Project
ENCODE: H3K4me3 ChIP-Seq of different mouse tissues during embryonic development
Navigation
Downloads
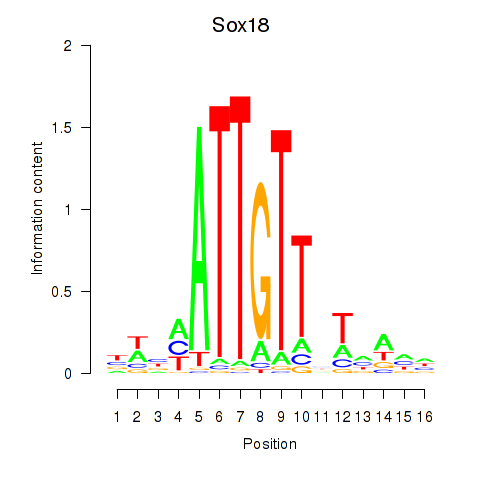
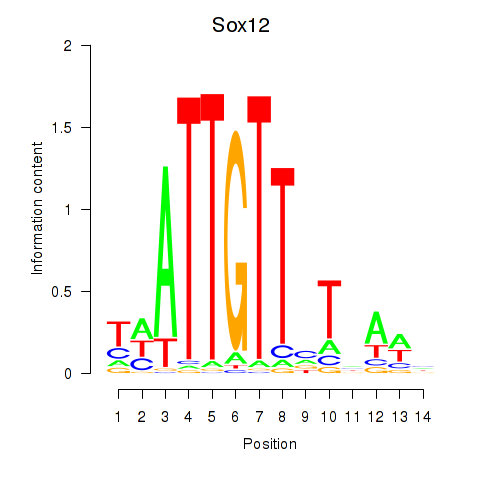
Results for Sox18_Sox12
Z-value: 1.78
Transcription factors associated with Sox18_Sox12
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Sox18
|
ENSMUSG00000046470.5 | SRY (sex determining region Y)-box 18 |
|
Sox12
|
ENSMUSG00000051817.8 | SRY (sex determining region Y)-box 12 |
Correlations of motif activity and signal intensity at CREs associated with the motif's TFs:
This plot shows correlation between observed signal intensity of a CRE associated with the transcription factor across all samples and activity of the motif.
For each TF, only the top 5 correlated CREs are shown.
| CRE | Gene | Distance | Association probability | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|---|---|
| chr2_152405648_152405895 | Sox12 | 7708 | 0.082670 | -0.38 | 2.8e-03 | Click! |
| chr2_152405939_152406162 | Sox12 | 7987 | 0.082092 | -0.34 | 8.2e-03 | Click! |
| chr2_152394551_152394958 | Sox12 | 3292 | 0.110176 | 0.30 | 2.2e-02 | Click! |
| chr2_152395546_152396207 | Sox12 | 2170 | 0.145788 | 0.23 | 7.9e-02 | Click! |
| chr2_152406328_152406620 | Sox12 | 8411 | 0.081268 | -0.21 | 1.0e-01 | Click! |
| chr2_181675377_181676046 | Sox18 | 4071 | 0.103898 | -0.43 | 6.0e-04 | Click! |
| chr2_181669407_181669715 | Sox18 | 2079 | 0.158464 | -0.22 | 9.4e-02 | Click! |
| chr2_181670114_181670265 | Sox18 | 1451 | 0.223742 | -0.15 | 2.5e-01 | Click! |
| chr2_181670330_181671688 | Sox18 | 631 | 0.519654 | -0.02 | 8.6e-01 | Click! |
Activity of the Sox18_Sox12 motif across conditions
Conditions sorted by the z-value of the Sox18_Sox12 motif activity
Move your cursor over a bar to see sample name and corresponding Z-value.
