of transcription from RNA polymerase II promoter involved in forebrain neuron fate commitment(GO:0021882) |
| 0.9 | 4.5 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.9 | 0.9 | GO:1904339 | negative regulation of dopaminergic neuron differentiation(GO:1904339) |
| 0.9 | 6.2 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.9 | 4.4 | GO:0048852 | diencephalon morphogenesis(GO:0048852) |
| 0.9 | 4.4 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.9 | 4.4 | GO:0098828 | positive regulation of inhibitory postsynaptic potential(GO:0097151) modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.9 | 3.5 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.9 | 3.5 | GO:2000852 | corticosterone secretion(GO:0035934) regulation of corticosterone secretion(GO:2000852) |
| 0.9 | 6.0 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.8 | 2.5 | GO:0060060 | post-embryonic retina morphogenesis in camera-type eye(GO:0060060) |
| 0.8 | 5.1 | GO:0002091 | negative regulation of receptor internalization(GO:0002091) |
| 0.8 | 2.4 | GO:0042126 | nitrate metabolic process(GO:0042126) |
| 0.8 | 6.4 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.8 | 0.8 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) |
| 0.8 | 2.3 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.8 | 2.3 | GO:0060478 | acrosomal vesicle exocytosis(GO:0060478) |
| 0.8 | 2.3 | GO:0036515 | serotonergic neuron axon guidance(GO:0036515) |
| 0.7 | 2.9 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.7 | 0.7 | GO:1904861 | excitatory synapse assembly(GO:1904861) |
| 0.7 | 1.4 | GO:0072309 | mesenchymal stem cell maintenance involved in metanephric nephron morphogenesis(GO:0072309) |
| 0.7 | 2.2 | GO:0002877 | regulation of acute inflammatory response to non-antigenic stimulus(GO:0002877) |
| 0.7 | 5.0 | GO:0007406 | negative regulation of neuroblast proliferation(GO:0007406) |
| 0.7 | 1.4 | GO:0071314 | cellular response to cocaine(GO:0071314) |
| 0.7 | 0.7 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.7 | 0.7 | GO:0060523 | prostate epithelial cord elongation(GO:0060523) |
| 0.7 | 2.1 | GO:0045794 | negative regulation of cell volume(GO:0045794) |
| 0.7 | 0.7 | GO:1902473 | regulation of protein localization to synapse(GO:1902473) positive regulation of protein localization to synapse(GO:1902474) |
| 0.7 | 2.0 | GO:0035696 | monocyte extravasation(GO:0035696) regulation of monocyte extravasation(GO:2000437) |
| 0.7 | 6.0 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 0.7 | 4.6 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.7 | 1.3 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.7 | 2.6 | GO:1903215 | negative regulation of protein targeting to mitochondrion(GO:1903215) |
| 0.7 | 2.0 | GO:1902071 | regulation of hypoxia-inducible factor-1alpha signaling pathway(GO:1902071) |
| 0.6 | 1.9 | GO:0032100 | positive regulation of response to food(GO:0032097) positive regulation of appetite(GO:0032100) |
| 0.6 | 1.9 | GO:1903223 | positive regulation of oxidative stress-induced neuron death(GO:1903223) |
| 0.6 | 1.9 | GO:0006842 | tricarboxylic acid transport(GO:0006842) citrate transport(GO:0015746) |
| 0.6 | 3.2 | GO:0050915 | sensory perception of sour taste(GO:0050915) |
| 0.6 | 2.5 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.6 | 3.1 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.6 | 1.2 | GO:0021797 | forebrain anterior/posterior pattern specification(GO:0021797) |
| 0.6 | 1.8 | GO:1903690 | negative regulation of wound healing, spreading of epidermal cells(GO:1903690) |
| 0.6 | 1.8 | GO:0050975 | sensory perception of touch(GO:0050975) |
| 0.6 | 2.3 | GO:0021891 | olfactory bulb interneuron development(GO:0021891) |
| 0.6 | 5.7 | GO:0001964 | startle response(GO:0001964) |
| 0.6 | 2.8 | GO:0021707 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.6 | 2.2 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.5 | 3.3 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.5 | 0.5 | GO:0010751 | negative regulation of nitric oxide mediated signal transduction(GO:0010751) |
| 0.5 | 27.3 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.5 | 2.1 | GO:0045113 | regulation of integrin biosynthetic process(GO:0045113) |
| 0.5 | 5.8 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.5 | 2.1 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.5 | 1.1 | GO:0060166 | olfactory pit development(GO:0060166) |
| 0.5 | 0.5 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.5 | 8.4 | GO:0021516 | dorsal spinal cord development(GO:0021516) |
| 0.5 | 1.6 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.5 | 1.0 | GO:0014075 | response to amine(GO:0014075) |
| 0.5 | 0.5 | GO:0010749 | regulation of nitric oxide mediated signal transduction(GO:0010749) |
| 0.5 | 10.5 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.5 | 2.0 | GO:0098904 | regulation of AV node cell action potential(GO:0098904) |
| 0.5 | 5.9 | GO:0001504 | neurotransmitter uptake(GO:0001504) |
| 0.5 | 7.1 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.5 | 2.8 | GO:0003139 | secondary heart field specification(GO:0003139) |
| 0.5 | 2.8 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.5 | 1.4 | GO:1902630 | regulation of membrane hyperpolarization(GO:1902630) |
| 0.5 | 0.9 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.5 | 1.4 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 0.5 | 4.2 | GO:0042428 | serotonin metabolic process(GO:0042428) |
| 0.5 | 5.5 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.5 | 2.3 | GO:0060023 | soft palate development(GO:0060023) |
| 0.4 | 3.1 | GO:0048671 | negative regulation of collateral sprouting(GO:0048671) |
| 0.4 | 1.8 | GO:0072592 | oxygen metabolic process(GO:0072592) |
| 0.4 | 0.9 | GO:0021871 | forebrain regionalization(GO:0021871) |
| 0.4 | 1.8 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 0.4 | 0.9 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.4 | 13.4 | GO:0021766 | hippocampus development(GO:0021766) |
| 0.4 | 4.3 | GO:0033523 | histone H2B ubiquitination(GO:0033523) |
| 0.4 | 2.6 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.4 | 0.9 | GO:0051902 | negative regulation of mitochondrial depolarization(GO:0051902) |
| 0.4 | 2.5 | GO:0007210 | serotonin receptor signaling pathway(GO:0007210) |
| 0.4 | 0.8 | GO:0021631 | optic nerve morphogenesis(GO:0021631) |
| 0.4 | 1.3 | GO:1904252 | negative regulation of bile acid biosynthetic process(GO:0070858) negative regulation of bile acid metabolic process(GO:1904252) |
| 0.4 | 5.0 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.4 | 1.2 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.4 | 1.2 | GO:1903977 | positive regulation of glial cell migration(GO:1903977) |
| 0.4 | 1.6 | GO:0034627 | 'de novo' NAD biosynthetic process(GO:0034627) |
| 0.4 | 0.8 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.4 | 0.7 | GO:0035633 | maintenance of blood-brain barrier(GO:0035633) |
| 0.4 | 28.3 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.4 | 6.8 | GO:0007616 | long-term memory(GO:0007616) |
| 0.4 | 1.4 | GO:0021562 | vestibulocochlear nerve development(GO:0021562) |
| 0.4 | 0.4 | GO:0097154 | GABAergic neuron differentiation(GO:0097154) |
| 0.3 | 1.0 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.3 | 2.1 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.3 | 2.1 | GO:0070050 | neuron cellular homeostasis(GO:0070050) |
| 0.3 | 0.3 | GO:0071677 | positive regulation of mononuclear cell migration(GO:0071677) |
| 0.3 | 1.0 | GO:0089700 | protein kinase D signaling(GO:0089700) |
| 0.3 | 1.0 | GO:2000463 | positive regulation of excitatory postsynaptic potential(GO:2000463) |
| 0.3 | 27.1 | GO:0007612 | learning(GO:0007612) |
| 0.3 | 0.7 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.3 | 0.3 | GO:0048696 | regulation of collateral sprouting in absence of injury(GO:0048696) |
| 0.3 | 0.3 | GO:0003051 | angiotensin-mediated drinking behavior(GO:0003051) |
| 0.3 | 1.3 | GO:0007619 | courtship behavior(GO:0007619) |
| 0.3 | 1.6 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.3 | 0.6 | GO:0032058 | positive regulation of translational initiation in response to stress(GO:0032058) |
| 0.3 | 2.2 | GO:0034776 | response to histamine(GO:0034776) cellular response to histamine(GO:0071420) |
| 0.3 | 1.0 | GO:0035582 | sequestering of BMP in extracellular matrix(GO:0035582) |
| 0.3 | 1.3 | GO:0034047 | regulation of protein phosphatase type 2A activity(GO:0034047) |
| 0.3 | 0.6 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.3 | 1.3 | GO:0006432 | phenylalanyl-tRNA aminoacylation(GO:0006432) |
| 0.3 | 1.3 | GO:0033603 | positive regulation of dopamine secretion(GO:0033603) |
| 0.3 | 0.9 | GO:0070649 | formin-nucleated actin cable assembly(GO:0070649) |
| 0.3 | 0.9 | GO:0051365 | cellular response to potassium ion starvation(GO:0051365) |
| 0.3 | 0.9 | GO:0018242 | protein O-linked glycosylation via serine(GO:0018242) |
| 0.3 | 0.9 | GO:0046013 | regulation of T cell homeostatic proliferation(GO:0046013) |
| 0.3 | 0.3 | GO:0070295 | renal water absorption(GO:0070295) |
| 0.3 | 2.6 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 0.3 | 1.1 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.3 | 7.4 | GO:0021954 | central nervous system neuron development(GO:0021954) |
| 0.3 | 0.3 | GO:0021877 | forebrain neuron fate commitment(GO:0021877) |
| 0.3 | 2.3 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.3 | 0.6 | GO:1902309 | negative regulation of peptidyl-serine dephosphorylation(GO:1902309) |
| 0.3 | 0.8 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 0.3 | 2.5 | GO:0072502 | cellular phosphate ion homeostasis(GO:0030643) cellular trivalent inorganic anion homeostasis(GO:0072502) |
| 0.3 | 2.5 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.3 | 0.6 | GO:0030432 | peristalsis(GO:0030432) |
| 0.3 | 3.8 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.3 | 1.6 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.3 | 0.8 | GO:0010046 | response to mycotoxin(GO:0010046) |
| 0.3 | 0.8 | GO:0034093 | positive regulation of maintenance of sister chromatid cohesion(GO:0034093) positive regulation of maintenance of mitotic sister chromatid cohesion(GO:0034184) |
| 0.3 | 0.3 | GO:1903261 | regulation of serine phosphorylation of STAT3 protein(GO:1903261) |
| 0.3 | 1.3 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.3 | 0.5 | GO:0015817 | histidine transport(GO:0015817) |
| 0.3 | 1.6 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.3 | 0.8 | GO:0015755 | fructose transport(GO:0015755) |
| 0.3 | 0.5 | GO:1903061 | positive regulation of protein lipidation(GO:1903061) |
| 0.3 | 0.5 | GO:0023021 | termination of signal transduction(GO:0023021) |
| 0.3 | 6.5 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.3 | 0.5 | GO:0003331 | regulation of extracellular matrix constituent secretion(GO:0003330) positive regulation of extracellular matrix constituent secretion(GO:0003331) |
| 0.3 | 1.0 | GO:0051919 | positive regulation of fibrinolysis(GO:0051919) |
| 0.3 | 0.8 | GO:0019464 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.3 | 0.5 | GO:0033693 | neurofilament bundle assembly(GO:0033693) |
| 0.3 | 1.0 | GO:0006564 | L-serine biosynthetic process(GO:0006564) |
| 0.3 | 1.0 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.3 | 0.8 | GO:0035609 | C-terminal protein deglutamylation(GO:0035609) |
| 0.2 | 0.5 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.2 | 0.2 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.2 | 3.5 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.2 | 0.7 | GO:0045650 | negative regulation of macrophage differentiation(GO:0045650) |
| 0.2 | 1.0 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.2 | 1.9 | GO:0034982 | mitochondrial protein processing(GO:0034982) |
| 0.2 | 1.0 | GO:0033030 | negative regulation of neutrophil apoptotic process(GO:0033030) |
| 0.2 | 0.7 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 0.2 | 6.1 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.2 | 0.2 | GO:0071608 | macrophage inflammatory protein-1 alpha production(GO:0071608) |
| 0.2 | 1.4 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.2 | 0.7 | GO:0060178 | regulation of exocyst localization(GO:0060178) |
| 0.2 | 0.5 | GO:0033239 | negative regulation of cellular amine metabolic process(GO:0033239) |
| 0.2 | 3.2 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.2 | 0.2 | GO:1903377 | negative regulation of oxidative stress-induced neuron intrinsic apoptotic signaling pathway(GO:1903377) |
| 0.2 | 0.7 | GO:0015803 | branched-chain amino acid transport(GO:0015803) leucine transport(GO:0015820) |
| 0.2 | 1.6 | GO:0021702 | cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.2 | 1.1 | GO:0032252 | secretory granule localization(GO:0032252) |
| 0.2 | 0.7 | GO:2000847 | negative regulation of steroid hormone secretion(GO:2000832) negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.2 | 2.3 | GO:0002523 | leukocyte migration involved in inflammatory response(GO:0002523) |
| 0.2 | 0.9 | GO:0042518 | negative regulation of tyrosine phosphorylation of Stat3 protein(GO:0042518) |
| 0.2 | 0.9 | GO:2000650 | negative regulation of sodium ion transmembrane transporter activity(GO:2000650) |
| 0.2 | 0.7 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.2 | 0.7 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.2 | 0.4 | GO:0061092 | regulation of phospholipid translocation(GO:0061091) positive regulation of phospholipid translocation(GO:0061092) |
| 0.2 | 5.1 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.2 | 1.3 | GO:0009068 | aspartate family amino acid catabolic process(GO:0009068) |
| 0.2 | 0.4 | GO:0060825 | fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060825) regulation of fibroblast growth factor receptor signaling pathway involved in neural plate anterior/posterior pattern formation(GO:2000313) |
| 0.2 | 0.6 | GO:1904217 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) regulation of serine C-palmitoyltransferase activity(GO:1904220) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.2 | 0.9 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.2 | 0.4 | GO:0042713 | sperm ejaculation(GO:0042713) |
| 0.2 | 0.9 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.2 | 1.5 | GO:1900016 | negative regulation of cytokine production involved in inflammatory response(GO:1900016) |
| 0.2 | 1.1 | GO:0071397 | cellular response to cholesterol(GO:0071397) |
| 0.2 | 3.5 | GO:0097352 | autophagosome maturation(GO:0097352) |
| 0.2 | 0.4 | GO:0042891 | antibiotic transport(GO:0042891) |
| 0.2 | 0.2 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.2 | 0.4 | GO:0030908 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.2 | 8.8 | GO:0061178 | regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061178) |
| 0.2 | 0.6 | GO:1902564 | negative regulation of neutrophil activation(GO:1902564) |
| 0.2 | 0.4 | GO:2000310 | regulation of N-methyl-D-aspartate selective glutamate receptor activity(GO:2000310) |
| 0.2 | 0.2 | GO:0051572 | negative regulation of histone H3-K4 methylation(GO:0051572) |
| 0.2 | 0.4 | GO:2000974 | negative regulation of pro-B cell differentiation(GO:2000974) |
| 0.2 | 0.4 | GO:0090135 | actin filament branching(GO:0090135) |
| 0.2 | 0.8 | GO:0006041 | glucosamine metabolic process(GO:0006041) |
| 0.2 | 0.2 | GO:1902308 | regulation of peptidyl-serine dephosphorylation(GO:1902308) |
| 0.2 | 0.4 | GO:0001997 | positive regulation of the force of heart contraction by epinephrine-norepinephrine(GO:0001997) positive regulation of the force of heart contraction by chemical signal(GO:0003099) |
| 0.2 | 2.0 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.2 | 1.9 | GO:0036065 | fucosylation(GO:0036065) |
| 0.2 | 0.6 | GO:0051835 | positive regulation of synapse structural plasticity(GO:0051835) |
| 0.2 | 0.6 | GO:0036481 | intrinsic apoptotic signaling pathway in response to hydrogen peroxide(GO:0036481) |
| 0.2 | 0.4 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.2 | 2.7 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.2 | 0.8 | GO:0045409 | negative regulation of interleukin-6 biosynthetic process(GO:0045409) |
| 0.2 | 0.2 | GO:1901727 | positive regulation of histone deacetylase activity(GO:1901727) |
| 0.2 | 0.8 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.2 | 0.9 | GO:1901026 | ripoptosome assembly(GO:0097343) ripoptosome assembly involved in necroptotic process(GO:1901026) |
| 0.2 | 0.2 | GO:0031547 | brain-derived neurotrophic factor receptor signaling pathway(GO:0031547) |
| 0.2 | 0.6 | GO:0060052 | neurofilament cytoskeleton organization(GO:0060052) |
| 0.2 | 1.1 | GO:0035773 | insulin secretion involved in cellular response to glucose stimulus(GO:0035773) |
| 0.2 | 1.6 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.2 | 0.4 | GO:0046098 | guanine metabolic process(GO:0046098) |
| 0.2 | 0.4 | GO:0035434 | copper ion transmembrane transport(GO:0035434) |
| 0.2 | 0.2 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) |
| 0.2 | 0.2 | GO:0072289 | metanephric nephron tubule formation(GO:0072289) |
| 0.2 | 1.4 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.2 | 0.5 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.2 | 0.5 | GO:0044849 | estrous cycle(GO:0044849) |
| 0.2 | 0.7 | GO:0014744 | positive regulation of muscle adaptation(GO:0014744) |
| 0.2 | 1.2 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.2 | 0.5 | GO:0070885 | negative regulation of calcineurin-NFAT signaling cascade(GO:0070885) |
| 0.2 | 0.2 | GO:0060278 | regulation of ovulation(GO:0060278) |
| 0.2 | 0.7 | GO:0014028 | notochord formation(GO:0014028) |
| 0.2 | 0.2 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.2 | 0.5 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.2 | 0.5 | GO:1900736 | regulation of phospholipase C-activating G-protein coupled receptor signaling pathway(GO:1900736) |
| 0.2 | 0.5 | GO:1903862 | positive regulation of oxidative phosphorylation(GO:1903862) |
| 0.2 | 0.8 | GO:0015816 | glycine transport(GO:0015816) |
| 0.2 | 0.2 | GO:0097051 | establishment of protein localization to endoplasmic reticulum membrane(GO:0097051) |
| 0.2 | 0.3 | GO:0001831 | trophectodermal cellular morphogenesis(GO:0001831) |
| 0.2 | 0.8 | GO:1900028 | negative regulation of ruffle assembly(GO:1900028) |
| 0.2 | 0.6 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.2 | 0.5 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.2 | 0.6 | GO:0032714 | negative regulation of interleukin-5 production(GO:0032714) |
| 0.2 | 0.2 | GO:0061502 | early endosome to recycling endosome transport(GO:0061502) |
| 0.2 | 0.3 | GO:0071895 | odontoblast differentiation(GO:0071895) |
| 0.2 | 0.5 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.2 | 0.6 | GO:0003383 | apical constriction(GO:0003383) |
| 0.2 | 0.8 | GO:0009446 | putrescine biosynthetic process(GO:0009446) |
| 0.2 | 1.5 | GO:0030828 | positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.1 | 0.7 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.1 | 0.4 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 0.3 | GO:0033326 | cerebrospinal fluid secretion(GO:0033326) |
| 0.1 | 1.2 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 0.3 | GO:0014059 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.1 | 0.4 | GO:1900094 | determination of left/right asymmetry in lateral mesoderm(GO:0003140) nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of transcription from RNA polymerase II promoter involved in determination of left/right symmetry(GO:1900094) nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900164) |
| 0.1 | 0.3 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.1 | 1.7 | GO:0048714 | positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.1 | 1.8 | GO:0030497 | fatty acid elongation(GO:0030497) |
| 0.1 | 0.1 | GO:0045163 | clustering of voltage-gated potassium channels(GO:0045163) |
| 0.1 | 0.8 | GO:0070327 | thyroid hormone transport(GO:0070327) |
| 0.1 | 0.3 | GO:0031296 | B cell costimulation(GO:0031296) |
| 0.1 | 0.3 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.1 | 0.4 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.1 | 0.4 | GO:2001013 | epithelial cell proliferation involved in renal tubule morphogenesis(GO:2001013) |
| 0.1 | 0.8 | GO:0032026 | response to magnesium ion(GO:0032026) |
| 0.1 | 0.5 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.1 | 0.1 | GO:0061196 | fungiform papilla development(GO:0061196) |
| 0.1 | 0.1 | GO:1900226 | negative regulation of NLRP3 inflammasome complex assembly(GO:1900226) |
| 0.1 | 0.4 | GO:0044860 | protein localization to plasma membrane raft(GO:0044860) |
| 0.1 | 0.1 | GO:0008065 | establishment of blood-nerve barrier(GO:0008065) |
| 0.1 | 0.4 | GO:1990168 | protein K29-linked deubiquitination(GO:0035523) protein K33-linked deubiquitination(GO:1990168) |
| 0.1 | 0.7 | GO:0002591 | positive regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002591) |
| 0.1 | 1.5 | GO:0010763 | positive regulation of fibroblast migration(GO:0010763) |
| 0.1 | 0.7 | GO:0032095 | regulation of response to food(GO:0032095) |
| 0.1 | 0.1 | GO:2000110 | negative regulation of macrophage apoptotic process(GO:2000110) |
| 0.1 | 1.0 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.1 | 11.5 | GO:0050770 | regulation of axonogenesis(GO:0050770) |
| 0.1 | 0.1 | GO:1902956 | regulation of mitochondrial electron transport, NADH to ubiquinone(GO:1902956) |
| 0.1 | 0.8 | GO:2000672 | negative regulation of motor neuron apoptotic process(GO:2000672) |
| 0.1 | 0.4 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 0.1 | 0.1 | GO:0014056 | acetylcholine secretion, neurotransmission(GO:0014055) regulation of acetylcholine secretion, neurotransmission(GO:0014056) |
| 0.1 | 0.1 | GO:0018243 | protein O-linked glycosylation via threonine(GO:0018243) |
| 0.1 | 0.3 | GO:0060079 | excitatory postsynaptic potential(GO:0060079) |
| 0.1 | 2.9 | GO:0034067 | protein localization to Golgi apparatus(GO:0034067) |
| 0.1 | 7.7 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 0.4 | GO:1903288 | regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) |
| 0.1 | 0.5 | GO:1900363 | regulation of mRNA polyadenylation(GO:1900363) negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.1 | 0.1 | GO:0019064 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.1 | 3.3 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 0.1 | GO:1905005 | regulation of epithelial to mesenchymal transition involved in endocardial cushion formation(GO:1905005) |
| 0.1 | 0.4 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.1 | 0.7 | GO:0035280 | miRNA loading onto RISC involved in gene silencing by miRNA(GO:0035280) |
| 0.1 | 0.1 | GO:0060763 | mammary duct terminal end bud growth(GO:0060763) |
| 0.1 | 0.1 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.1 | 0.2 | GO:0010908 | regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010908) positive regulation of heparan sulfate proteoglycan biosynthetic process(GO:0010909) positive regulation of proteoglycan biosynthetic process(GO:1902730) |
| 0.1 | 6.4 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.1 | 0.5 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.1 | 0.5 | GO:0070837 | dehydroascorbic acid transport(GO:0070837) |
| 0.1 | 0.2 | GO:1903336 | negative regulation of vacuolar transport(GO:1903336) |
| 0.1 | 0.1 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.1 | 0.3 | GO:0006563 | L-serine metabolic process(GO:0006563) |