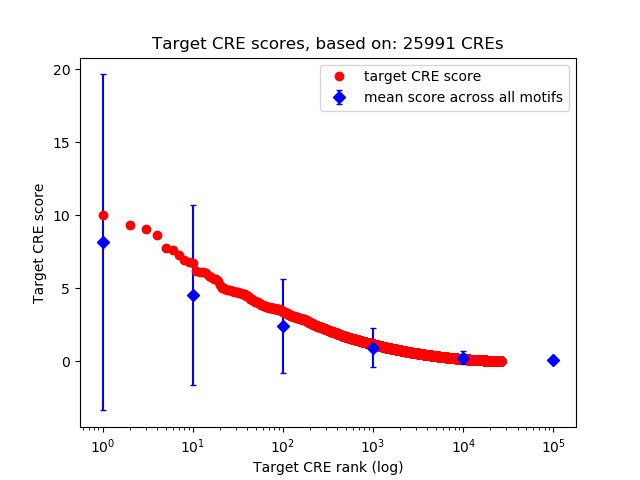
Top target CREs of the motif:
| Cis Regulatory Element (CRE) | Target Score | Top associated gene | Gene Info | Distance of CRE to TSS | CRE/Gene association probability |
|---|---|---|---|---|---|
| chr15_95525022_95525792 | 10.04 |
Nell2 |
NEL-like 2 |
2771 |
0.35 |
| chr2_65620767_65621991 | 9.31 |
Scn2a |
sodium channel, voltage-gated, type II, alpha |
568 |
0.82 |
| chrX_152643367_152644550 | 9.04 |
Shroom2 |
shroom family member 2 |
34 |
0.98 |
| chr12_72234504_72235243 | 8.66 |
Rtn1 |
reticulon 1 |
866 |
0.66 |
| chr10_92160735_92161461 | 7.75 |
Rmst |
rhabdomyosarcoma 2 associated transcript (non-coding RNA) |
1663 |
0.4 |
| chrX_143930842_143933141 | 7.60 |
Dcx |
doublecortin |
1059 |
0.64 |
| chr16_77594640_77595970 | 7.32 |
Mir99ahg |
Mir99a and Mirlet7c-1 host gene (non-protein coding) |
403 |
0.71 |
| chr4_22485441_22485749 | 6.97 |
Pou3f2 |
POU domain, class 3, transcription factor 2 |
2771 |
0.23 |
| chr8_54957303_54957776 | 6.82 |
Gm45263 |
predicted gene 45263 |
2280 |
0.24 |
| chr14_66344363_66345813 | 6.72 |
Stmn4 |
stathmin-like 4 |
707 |
0.65 |
| chr2_158606690_158608449 | 6.17 |
Gm14204 |
predicted gene 14204 |
3021 |
0.15 |
| chr7_62366195_62366527 | 6.11 |
Magel2 |
melanoma antigen, family L, 2 |
10649 |
0.18 |
| chr5_88583963_88584847 | 6.09 |
Rufy3 |
RUN and FYVE domain containing 3 |
611 |
0.7 |
| chr10_111249619_111250191 | 6.05 |
Osbpl8 |
oxysterol binding protein-like 8 |
1837 |
0.35 |
| chr2_181767278_181768191 | 5.87 |
Myt1 |
myelin transcription factor 1 |
222 |
0.91 |
| chr1_42701498_42701775 | 5.78 |
Pou3f3 |
POU domain, class 3, transcription factor 3 |
5868 |
0.14 |
| chr11_54596874_54597434 | 5.67 |
Rapgef6 |
Rap guanine nucleotide exchange factor (GEF) 6 |
980 |
0.6 |
| chr9_52148115_52149635 | 5.63 |
Zc3h12c |
zinc finger CCCH type containing 12C |
19236 |
0.18 |
| chr4_22481292_22481918 | 5.47 |
Pou3f2 |
POU domain, class 3, transcription factor 2 |
6761 |
0.17 |
| chr13_28419787_28420291 | 5.24 |
Gm40841 |
predicted gene, 40841 |
27 |
0.98 |
| chr8_85698168_85698436 | 5.05 |
Neto2 |
neuropilin (NRP) and tolloid (TLL)-like 2 |
2452 |
0.21 |
| chr7_144238658_144240098 | 5.02 |
Shank2 |
SH3 and multiple ankyrin repeat domains 2 |
653 |
0.8 |
| chr4_82501450_82502014 | 4.98 |
Nfib |
nuclear factor I/B |
2416 |
0.3 |
| chr1_9601164_9602408 | 4.90 |
Vxn |
vexin |
587 |
0.67 |
| chr8_54956010_54956394 | 4.90 |
Gpm6a |
glycoprotein m6a |
1359 |
0.38 |
| chr9_52145956_52147029 | 4.87 |
Zc3h12c |
zinc finger CCCH type containing 12C |
21619 |
0.17 |
| chr2_6881042_6881689 | 4.85 |
Gm13389 |
predicted gene 13389 |
2905 |
0.24 |
| chr4_24429901_24430719 | 4.81 |
Gm27243 |
predicted gene 27243 |
580 |
0.79 |
| chr3_17790150_17790808 | 4.75 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
522 |
0.77 |
| chr2_79452354_79452909 | 4.73 |
Neurod1 |
neurogenic differentiation 1 |
4120 |
0.24 |
| chr12_29529828_29531185 | 4.72 |
Gm20208 |
predicted gene, 20208 |
609 |
0.74 |
| chrX_166346283_166346827 | 4.71 |
Gpm6b |