| 0.1 | 0.1 | GO:0097466 | glycoprotein ERAD pathway(GO:0097466) response to glycoprotein(GO:1904587) |
| 0.1 | 0.1 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.1 | 0.2 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.1 | 0.4 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.1 | 0.3 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 0.2 | GO:0072367 | regulation of lipid transport by regulation of transcription from RNA polymerase II promoter(GO:0072367) |
| 0.1 | 0.4 | GO:0070417 | cellular response to cold(GO:0070417) |
| 0.1 | 0.4 | GO:0032929 | negative regulation of superoxide anion generation(GO:0032929) |
| 0.1 | 0.1 | GO:0090045 | positive regulation of deacetylase activity(GO:0090045) |
| 0.1 | 0.1 | GO:0097195 | pilomotor reflex(GO:0097195) |
| 0.1 | 0.1 | GO:0002752 | cell surface pattern recognition receptor signaling pathway(GO:0002752) |
| 0.1 | 2.4 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.1 | 0.4 | GO:0035247 | peptidyl-arginine methylation, to asymmetrical-dimethyl arginine(GO:0019919) peptidyl-arginine omega-N-methylation(GO:0035247) |
| 0.1 | 0.3 | GO:0045760 | positive regulation of action potential(GO:0045760) |
| 0.1 | 0.7 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.1 | 0.6 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.1 | 0.2 | GO:0061302 | smooth muscle cell-matrix adhesion(GO:0061302) |
| 0.1 | 0.1 | GO:0045200 | establishment or maintenance of neuroblast polarity(GO:0045196) establishment of neuroblast polarity(GO:0045200) |
| 0.1 | 0.3 | GO:0002913 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.1 | 0.9 | GO:0071526 | semaphorin-plexin signaling pathway(GO:0071526) |
| 0.1 | 0.4 | GO:0048934 | peripheral nervous system neuron differentiation(GO:0048934) peripheral nervous system neuron development(GO:0048935) |
| 0.1 | 3.6 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.1 | 0.2 | GO:0071898 | regulation of estrogen receptor binding(GO:0071898) negative regulation of estrogen receptor binding(GO:0071899) |
| 0.1 | 0.3 | GO:0036015 | response to interleukin-3(GO:0036015) cellular response to interleukin-3(GO:0036016) |
| 0.1 | 0.2 | GO:0042048 | olfactory behavior(GO:0042048) |
| 0.1 | 0.3 | GO:0070535 | histone H2A K63-linked ubiquitination(GO:0070535) |
| 0.1 | 0.2 | GO:1902953 | positive regulation of ER to Golgi vesicle-mediated transport(GO:1902953) |
| 0.1 | 0.3 | GO:0016103 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
| 0.1 | 0.3 | GO:0006562 | proline catabolic process(GO:0006562) |
| 0.1 | 0.2 | GO:0061577 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) |
| 0.1 | 0.1 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.1 | 0.4 | GO:0060830 | ciliary receptor clustering involved in smoothened signaling pathway(GO:0060830) |
| 0.1 | 0.2 | GO:0019344 | cysteine biosynthetic process(GO:0019344) |
| 0.1 | 0.2 | GO:0032310 | prostaglandin transport(GO:0015732) prostaglandin secretion(GO:0032310) |
| 0.1 | 0.5 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 0.2 | GO:0009629 | response to gravity(GO:0009629) |
| 0.1 | 0.1 | GO:0002121 | inter-male aggressive behavior(GO:0002121) |
| 0.1 | 0.4 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.1 | 0.4 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.1 | 0.6 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.1 | 0.2 | GO:0061038 | uterus morphogenesis(GO:0061038) |
| 0.1 | 0.1 | GO:0010725 | regulation of primitive erythrocyte differentiation(GO:0010725) |
| 0.1 | 0.1 | GO:0031441 | negative regulation of mRNA 3'-end processing(GO:0031441) |
| 0.1 | 0.2 | GO:0042536 | negative regulation of tumor necrosis factor biosynthetic process(GO:0042536) |
| 0.1 | 0.2 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) |
| 0.1 | 0.3 | GO:0015871 | choline transport(GO:0015871) |
| 0.1 | 0.3 | GO:1901724 | positive regulation of cell proliferation involved in kidney development(GO:1901724) |
| 0.1 | 0.4 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.1 | 0.3 | GO:0050884 | neuromuscular process controlling posture(GO:0050884) |
| 0.1 | 0.2 | GO:0035865 | response to potassium ion(GO:0035864) cellular response to potassium ion(GO:0035865) |
| 0.1 | 0.2 | GO:0001920 | negative regulation of receptor recycling(GO:0001920) |
| 0.1 | 0.2 | GO:0002248 | connective tissue replacement involved in inflammatory response wound healing(GO:0002248) connective tissue replacement(GO:0097709) |
| 0.1 | 0.1 | GO:0071926 | cannabinoid signaling pathway(GO:0038171) endocannabinoid signaling pathway(GO:0071926) |
| 0.1 | 0.2 | GO:0072531 | pyrimidine-containing compound transmembrane transport(GO:0072531) |
| 0.1 | 0.3 | GO:0015879 | carnitine transport(GO:0015879) |
| 0.1 | 0.2 | GO:0001555 | oocyte growth(GO:0001555) |
| 0.1 | 0.4 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.1 | 0.4 | GO:0045631 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.1 | 0.8 | GO:0000038 | very long-chain fatty acid metabolic process(GO:0000038) |
| 0.1 | 0.5 | GO:1902222 | L-phenylalanine metabolic process(GO:0006558) L-phenylalanine catabolic process(GO:0006559) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 0.1 | 0.1 | GO:0015793 | glycerol transport(GO:0015793) cellular response to copper ion(GO:0071280) cellular response to mercury ion(GO:0071288) |
| 0.1 | 0.2 | GO:0045080 | positive regulation of chemokine biosynthetic process(GO:0045080) |
| 0.1 | 0.2 | GO:0050847 | progesterone receptor signaling pathway(GO:0050847) |
| 0.1 | 0.2 | GO:0019287 | isopentenyl diphosphate biosynthetic process(GO:0009240) isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.1 | 0.1 | GO:2000259 | positive regulation of complement activation(GO:0045917) positive regulation of protein activation cascade(GO:2000259) |
| 0.1 | 0.1 | GO:2000304 | positive regulation of sphingolipid biosynthetic process(GO:0090154) positive regulation of ceramide biosynthetic process(GO:2000304) |
| 0.1 | 0.3 | GO:0051918 | negative regulation of fibrinolysis(GO:0051918) |
| 0.1 | 0.4 | GO:0042136 | neurotransmitter biosynthetic process(GO:0042136) |
| 0.1 | 0.2 | GO:0003105 | negative regulation of glomerular filtration(GO:0003105) |
| 0.1 | 0.7 | GO:0046688 | response to copper ion(GO:0046688) |
| 0.1 | 0.4 | GO:0050774 | negative regulation of dendrite morphogenesis(GO:0050774) |
| 0.1 | 0.4 | GO:2000254 | regulation of male germ cell proliferation(GO:2000254) |
| 0.1 | 0.3 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.1 | 0.1 | GO:0045990 | carbon catabolite regulation of transcription(GO:0045990) |
| 0.1 | 0.2 | GO:0097411 | hypoxia-inducible factor-1alpha signaling pathway(GO:0097411) |
| 0.1 | 3.3 | GO:0050808 | synapse organization(GO:0050808) |
| 0.1 | 0.2 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.1 | 0.3 | GO:0046836 | glycolipid transport(GO:0046836) |
| 0.1 | 0.2 | GO:0010966 | regulation of phosphate transport(GO:0010966) |
| 0.1 | 0.7 | GO:0006071 | glycerol metabolic process(GO:0006071) |
| 0.1 | 0.1 | GO:0072393 | microtubule anchoring at microtubule organizing center(GO:0072393) |
| 0.1 | 0.2 | GO:0044828 | negative regulation by host of viral genome replication(GO:0044828) |
| 0.1 | 0.3 | GO:0033159 | negative regulation of protein import into nucleus, translocation(GO:0033159) |
| 0.1 | 0.1 | GO:0060300 | regulation of cytokine activity(GO:0060300) |
| 0.1 | 0.7 | GO:0010971 | positive regulation of G2/M transition of mitotic cell cycle(GO:0010971) |
| 0.1 | 0.6 | GO:1901522 | positive regulation of transcription from RNA polymerase II promoter involved in cellular response to chemical stimulus(GO:1901522) |
| 0.1 | 0.1 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.1 | 0.6 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 0.3 | GO:1903352 | ornithine transport(GO:0015822) L-ornithine transmembrane transport(GO:1903352) |
| 0.1 | 0.1 | GO:0035905 | ascending aorta development(GO:0035905) ascending aorta morphogenesis(GO:0035910) |
| 0.1 | 1.1 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.1 | 0.1 | GO:1900449 | regulation of glutamate receptor signaling pathway(GO:1900449) |
| 0.1 | 0.3 | GO:1902969 | mitotic DNA replication(GO:1902969) |
| 0.1 | 0.8 | GO:0043457 | regulation of cellular respiration(GO:0043457) |
| 0.1 | 0.2 | GO:0072697 | protein localization to cell cortex(GO:0072697) |
| 0.1 | 0.1 | GO:1904528 | regulation of microtubule binding(GO:1904526) positive regulation of microtubule binding(GO:1904528) |
| 0.1 | 0.1 | GO:0030382 | sperm mitochondrion organization(GO:0030382) |
| 0.1 | 0.1 | GO:0002476 | antigen processing and presentation of endogenous peptide antigen via MHC class Ib(GO:0002476) |
| 0.1 | 0.4 | GO:0035428 | hexose transmembrane transport(GO:0035428) |
| 0.1 | 0.1 | GO:0035092 | sperm chromatin condensation(GO:0035092) |
| 0.1 | 0.4 | GO:0007207 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) phospholipase C-activating G-protein coupled acetylcholine receptor signaling pathway(GO:0007207) |
| 0.1 | 0.1 | GO:0002606 | positive regulation of dendritic cell antigen processing and presentation(GO:0002606) |
| 0.1 | 0.2 | GO:0051454 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.1 | 0.1 | GO:0048865 | stem cell fate commitment(GO:0048865) |
| 0.1 | 0.4 | GO:0032094 | response to food(GO:0032094) |
| 0.1 | 0.5 | GO:0030238 | male sex determination(GO:0030238) |
| 0.1 | 0.1 | GO:0046490 | isopentenyl diphosphate metabolic process(GO:0046490) |
| 0.1 | 0.1 | GO:0051799 | negative regulation of hair follicle development(GO:0051799) |
| 0.1 | 0.2 | GO:0039689 | negative stranded viral RNA replication(GO:0039689) multi-organism biosynthetic process(GO:0044034) |
| 0.1 | 0.1 | GO:0060120 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.1 | 0.2 | GO:0032747 | positive regulation of interleukin-23 production(GO:0032747) |
| 0.1 | 0.1 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 3.0 | GO:0007269 | neurotransmitter secretion(GO:0007269) presynaptic process involved in chemical synaptic transmission(GO:0099531) signal release from synapse(GO:0099643) |
| 0.1 | 0.4 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.1 | 0.1 | GO:0060916 | mesenchymal cell proliferation involved in lung development(GO:0060916) |
| 0.1 | 0.1 | GO:0002461 | tolerance induction dependent upon immune response(GO:0002461) |
| 0.1 | 0.1 | GO:0001834 | trophectodermal cell proliferation(GO:0001834) |
| 0.1 | 0.2 | GO:0061511 | centriole elongation(GO:0061511) |
| 0.1 | 0.1 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.1 | 0.1 | GO:1903071 | positive regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903071) |
| 0.1 | 0.2 | GO:0043097 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine nucleoside salvage(GO:0043097) |
| 0.1 | 0.3 | GO:2000270 | negative regulation of fibroblast apoptotic process(GO:2000270) |
| 0.1 | 0.1 | GO:0051385 | response to mineralocorticoid(GO:0051385) |
| 0.1 | 0.3 | GO:0010452 | histone H3-K36 methylation(GO:0010452) |
| 0.1 | 0.2 | GO:0090080 | positive regulation of MAPKKK cascade by fibroblast growth factor receptor signaling pathway(GO:0090080) |
| 0.1 | 0.1 | GO:0046864 | retinol transport(GO:0034633) isoprenoid transport(GO:0046864) terpenoid transport(GO:0046865) |
| 0.1 | 0.2 | GO:0033762 | response to glucagon(GO:0033762) |
| 0.1 | 0.5 | GO:0035563 | positive regulation of chromatin binding(GO:0035563) |
| 0.1 | 0.1 | GO:0007341 | penetration of zona pellucida(GO:0007341) |
| 0.1 | 0.1 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.2 | GO:2000325 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.1 | 0.2 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.1 | 0.1 | GO:0060468 | prevention of polyspermy(GO:0060468) |
| 0.1 | 0.2 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.1 | 0.2 | GO:1903964 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.0 | 0.1 | GO:0051344 | negative regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051344) |
| 0.0 | 0.6 | GO:0051905 | establishment of pigment granule localization(GO:0051905) |
| 0.0 | 0.1 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.0 | 0.1 | GO:0006531 | aspartate metabolic process(GO:0006531) |
| 0.0 | 0.4 | GO:0032305 | positive regulation of icosanoid secretion(GO:0032305) |
| 0.0 | 0.0 | GO:0010897 | negative regulation of triglyceride catabolic process(GO:0010897) |
| 0.0 | 2.3 | GO:0001580 | detection of chemical stimulus involved in sensory perception of bitter taste(GO:0001580) |
| 0.0 | 0.0 | GO:0015819 | lysine transport(GO:0015819) |
| 0.0 | 0.1 | GO:0021794 | thalamus development(GO:0021794) |
| 0.0 | 0.4 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.0 | 0.2 | GO:0048194 | Golgi vesicle budding(GO:0048194) |
| 0.0 | 0.1 | GO:0006216 | cytidine catabolic process(GO:0006216) cytidine deamination(GO:0009972) cytidine metabolic process(GO:0046087) |
| 0.0 | 0.1 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.0 | 0.0 | GO:1903236 | regulation of leukocyte tethering or rolling(GO:1903236) |
| 0.0 | 0.1 | GO:0070268 | cornification(GO:0070268) |
| 0.0 | 0.1 | GO:0070164 | negative regulation of adiponectin secretion(GO:0070164) |
| 0.0 | 0.1 | GO:0097062 | dendritic spine maintenance(GO:0097062) |
| 0.0 | 0.1 | GO:0032661 | regulation of interleukin-18 production(GO:0032661) |
| 0.0 | 0.1 | GO:0030167 | proteoglycan catabolic process(GO:0030167) |
| 0.0 | 0.1 | GO:0032377 | regulation of intracellular lipid transport(GO:0032377) regulation of intracellular sterol transport(GO:0032380) regulation of intracellular cholesterol transport(GO:0032383) |
| 0.0 | 0.2 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.0 | 0.0 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 0.0 | 0.1 | GO:0046075 | dTTP biosynthetic process(GO:0006235) pyrimidine deoxyribonucleoside triphosphate biosynthetic process(GO:0009212) dTTP metabolic process(GO:0046075) |
| 0.0 | 0.2 | GO:0018101 | protein citrullination(GO:0018101) |
| 0.0 | 0.1 | GO:0090209 | negative regulation of triglyceride metabolic process(GO:0090209) |
| 0.0 | 0.1 | GO:0097210 | response to gonadotropin-releasing hormone(GO:0097210) cellular response to gonadotropin-releasing hormone(GO:0097211) |
| 0.0 | 0.1 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.0 | 0.3 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.0 | 0.1 | GO:0071873 | response to norepinephrine(GO:0071873) |
| 0.0 | 0.3 | GO:0033033 | negative regulation of myeloid cell apoptotic process(GO:0033033) |
| 0.0 | 0.0 | GO:0007215 | glutamate receptor signaling pathway(GO:0007215) |
| 0.0 | 0.1 | GO:0060295 | regulation of cilium movement involved in cell motility(GO:0060295) regulation of cilium beat frequency involved in ciliary motility(GO:0060296) regulation of cilium-dependent cell motility(GO:1902019) |
| 0.0 | 0.0 | GO:1901725 | regulation of histone deacetylase activity(GO:1901725) |
| 0.0 | 0.0 | GO:0015840 | urea transport(GO:0015840) |
| 0.0 | 0.1 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.0 | 0.0 | GO:2000909 | regulation of cholesterol import(GO:0060620) regulation of sterol import(GO:2000909) |
| 0.0 | 0.1 | GO:0060339 | negative regulation of type I interferon-mediated signaling pathway(GO:0060339) |
| 0.0 | 0.3 | GO:0046040 | IMP metabolic process(GO:0046040) |
| 0.0 | 0.5 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.0 | GO:2001025 | positive regulation of response to drug(GO:2001025) |
| 0.0 | 0.1 | GO:0060750 | epithelial cell proliferation involved in mammary gland duct elongation(GO:0060750) branch elongation involved in mammary gland duct branching(GO:0060751) |
| 0.0 | 1.3 | GO:0070098 | chemokine-mediated signaling pathway(GO:0070098) |
| 0.0 | 0.2 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.0 | 0.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.0 | 0.1 | GO:0048675 | axon extension(GO:0048675) |
| 0.0 | 0.1 | GO:0009298 | GDP-mannose biosynthetic process(GO:0009298) |
| 0.0 | 0.1 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.0 | 0.1 | GO:0003356 | regulation of cilium beat frequency(GO:0003356) |
| 0.0 | 0.0 | GO:0016237 | lysosomal microautophagy(GO:0016237) piecemeal microautophagy of nucleus(GO:0034727) late nucleophagy(GO:0044805) |
| 0.0 | 0.3 | GO:0033700 | phospholipid efflux(GO:0033700) |
| 0.0 | 0.0 | GO:0032227 | negative regulation of synaptic transmission, dopaminergic(GO:0032227) |
| 0.0 | 0.3 | GO:0097484 | dendrite extension(GO:0097484) |
| 0.0 | 0.2 | GO:0016048 | detection of temperature stimulus(GO:0016048) |
| 0.0 | 0.0 | GO:0031587 | positive regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0031587) |
| 0.0 | 0.0 | GO:1900425 | negative regulation of defense response to bacterium(GO:1900425) |
| 0.0 | 0.0 | GO:0002386 | immune response in mucosal-associated lymphoid tissue(GO:0002386) |
| 0.0 | 0.2 | GO:0098734 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.0 | 0.2 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.2 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.0 | 0.1 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.0 | 0.1 | GO:1901874 | regulation of post-translational protein modification(GO:1901873) negative regulation of post-translational protein modification(GO:1901874) |
| 0.0 | 0.0 | GO:0043396 | corticotropin-releasing hormone secretion(GO:0043396) regulation of corticotropin-releasing hormone secretion(GO:0043397) |
| 0.0 | 0.1 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.0 | 0.1 | GO:2000009 | negative regulation of protein localization to cell surface(GO:2000009) |
| 0.0 | 0.0 | GO:0017085 | response to insecticide(GO:0017085) |
| 0.0 | 0.3 | GO:0006182 | cGMP biosynthetic process(GO:0006182) |
| 0.0 | 0.1 | GO:0035984 | response to trichostatin A(GO:0035983) cellular response to trichostatin A(GO:0035984) |
| 0.0 | 0.2 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.0 | 0.1 | GO:0000711 | meiotic DNA repair synthesis(GO:0000711) |
| 0.0 | 0.1 | GO:0010989 | negative regulation of low-density lipoprotein particle clearance(GO:0010989) |
| 0.0 | 0.0 | GO:0006701 | progesterone biosynthetic process(GO:0006701) |
| 0.0 | 0.1 | GO:0060831 | smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:0060831) |
| 0.0 | 0.1 | GO:0019276 | UDP-N-acetylgalactosamine metabolic process(GO:0019276) |
| 0.0 | 0.2 | GO:0042438 | melanin biosynthetic process(GO:0042438) |
| 0.0 | 0.7 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 0.3 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.1 | GO:0002727 | natural killer cell cytokine production(GO:0002370) regulation of natural killer cell cytokine production(GO:0002727) |
| 0.0 | 0.0 | GO:0034442 | regulation of lipoprotein oxidation(GO:0034442) negative regulation of lipoprotein oxidation(GO:0034443) |
| 0.0 | 0.0 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.0 | 0.1 | GO:0048261 | negative regulation of receptor-mediated endocytosis(GO:0048261) |
| 0.0 | 0.1 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.0 | 0.0 | GO:1901679 | nucleotide transmembrane transport(GO:1901679) |
| 0.0 | 0.5 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.0 | GO:0018125 | peptidyl-cysteine methylation(GO:0018125) |
| 0.0 | 0.1 | GO:0030007 | cellular potassium ion homeostasis(GO:0030007) |
| 0.0 | 0.1 | GO:0002924 | negative regulation of humoral immune response mediated by circulating immunoglobulin(GO:0002924) |
| 0.0 | 0.0 | GO:1900619 | acetylcholine metabolic process(GO:0008291) acetate ester metabolic process(GO:1900619) |
| 0.0 | 0.1 | GO:0002426 | immunoglobulin production in mucosal tissue(GO:0002426) |
| 0.0 | 0.0 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.0 | 0.1 | GO:0031049 | programmed DNA elimination(GO:0031049) chromosome breakage(GO:0031052) |
| 0.0 | 0.3 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.0 | GO:2001170 | negative regulation of ATP biosynthetic process(GO:2001170) |
| 0.0 | 0.1 | GO:0060736 | prostate gland growth(GO:0060736) |
| 0.0 | 0.1 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.1 | GO:1902809 | regulation of skeletal muscle fiber differentiation(GO:1902809) |
| 0.0 | 0.0 | GO:0070947 | neutrophil mediated killing of fungus(GO:0070947) |
| 0.0 | 0.0 | GO:0090383 | phagosome acidification(GO:0090383) |
| 0.0 | 0.0 | GO:0051970 | negative regulation of transmission of nerve impulse(GO:0051970) |
| 0.0 | 0.0 | GO:0007066 | female meiosis sister chromatid cohesion(GO:0007066) |
| 0.0 | 0.1 | GO:0001771 | immunological synapse formation(GO:0001771) |
| 0.0 | 0.0 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.0 | 0.1 | GO:0007603 | phototransduction, visible light(GO:0007603) |
| 0.0 | 0.1 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 0.0 | 0.1 | GO:0070920 | regulation of production of small RNA involved in gene silencing by RNA(GO:0070920) |
| 0.0 | 0.1 | GO:0009052 | pentose-phosphate shunt, non-oxidative branch(GO:0009052) |
| 0.0 | 0.1 | GO:0007135 | meiosis II(GO:0007135) |
| 0.0 | 0.1 | GO:0006477 | protein sulfation(GO:0006477) |
| 0.0 | 0.0 | GO:0044351 | macropinocytosis(GO:0044351) |
| 0.0 | 0.0 | GO:1904970 | brush border assembly(GO:1904970) |
| 0.0 | 0.1 | GO:0032826 | natural killer cell differentiation involved in immune response(GO:0002325) negative regulation of natural killer cell differentiation(GO:0032824) regulation of natural killer cell differentiation involved in immune response(GO:0032826) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 0.0 | 0.1 | GO:0002765 | immune response-inhibiting signal transduction(GO:0002765) |
| 0.0 | 0.4 | GO:0008333 | endosome to lysosome transport(GO:0008333) |
| 0.0 | 0.1 | GO:0051105 | regulation of DNA ligation(GO:0051105) positive regulation of DNA ligation(GO:0051106) |
| 0.0 | 0.3 | GO:0050650 | chondroitin sulfate proteoglycan biosynthetic process(GO:0050650) |
| 0.0 | 0.1 | GO:0034085 | establishment of sister chromatid cohesion(GO:0034085) cohesin loading(GO:0071921) regulation of cohesin loading(GO:0071922) |
| 0.0 | 0.0 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.0 | 0.1 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.0 | GO:0036257 | multivesicular body organization(GO:0036257) |
| 0.0 | 0.1 | GO:0032485 | regulation of Ral protein signal transduction(GO:0032485) |
| 0.0 | 0.0 | GO:0042940 | D-amino acid transport(GO:0042940) |
| 0.0 | 0.0 | GO:0060128 | corticotropin hormone secreting cell differentiation(GO:0060128) |
| 0.0 | 0.1 | GO:2000676 | positive regulation of type B pancreatic cell apoptotic process(GO:2000676) |
| 0.0 | 0.1 | GO:0017183 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.0 | 0.1 | GO:0032202 | telomere assembly(GO:0032202) |
| 0.0 | 0.1 | GO:0031115 | negative regulation of microtubule polymerization(GO:0031115) |
| 0.0 | 0.0 | GO:1901896 | positive regulation of calcium-transporting ATPase activity(GO:1901896) |
| 0.0 | 0.0 | GO:0046340 | diacylglycerol catabolic process(GO:0046340) |
| 0.0 | 0.7 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.1 | GO:0060081 | membrane hyperpolarization(GO:0060081) |
| 0.0 | 0.0 | GO:0002579 | positive regulation of antigen processing and presentation(GO:0002579) |
| 0.0 | 0.1 | GO:1902572 | regulation of serine-type endopeptidase activity(GO:1900003) negative regulation of serine-type endopeptidase activity(GO:1900004) regulation of serine-type peptidase activity(GO:1902571) negative regulation of serine-type peptidase activity(GO:1902572) |
| 0.0 | 0.0 | GO:0001927 | exocyst assembly(GO:0001927) |
| 0.0 | 0.1 | GO:0070197 | meiotic telomere tethering at nuclear periphery(GO:0044821) meiotic attachment of telomere to nuclear envelope(GO:0070197) chromosome attachment to the nuclear envelope(GO:0097240) |
| 0.0 | 0.0 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.0 | 0.0 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.0 | 0.1 | GO:0042737 | drug catabolic process(GO:0042737) |
| 0.0 | 0.0 | GO:0019042 | viral latency(GO:0019042) |
| 0.0 | 0.0 | GO:0000320 | re-entry into mitotic cell cycle(GO:0000320) |
| 0.0 | 0.0 | GO:0035524 | proline transmembrane transport(GO:0035524) |
| 0.0 | 0.1 | GO:0031498 | chromatin disassembly(GO:0031498) |
| 0.0 | 0.0 | GO:0021819 | layer formation in cerebral cortex(GO:0021819) |
| 0.0 | 0.0 | GO:0006083 | acetate metabolic process(GO:0006083) |
| 0.0 | 0.1 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.0 | 0.0 | GO:1900157 | regulation of bone mineralization involved in bone maturation(GO:1900157) |
| 0.0 | 0.1 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 0.0 | GO:1900368 | regulation of RNA interference(GO:1900368) |
| 0.0 | 0.1 | GO:0090160 | Golgi to lysosome transport(GO:0090160) |
| 0.0 | 0.1 | GO:0006002 | fructose 6-phosphate metabolic process(GO:0006002) |
| 0.0 | 0.1 | GO:0033313 | meiotic cell cycle checkpoint(GO:0033313) |
| 0.0 | 0.1 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.0 | 0.0 | GO:0050717 | positive regulation of interleukin-1 alpha secretion(GO:0050717) |
| 0.0 | 0.1 | GO:0052697 | xenobiotic glucuronidation(GO:0052697) |
| 0.0 | 0.0 | GO:0014724 | regulation of twitch skeletal muscle contraction(GO:0014724) |
| 0.0 | 0.0 | GO:0048382 | mesendoderm development(GO:0048382) |
| 0.0 | 0.0 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.0 | 0.0 | GO:0071294 | cellular response to zinc ion(GO:0071294) |
| 0.0 | 0.0 | GO:0006449 | regulation of translational termination(GO:0006449) |
| 0.0 | 0.0 | GO:0046051 | UTP metabolic process(GO:0046051) |
| 0.0 | 0.1 | GO:0090084 | negative regulation of inclusion body assembly(GO:0090084) |
| 0.0 | 0.2 | GO:0043631 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.0 | GO:0051654 | establishment of mitochondrion localization(GO:0051654) |
| 0.0 | 0.1 | GO:0006307 | DNA dealkylation involved in DNA repair(GO:0006307) |
| 0.0 | 0.0 | GO:0002689 | negative regulation of leukocyte chemotaxis(GO:0002689) |
| 0.0 | 0.0 | GO:0034370 | triglyceride-rich lipoprotein particle remodeling(GO:0034370) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 13.4 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 1.6 | 1.6 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 1.4 | 14.0 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 1.3 | 4.0 | GO:0097451 | glial limiting end-foot(GO:0097451) |
| 1.2 | 4.9 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 1.2 | 3.7 | GO:0030981 | cortical microtubule cytoskeleton(GO:0030981) |
| 1.2 | 22.7 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 1.1 | 5.4 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 1.1 | 3.2 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 1.0 | 52.7 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.9 | 6.2 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.9 | 12.1 | GO:0043196 | varicosity(GO:0043196) |
| 0.8 | 5.0 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.8 | 2.4 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.8 | 7.5 | GO:0030673 | axolemma(GO:0030673) |
| 0.7 | 2.1 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.7 | 8.2 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.7 | 3.4 | GO:0097433 | dense body(GO:0097433) |
| 0.7 | 4.7 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.7 | 4.6 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.7 | 9.8 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.6 | 7.9 | GO:0031045 | dense core granule(GO:0031045) |
| 0.6 | 14.5 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.6 | 2.4 | GO:0002142 | stereocilia ankle link complex(GO:0002142) |
| 0.6 | 2.9 | GO:0005883 | neurofilament(GO:0005883) |
| 0.6 | 4.0 | GO:0097449 | astrocyte projection(GO:0097449) |
| 0.5 | 6.4 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.5 | 4.7 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.5 | 4.5 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.5 | 12.5 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.5 | 13.9 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.5 | 19.3 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.5 | 71.9 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.5 | 6.6 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.4 | 3.0 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.4 | 2.5 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.4 | 1.5 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.4 | 7.3 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.4 | 2.1 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.3 | 1.0 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.3 | 0.3 | GO:0008328 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.3 | 31.6 | GO:0030427 | site of polarized growth(GO:0030427) |
| 0.3 | 1.0 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.3 | 0.6 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.3 | 3.5 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.3 | 4.1 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.3 | 0.6 | GO:0097169 | AIM2 inflammasome complex(GO:0097169) |
| 0.3 | 23.4 | GO:0043204 | perikaryon(GO:0043204) |
| 0.3 | 7.3 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.3 | 2.3 | GO:0016600 | flotillin complex(GO:0016600) |
| 0.3 | 12.2 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.3 | 0.5 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.2 | 1.4 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.2 | 0.4 | GO:0031088 | platelet dense granule membrane(GO:0031088) |
| 0.2 | 1.8 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.2 | 1.6 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.2 | 0.9 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.2 | 1.1 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.2 | 0.4 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.2 | 1.8 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.2 | 0.2 | GO:0070033 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 0.2 | 3.6 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.2 | 0.5 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 0.2 | 0.2 | GO:0032010 | phagolysosome(GO:0032010) |
| 0.2 | 2.2 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.2 | 0.6 | GO:0014731 | spectrin-associated cytoskeleton(GO:0014731) |
| 0.2 | 2.5 | GO:0090545 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.2 | 0.9 | GO:0036056 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.2 | 0.5 | GO:0097427 | microtubule bundle(GO:0097427) |
| 0.2 | 0.6 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 0.7 | GO:0097440 | apical dendrite(GO:0097440) |
| 0.1 | 0.4 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.1 | 0.4 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.1 | 0.1 | GO:0017071 | intracellular cyclic nucleotide activated cation channel complex(GO:0017071) |
| 0.1 | 0.4 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.1 | 0.3 | GO:0033648 | host intracellular organelle(GO:0033647) host intracellular membrane-bounded organelle(GO:0033648) |
| 0.1 | 17.8 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 0.2 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.1 | 0.4 | GO:0020018 | ciliary pocket(GO:0020016) ciliary pocket membrane(GO:0020018) |
| 0.1 | 1.1 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.1 | 1.7 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.1 | 6.5 | GO:0043679 | axon terminus(GO:0043679) |
| 0.1 | 0.4 | GO:0045293 | mRNA editing complex(GO:0045293) |
| 0.1 | 0.8 | GO:1904115 | axon cytoplasm(GO:1904115) |
| 0.1 | 0.9 | GO:0042582 | primary lysosome(GO:0005766) azurophil granule(GO:0042582) |
| 0.1 | 0.1 | GO:0042827 | platelet dense granule(GO:0042827) |
| 0.1 | 0.1 | GO:0032280 | symmetric synapse(GO:0032280) |
| 0.1 | 0.3 | GO:1990075 | periciliary membrane compartment(GO:1990075) |
| 0.1 | 0.3 | GO:0008275 | gamma-tubulin small complex(GO:0008275) |
| 0.1 | 0.3 | GO:0044322 | endoplasmic reticulum quality control compartment(GO:0044322) |
| 0.1 | 0.3 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 0.1 | 3.8 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 0.3 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.1 | 0.2 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.1 | 0.1 | GO:0044393 | microspike(GO:0044393) |
| 0.1 | 0.7 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.1 | 0.6 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.1 | 0.2 | GO:0034673 | inhibin-betaglycan-ActRII complex(GO:0034673) |
| 0.1 | 3.8 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.1 | 0.1 | GO:0043219 | lateral loop(GO:0043219) |
| 0.1 | 3.8 | GO:0001750 | photoreceptor outer segment(GO:0001750) |
| 0.1 | 0.9 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 0.6 | GO:0032045 | guanyl-nucleotide exchange factor complex(GO:0032045) |
| 0.1 | 0.2 | GO:0097629 | extrinsic component of omegasome membrane(GO:0097629) |
| 0.1 | 0.1 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.1 | 0.1 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.1 | 0.5 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.1 | 0.4 | GO:0001931 | uropod(GO:0001931) cell trailing edge(GO:0031254) |
| 0.1 | 14.7 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.1 | 0.4 | GO:0000801 | central element(GO:0000801) |
| 0.1 | 0.2 | GO:1990716 | axonemal central apparatus(GO:1990716) |
| 0.1 | 0.6 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.1 | 0.8 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.1 | 0.1 | GO:0097425 | smooth endoplasmic reticulum membrane(GO:0030868) smooth endoplasmic reticulum part(GO:0097425) |
| 0.1 | 0.3 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.1 | 0.2 | GO:0031465 | Cul4B-RING E3 ubiquitin ligase complex(GO:0031465) |
| 0.1 | 1.4 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 1.2 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 0.4 | GO:0031332 | RISC complex(GO:0016442) RNAi effector complex(GO:0031332) |
| 0.0 | 0.3 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 0.2 | GO:0070695 | FHF complex(GO:0070695) |
| 0.0 | 0.3 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) |
| 0.0 | 0.7 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 0.2 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 0.0 | 0.1 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.0 | 0.3 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.0 | 0.4 | GO:0046581 | intercellular canaliculus(GO:0046581) |
| 0.0 | 0.1 | GO:0030130 | clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.4 | GO:1990023 | mitotic spindle midzone(GO:1990023) |
| 0.0 | 0.1 | GO:0071203 | WASH complex(GO:0071203) |
| 0.0 | 0.3 | GO:0098793 | presynapse(GO:0098793) |
| 0.0 | 0.1 | GO:0043293 | apoptosome(GO:0043293) |
| 0.0 | 0.4 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 1.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.3 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 0.3 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 0.1 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.0 | 0.2 | GO:0071547 | piP-body(GO:0071547) |
| 0.0 | 0.1 | GO:0000811 | GINS complex(GO:0000811) |
| 0.0 | 0.1 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.3 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.2 | GO:0042824 | MHC class I peptide loading complex(GO:0042824) |
| 0.0 | 0.2 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.0 | 0.0 | GO:0097386 | glial cell projection(GO:0097386) |
| 0.0 | 1.1 | GO:0045171 | intercellular bridge(GO:0045171) |
| 0.0 | 0.2 | GO:0030061 | mitochondrial crista(GO:0030061) |
| 0.0 | 0.1 | GO:0071942 | XPC complex(GO:0071942) |
| 0.0 | 0.1 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 0.1 | GO:0042581 | specific granule(GO:0042581) |
| 0.0 | 0.3 | GO:0034385 | very-low-density lipoprotein particle(GO:0034361) triglyceride-rich lipoprotein particle(GO:0034385) |
| 0.0 | 0.1 | GO:0005663 | DNA replication factor C complex(GO:0005663) |
| 0.0 | 0.4 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 0.1 | GO:0044194 | cytolytic granule(GO:0044194) |
| 0.0 | 0.0 | GO:0072558 | NLRP1 inflammasome complex(GO:0072558) |
| 0.0 | 0.1 | GO:0002177 | manchette(GO:0002177) |
| 0.0 | 0.1 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.0 | 0.1 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 0.2 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.1 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.0 | 0.1 | GO:0005945 | 6-phosphofructokinase complex(GO:0005945) |
| 0.0 | 0.1 | GO:0098576 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) lumenal side of membrane(GO:0098576) |
| 0.0 | 0.1 | GO:0005796 | Golgi lumen(GO:0005796) |
| 0.0 | 0.0 | GO:0044614 | nuclear pore cytoplasmic filaments(GO:0044614) |
| 0.0 | 0.0 | GO:0005745 | m-AAA complex(GO:0005745) |
| 0.0 | 0.8 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 0.0 | GO:0031094 | platelet dense tubular network(GO:0031094) |
| 0.0 | 0.1 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.0 | 0.1 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.0 | 0.1 | GO:0097209 | epidermal lamellar body(GO:0097209) |
| 0.0 | 0.0 | GO:0005767 | secondary lysosome(GO:0005767) |
| 0.0 | 0.1 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.6 | 22.8 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 4.2 | 8.5 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 3.3 | 9.8 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 3.3 | 16.3 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 2.8 | 8.4 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 2.4 | 12.1 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 2.4 | 7.2 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 2.1 | 18.9 | GO:0001091 | RNA polymerase II basal transcription factor binding(GO:0001091) |
| 2.1 | 14.7 | GO:0003680 | AT DNA binding(GO:0003680) |
| 2.0 | 6.1 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 1.7 | 10.0 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 1.6 | 9.6 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 1.1 | 4.3 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 1.0 | 7.3 | GO:0043495 | protein anchor(GO:0043495) |
| 1.0 | 2.9 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 1.0 | 4.8 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 1.0 | 2.9 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 0.9 | 3.7 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.9 | 7.4 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 0.9 | 4.6 | GO:0015185 | gamma-aminobutyric acid transmembrane transporter activity(GO:0015185) |
| 0.9 | 10.7 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) |
| 0.8 | 9.3 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.8 | 6.5 | GO:0098988 | G-protein coupled glutamate receptor activity(GO:0098988) |
| 0.8 | 3.2 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) glycine:sodium symporter activity(GO:0015375) |
| 0.8 | 6.1 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.7 | 4.3 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.7 | 3.4 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.7 | 2.0 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.7 | 3.4 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.6 | 0.6 | GO:0005283 | sodium:amino acid symporter activity(GO:0005283) |
| 0.6 | 1.9 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.6 | 17.2 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.6 | 1.9 | GO:0015142 | citrate transmembrane transporter activity(GO:0015137) tricarboxylic acid transmembrane transporter activity(GO:0015142) |
| 0.6 | 1.8 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.6 | 1.7 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.6 | 13.5 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.6 | 7.9 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) serotonin receptor activity(GO:0099589) |
| 0.6 | 11.2 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.5 | 3.8 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.5 | 2.2 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.5 | 2.7 | GO:0048273 | mitogen-activated protein kinase p38 binding(GO:0048273) |
| 0.5 | 2.6 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.5 | 1.5 | GO:0008066 | glutamate receptor activity(GO:0008066) |
| 0.5 | 2.5 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.5 | 1.4 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.5 | 2.4 | GO:0003958 | NADPH-hemoprotein reductase activity(GO:0003958) |
| 0.5 | 2.4 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.5 | 5.1 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.5 | 1.9 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.5 | 3.2 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.5 | 1.8 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.5 | 1.4 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.4 | 0.9 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.4 | 1.8 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.4 | 4.0 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.4 | 2.2 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.4 | 8.6 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.4 | 1.7 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.4 | 10.4 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.4 | 1.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.4 | 7.1 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.4 | 1.2 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.4 | 10.1 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.4 | 0.8 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.4 | 1.6 | GO:0008486 | diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) |
| 0.4 | 4.3 | GO:0005522 | profilin binding(GO:0005522) |
| 0.4 | 1.9 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.4 | 2.3 | GO:0005225 | volume-sensitive anion channel activity(GO:0005225) |
| 0.4 | 1.2 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.4 | 2.7 | GO:0004952 | dopamine neurotransmitter receptor activity(GO:0004952) |
| 0.4 | 13.6 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.4 | 1.9 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.4 | 3.0 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.4 | 4.7 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.4 | 11.9 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.3 | 1.0 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.3 | 5.4 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.3 | 1.3 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.3 | 6.0 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.3 | 3.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 1.6 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.3 | 1.0 | GO:0004980 | melanocyte-stimulating hormone receptor activity(GO:0004980) |
| 0.3 | 1.6 | GO:0015204 | urea transmembrane transporter activity(GO:0015204) |
| 0.3 | 0.6 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 0.3 | 5.0 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.3 | 1.2 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.3 | 1.2 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.3 | 2.4 | GO:0103116 | alpha-D-galactofuranose transporter activity(GO:0103116) |
| 0.3 | 1.5 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.3 | 0.6 | GO:0043125 | ErbB-3 class receptor binding(GO:0043125) |
| 0.3 | 0.3 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.3 | 1.4 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.3 | 1.3 | GO:0004826 | phenylalanine-tRNA ligase activity(GO:0004826) |
| 0.3 | 3.6 | GO:0022841 | potassium ion leak channel activity(GO:0022841) |
| 0.3 | 1.3 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.3 | 6.0 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.2 | 1.9 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.2 | 3.8 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.2 | 1.7 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.2 | 1.4 | GO:0016775 | phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.2 | 2.2 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.2 | 0.7 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.2 | 0.9 | GO:0008934 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.2 | 4.1 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.2 | 0.2 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.2 | 2.6 | GO:0004000 | adenosine deaminase activity(GO:0004000) |
| 0.2 | 0.6 | GO:0004920 | interleukin-10 receptor activity(GO:0004920) |
| 0.2 | 0.4 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.2 | 0.8 | GO:0031694 | alpha-2A adrenergic receptor binding(GO:0031694) |
| 0.2 | 1.8 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.2 | 0.6 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.2 | 0.6 | GO:0008184 | glycogen phosphorylase activity(GO:0008184) |
| 0.2 | 0.6 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.2 | 1.3 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.2 | 2.3 | GO:0031005 | filamin binding(GO:0031005) |
| 0.2 | 0.2 | GO:0072542 | protein phosphatase activator activity(GO:0072542) |
| 0.2 | 0.9 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.2 | 1.5 | GO:0050544 | arachidonic acid binding(GO:0050544) |
| 0.2 | 2.2 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.2 | 2.2 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.2 | 0.4 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 0.2 | 1.8 | GO:0042301 | phosphate ion binding(GO:0042301) |
| 0.2 | 0.7 | GO:0033192 | calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.2 | 2.4 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.2 | 1.0 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.2 | 0.5 | GO:0005001 | transmembrane receptor protein tyrosine phosphatase activity(GO:0005001) |
| 0.2 | 0.5 | GO:0005290 | L-histidine transmembrane transporter activity(GO:0005290) |
| 0.2 | 0.7 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.2 | 1.3 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.2 | 1.2 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.2 | 3.1 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.2 | 0.5 | GO:0031493 | nucleosomal histone binding(GO:0031493) |
| 0.2 | 1.0 | GO:0015198 | oligopeptide transporter activity(GO:0015198) |
| 0.2 | 1.0 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.2 | 0.5 | GO:0019961 | interferon binding(GO:0019961) |
| 0.2 | 3.7 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.2 | 0.5 | GO:0004515 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.2 | 0.5 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.2 | 0.3 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.2 | 3.5 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.2 | 0.5 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.2 | 0.5 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.2 | 0.3 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.2 | 5.4 | GO:0018727 | N-acetylglucosamine 6-O-sulfotransferase activity(GO:0001517) heparan sulfate 2-O-sulfotransferase activity(GO:0004394) HNK-1 sulfotransferase activity(GO:0016232) heparan sulfate 6-O-sulfotransferase activity(GO:0017095) trans-9R,10R-dihydrodiolphenanthrene sulfotransferase activity(GO:0018721) 1-phenanthrol sulfotransferase activity(GO:0018722) 3-phenanthrol sulfotransferase activity(GO:0018723) 4-phenanthrol sulfotransferase activity(GO:0018724) trans-3,4-dihydrodiolphenanthrene sulfotransferase activity(GO:0018725) 9-phenanthrol sulfotransferase activity(GO:0018726) 2-phenanthrol sulfotransferase activity(GO:0018727) phenanthrol sulfotransferase activity(GO:0019111) 1-hydroxypyrene sulfotransferase activity(GO:0034930) proteoglycan sulfotransferase activity(GO:0050698) cholesterol sulfotransferase activity(GO:0051922) hydroxyjasmonate sulfotransferase activity(GO:0080131) |
| 0.2 | 0.5 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.2 | 0.5 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.2 | 0.6 | GO:0070878 | primary miRNA binding(GO:0070878) |
| 0.2 | 0.9 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 1.2 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.1 | 0.3 | GO:1990829 | C-rich single-stranded DNA binding(GO:1990829) |
| 0.1 | 1.8 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.1 | 0.6 | GO:0001602 | pancreatic polypeptide receptor activity(GO:0001602) |
| 0.1 | 0.1 | GO:0001601 | peptide YY receptor activity(GO:0001601) |
| 0.1 | 1.6 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.1 | 0.4 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.1 | 3.9 | GO:0071855 | neuropeptide receptor binding(GO:0071855) |
| 0.1 | 0.6 | GO:0019145 | aminobutyraldehyde dehydrogenase activity(GO:0019145) 4-trimethylammoniobutyraldehyde dehydrogenase activity(GO:0047105) |
| 0.1 | 0.4 | GO:0005119 | smoothened binding(GO:0005119) |
| 0.1 | 0.5 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.1 | 2.9 | GO:0022839 | ion gated channel activity(GO:0022839) |
| 0.1 | 1.9 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.1 | 3.3 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 0.3 | GO:0048531 | beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.1 | 0.5 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.1 | 0.4 | GO:0051011 | microtubule minus-end binding(GO:0051011) |
| 0.1 | 0.7 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) |
| 0.1 | 0.1 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.1 | 2.2 | GO:0005527 | macrolide binding(GO:0005527) FK506 binding(GO:0005528) |
| 0.1 | 0.4 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.1 | 0.3 | GO:0004027 | alcohol sulfotransferase activity(GO:0004027) |
| 0.1 | 0.3 | GO:0048248 | CXCR3 chemokine receptor binding(GO:0048248) |
| 0.1 | 1.1 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.1 | 0.5 | GO:0050693 | LBD domain binding(GO:0050693) |
| 0.1 | 3.2 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.1 | 1.7 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.1 | 0.9 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 1.1 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.1 | 0.1 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.1 | 2.9 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 2.0 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 0.6 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 0.3 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.1 | 3.3 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.1 | 0.5 | GO:0033300 | dehydroascorbic acid transporter activity(GO:0033300) |
| 0.1 | 0.1 | GO:1901611 | phosphatidylglycerol binding(GO:1901611) cardiolipin binding(GO:1901612) |
| 0.1 | 1.0 | GO:0015245 | fatty acid transporter activity(GO:0015245) |
| 0.1 | 1.4 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 0.7 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 0.1 | 2.5 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 0.7 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) glycerol channel activity(GO:0015254) |
| 0.1 | 0.5 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.1 | 0.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 0.4 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.1 | 0.7 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) |
| 0.1 | 0.4 | GO:0043262 | adenosine-diphosphatase activity(GO:0043262) |
| 0.1 | 0.1 | GO:0033142 | progesterone receptor binding(GO:0033142) |
| 0.1 | 0.3 | GO:0016520 | growth hormone-releasing hormone receptor activity(GO:0016520) |
| 0.1 | 1.0 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.1 | 0.7 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 0.5 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.1 | 0.4 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 0.1 | 0.2 | GO:0050252 | retinol O-fatty-acyltransferase activity(GO:0050252) |
| 0.1 | 2.5 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 0.6 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.1 | 0.3 | GO:0005167 | neurotrophin TRK receptor binding(GO:0005167) |
| 0.1 | 0.7 | GO:0022889 | L-serine transmembrane transporter activity(GO:0015194) serine transmembrane transporter activity(GO:0022889) |
| 0.1 | 0.3 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.1 | 0.9 | GO:0016717 | oxidoreductase activity, acting on paired donors, with oxidation of a pair of donors resulting in the reduction of molecular oxygen to two molecules of water(GO:0016717) |
| 0.1 | 1.1 | GO:0070402 | NADPH binding(GO:0070402) |
| 0.1 | 0.4 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.1 | 0.2 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
| 0.1 | 0.3 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.1 | 0.8 | GO:0016493 | C-C chemokine receptor activity(GO:0016493) |
| 0.1 | 0.3 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.1 | 0.3 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.1 | 0.2 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.1 | 0.3 | GO:0017089 | glycolipid transporter activity(GO:0017089) |
| 0.1 | 0.5 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.1 | 1.1 | GO:0030955 | potassium ion binding(GO:0030955) |
| 0.1 | 11.2 | GO:0003729 | mRNA binding(GO:0003729) |
| 0.1 | 1.1 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 1.0 | GO:0008601 | protein phosphatase type 2A regulator activity(GO:0008601) |
| 0.1 | 0.3 | GO:0005275 | amine transmembrane transporter activity(GO:0005275) |
| 0.1 | 0.3 | GO:0031727 | CCR2 chemokine receptor binding(GO:0031727) |
| 0.1 | 2.2 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.1 | 0.4 | GO:0032795 | heterotrimeric G-protein binding(GO:0032795) |
| 0.1 | 0.1 | GO:0055131 | C3HC4-type RING finger domain binding(GO:0055131) |
| 0.1 | 0.7 | GO:0005326 | neurotransmitter transporter activity(GO:0005326) |
| 0.1 | 0.1 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.1 | 0.2 | GO:0030292 | protein tyrosine kinase inhibitor activity(GO:0030292) |
| 0.1 | 0.4 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.1 | 0.4 | GO:0048156 | tau protein binding(GO:0048156) |
| 0.1 | 0.4 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.1 | 0.4 | GO:0047760 | butyrate-CoA ligase activity(GO:0047760) |
| 0.1 | 0.1 | GO:0051033 | nucleic acid transmembrane transporter activity(GO:0051032) RNA transmembrane transporter activity(GO:0051033) |
| 0.1 | 0.2 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.1 | 0.8 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 0.5 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.1 | 0.3 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.1 | 0.1 | GO:0031698 | beta-2 adrenergic receptor binding(GO:0031698) |
| 0.1 | 0.1 | GO:0004645 | phosphorylase activity(GO:0004645) |
| 0.1 | 0.5 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 0.2 | GO:1904680 | peptide transmembrane transporter activity(GO:1904680) |
| 0.1 | 0.1 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.1 | 0.3 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.1 | 0.2 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 0.1 | 0.3 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
| 0.1 | 0.2 | GO:0060230 | lipoprotein lipase activator activity(GO:0060230) |
| 0.1 | 0.1 | GO:0004611 | phosphoenolpyruvate carboxykinase activity(GO:0004611) |
| 0.1 | 0.2 | GO:0035033 | histone deacetylase regulator activity(GO:0035033) |
| 0.1 | 1.2 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 0.2 | GO:0089720 | caspase binding(GO:0089720) |
| 0.1 | 0.3 | GO:0015189 | arginine transmembrane transporter activity(GO:0015181) L-lysine transmembrane transporter activity(GO:0015189) |
| 0.1 | 0.7 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.1 | 0.2 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.1 | 1.3 | GO:0033558 | histone deacetylase activity(GO:0004407) protein deacetylase activity(GO:0033558) |
| 0.1 | 0.4 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.1 | 0.3 | GO:0001135 | transcription factor activity, RNA polymerase II transcription factor recruiting(GO:0001135) |
| 0.1 | 0.2 | GO:0050220 | prostaglandin-E synthase activity(GO:0050220) |
| 0.1 | 0.1 | GO:0050815 | phosphoserine binding(GO:0050815) |
| 0.1 | 0.3 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.1 | 0.2 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.1 | 1.6 | GO:0043883 | N-ethylmaleimide reductase activity(GO:0008748) reduced coenzyme F420 dehydrogenase activity(GO:0043738) sulfur oxygenase reductase activity(GO:0043826) malolactic enzyme activity(GO:0043883) epoxyqueuosine reductase activity(GO:0052693) |
| 0.1 | 0.9 | GO:0019239 | deaminase activity(GO:0019239) |
| 0.1 | 0.9 | GO:0004012 | phospholipid-translocating ATPase activity(GO:0004012) |
| 0.1 | 0.1 | GO:0046978 | TAP1 binding(GO:0046978) TAP2 binding(GO:0046979) |
| 0.1 | 0.2 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.1 | 1.0 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.1 | 2.7 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.1 | 0.3 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.1 | 0.3 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.1 | 0.2 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.1 | 0.2 | GO:0015368 | calcium:cation antiporter activity(GO:0015368) |
| 0.0 | 0.2 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 0.1 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) |
| 0.0 | 0.1 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 0.0 | 0.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.1 | GO:0016880 | acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.0 | 0.5 | GO:0030676 | Rac guanyl-nucleotide exchange factor activity(GO:0030676) |
| 0.0 | 0.1 | GO:0005087 | Ran guanyl-nucleotide exchange factor activity(GO:0005087) |
| 0.0 | 1.1 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.0 | 0.1 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.0 | 0.5 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 0.1 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.2 | GO:0048403 | brain-derived neurotrophic factor binding(GO:0048403) |
| 0.0 | 1.4 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 0.0 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.0 | 0.0 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.1 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.0 | 0.3 | GO:0015321 | sodium-dependent phosphate transmembrane transporter activity(GO:0015321) |
| 0.0 | 0.8 | GO:0005537 | mannose binding(GO:0005537) |
| 0.0 | 0.7 | GO:0016247 | channel regulator activity(GO:0016247) |
| 0.0 | 2.2 | GO:0005261 | cation channel activity(GO:0005261) |
| 0.0 | 0.3 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.0 | 0.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.9 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.2 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.3 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.1 | GO:0015665 | alcohol transmembrane transporter activity(GO:0015665) |
| 0.0 | 0.7 | GO:0042923 | neuropeptide binding(GO:0042923) |
| 0.0 | 0.1 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.0 | 0.3 | GO:0005452 | inorganic anion exchanger activity(GO:0005452) |
| 0.0 | 0.4 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.0 | 0.0 | GO:0043398 | HLH domain binding(GO:0043398) |
| 0.0 | 0.3 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 0.0 | 0.0 | GO:0031748 | D1 dopamine receptor binding(GO:0031748) |
| 0.0 | 0.1 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.3 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.1 | GO:0099602 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.0 | 0.1 | GO:0043842 | Kdo transferase activity(GO:0043842) |
| 0.0 | 0.2 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 0.3 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 0.1 | GO:0008147 | structural constituent of bone(GO:0008147) |
| 0.0 | 0.3 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 0.0 | GO:0035175 | histone kinase activity (H3-S10 specific)(GO:0035175) |
| 0.0 | 0.4 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.2 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.5 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.0 | 0.7 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) |
| 0.0 | 0.1 | GO:0050733 | RS domain binding(GO:0050733) |
| 0.0 | 0.4 | GO:0016805 | dipeptidase activity(GO:0016805) |
| 0.0 | 0.1 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.0 | 0.2 | GO:0070891 | lipoteichoic acid binding(GO:0070891) |
| 0.0 | 0.0 | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen(GO:0016701) |
| 0.0 | 0.0 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.0 | 5.6 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.1 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.1 | GO:0004439 | phosphatidylinositol-4,5-bisphosphate 5-phosphatase activity(GO:0004439) |
| 0.0 | 0.6 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.1 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.3 | GO:0016290 | palmitoyl-CoA hydrolase activity(GO:0016290) |
| 0.0 | 0.1 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) |
| 0.0 | 0.2 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.0 | 0.0 | GO:0016595 | glutamate binding(GO:0016595) |
| 0.0 | 0.1 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 0.1 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 0.0 | GO:0034618 | arginine binding(GO:0034618) |
| 0.0 | 0.1 | GO:0019767 | IgE receptor activity(GO:0019767) |
| 0.0 | 0.0 | GO:0008035 | high-density lipoprotein particle binding(GO:0008035) |
| 0.0 | 0.6 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.0 | 0.1 | GO:0003872 | 6-phosphofructokinase activity(GO:0003872) |
| 0.0 | 0.4 | GO:0004181 | metallocarboxypeptidase activity(GO:0004181) |
| 0.0 | 0.1 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.0 | 0.1 | GO:0008865 | fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.0 | 0.1 | GO:0015038 | glutathione disulfide oxidoreductase activity(GO:0015038) |
| 0.0 | 0.2 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.2 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.0 | 0.1 | GO:0030158 | protein xylosyltransferase activity(GO:0030158) |
| 0.0 | 0.0 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.0 | 0.1 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.1 | GO:0015464 | acetylcholine receptor activity(GO:0015464) |
| 0.0 | 0.5 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.0 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) |
| 0.0 | 0.3 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.1 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 0.1 | GO:0002094 | polyprenyltransferase activity(GO:0002094) |
| 0.0 | 0.1 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.0 | 0.0 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.0 | 0.0 | GO:0047276 | N-acetyllactosaminide 3-alpha-galactosyltransferase activity(GO:0047276) |
| 0.0 | 0.8 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.0 | 0.0 | GO:0004337 | geranyltranstransferase activity(GO:0004337) |
| 0.0 | 0.0 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.0 | 0.0 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.1 | GO:0043138 | 3'-5' DNA helicase activity(GO:0043138) |
| 0.0 | 0.1 | GO:0001162 | RNA polymerase II intronic transcription regulatory region sequence-specific DNA binding(GO:0001162) |
| 0.0 | 0.0 | GO:0050051 | leukotriene-B4 20-monooxygenase activity(GO:0050051) |
| 0.0 | 0.1 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 18.5 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.6 | 18.7 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.5 | 7.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.3 | 0.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.3 | 3.0 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.3 | 19.2 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.3 | 11.0 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.3 | 3.6 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.3 | 2.3 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.2 | 0.2 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.2 | 11.5 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.2 | 2.9 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.2 | 3.5 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.2 | 2.6 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.1 | 1.3 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.1 | 3.1 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 0.1 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.1 | 2.2 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.1 | 1.2 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.1 | 0.1 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 0.1 | PID FAS PATHWAY | FAS (CD95) signaling pathway |
| 0.1 | 2.3 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 1.4 | PID HDAC CLASSII PATHWAY | Signaling events mediated by HDAC Class II |
| 0.1 | 0.9 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.0 | 0.2 | PID ER NONGENOMIC PATHWAY | Plasma membrane estrogen receptor signaling |
| 0.0 | 0.7 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.8 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.7 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.0 | 0.3 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 0.4 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.7 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.2 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.0 | PID EPHA2 FWD PATHWAY | EPHA2 forward signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 18.1 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 1.1 | 13.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 1.0 | 25.1 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.9 | 3.8 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.9 | 0.9 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.9 | 7.8 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.7 | 7.8 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.7 | 12.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.7 | 13.6 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.6 | 9.1 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.5 | 3.3 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.5 | 5.8 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.5 | 6.7 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.4 | 4.6 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.4 | 8.1 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.4 | 5.5 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.4 | 4.9 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.3 | 5.5 | REACTOME PKA MEDIATED PHOSPHORYLATION OF CREB | Genes involved in PKA-mediated phosphorylation of CREB |
| 0.3 | 2.3 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.3 | 0.3 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.3 | 5.0 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.3 | 3.1 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.3 | 3.1 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 4.6 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.2 | 0.7 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.2 | 3.5 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.2 | 0.9 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.2 | 1.9 | REACTOME GABA B RECEPTOR ACTIVATION | Genes involved in GABA B receptor activation |
| 0.2 | 8.4 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.2 | 2.3 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 1.2 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.2 | 0.5 | REACTOME ADENYLATE CYCLASE INHIBITORY PATHWAY | Genes involved in Adenylate cyclase inhibitory pathway |
| 0.2 | 4.1 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.2 | 6.7 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.2 | 3.1 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.2 | 2.2 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.2 | 6.3 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.2 | 2.0 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 1.9 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.1 | 0.9 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.1 | 0.1 | REACTOME OPSINS | Genes involved in Opsins |
| 0.1 | 2.0 | REACTOME THE ROLE OF NEF IN HIV1 REPLICATION AND DISEASE PATHOGENESIS | Genes involved in The role of Nef in HIV-1 replication and disease pathogenesis |
| 0.1 | 2.3 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 2.1 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 0.1 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.1 | 0.3 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |
| 0.1 | 1.4 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |
| 0.1 | 0.4 | REACTOME INCRETIN SYNTHESIS SECRETION AND INACTIVATION | Genes involved in Incretin Synthesis, Secretion, and Inactivation |
| 0.1 | 4.3 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.1 | 2.3 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.1 | 0.6 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.1 | 1.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 0.3 | REACTOME SHC MEDIATED SIGNALLING | Genes involved in SHC-mediated signalling |
| 0.1 | 1.9 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.1 | 0.3 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.1 | 0.8 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.1 | 1.1 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.1 | 7.4 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.1 | 0.6 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 0.4 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.1 | 0.4 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.0 | 0.5 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.0 | 0.4 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.1 | REACTOME XENOBIOTICS | Genes involved in Xenobiotics |
| 0.0 | 0.6 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.0 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.0 | REACTOME REGULATION OF INSULIN SECRETION | Genes involved in Regulation of Insulin Secretion |
| 0.0 | 0.1 | REACTOME PLATELET HOMEOSTASIS | Genes involved in Platelet homeostasis |
| 0.0 | 0.0 | REACTOME SHC1 EVENTS IN EGFR SIGNALING | Genes involved in SHC1 events in EGFR signaling |
| 0.0 | 0.1 | REACTOME REGULATED PROTEOLYSIS OF P75NTR | Genes involved in Regulated proteolysis of p75NTR |
| 0.0 | 0.3 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.3 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.0 | REACTOME DOWNSTREAM TCR SIGNALING | Genes involved in Downstream TCR signaling |