glycoprotein m6b |
1713 |
0.43 |
| chr13_34126566_34127191 | 4.66 |
Tubb2b |
tubulin, beta 2B class IIB |
3476 |
0.13 |
| chr9_41378412_41379411 | 4.66 |
Mir100hg |
Mir100 Mirlet7a-2 Mir125b-1 cluster host gene |
2350 |
0.27 |
| chr1_143640264_143641520 | 4.64 |
B3galt2 |
UDP-Gal:betaGlcNAc beta 1,3-galactosyltransferase, polypeptide 2 |
228 |
0.59 |
| chrX_23284413_23285126 | 4.62 |
Klhl13 |
kelch-like 13 |
60 |
0.99 |
| chr18_35212708_35213458 | 4.61 |
Ctnna1 |
catenin (cadherin associated protein), alpha 1 |
1847 |
0.28 |
| chr18_72347538_72348154 | 4.59 |
Dcc |
deleted in colorectal carcinoma |
3171 |
0.38 |
| chr1_137901653_137901959 | 4.51 |
Gm4258 |
predicted gene 4258 |
3208 |
0.12 |
| chr1_66388178_66389004 | 4.47 |
Map2 |
microtubule-associated protein 2 |
1580 |
0.42 |
| chr17_91085493_91086001 | 4.45 |
Gm47307 |
predicted gene, 47307 |
2659 |
0.21 |
| chr14_98164357_98165375 | 4.41 |
Dach1 |
dachshund family transcription factor 1 |
4677 |
0.28 |
| chr8_41054476_41055299 | 4.30 |
Mtus1 |
mitochondrial tumor suppressor 1 |
93 |
0.95 |
| chrX_110818058_110818665 | 4.28 |
Pou3f4 |
POU domain, class 3, transcription factor 4 |
4081 |
0.27 |
| chr5_131532921_131534054 | 4.25 |
Auts2 |
autism susceptibility candidate 2 |
910 |
0.58 |
| chr18_31445651_31446131 | 4.17 |
Syt4 |
synaptotagmin IV |
1515 |
0.34 |
| chr18_77560987_77561705 | 4.17 |
Rnf165 |
ring finger protein 165 |
3263 |
0.29 |
| chr3_5224377_5225076 | 4.10 |
Zfhx4 |
zinc finger homeodomain 4 |
3221 |
0.24 |
| chr4_110282527_110283235 | 4.08 |
Elavl4 |
ELAV like RNA binding protein 4 |
3735 |
0.36 |
| chr13_109443973_109444887 | 4.07 |
Pde4d |
phosphodiesterase 4D, cAMP specific |
2247 |
0.46 |
| chr5_111421306_111422790 | 4.06 |
Gm43119 |
predicted gene 43119 |
1541 |
0.35 |
| chr19_47017205_47017356 | 4.04 |
Nt5c2 |
5'-nucleotidase, cytosolic II |
2127 |
0.2 |
| chr14_122477033_122477655 | 4.04 |
Zic2 |
zinc finger protein of the cerebellum 2 |
756 |
0.48 |
| chr5_131531462_131532675 | 4.03 |
Auts2 |
autism susceptibility candidate 2 |
2329 |
0.29 |
| chr13_84063384_84064052 | 4.00 |
Gm17750 |
predicted gene, 17750 |
1054 |
0.58 |
| chr8_94268148_94268983 | 3.91 |
Nup93 |
nucleoporin 93 |
41 |
0.96 |
| chr1_137902039_137902674 | 3.85 |
Gm4258 |
predicted gene 4258 |
3758 |
0.11 |
| chr15_30458403_30458947 | 3.85 |
Ctnnd2 |
catenin (cadherin associated protein), delta 2 |
887 |
0.67 |
| chr10_45889498_45890055 | 3.83 |
Gpx4-ps2 |
glutathione peroxidase 4, pseudogene 2 |
14471 |
0.23 |
| chr11_108607202_108607707 | 3.83 |
Cep112 |
centrosomal protein 112 |
2227 |
0.37 |
| chr1_66386919_66387899 | 3.82 |
Map2 |
microtubule-associated protein 2 |
398 |
0.87 |
| chr5_88583326_88583955 | 3.82 |
Rufy3 |
RUN and FYVE domain containing 3 |
62 |
0.97 |
| chr5_117415205_117415356 | 3.80 |
Ksr2 |
kinase suppressor of ras 2 |
1280 |
0.37 |
| chr2_65566848_65567533 | 3.79 |
Scn3a |
sodium channel, voltage-gated, type III, alpha |
302 |
0.92 |
| chr13_83734808_83735118 | 3.73 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
2397 |
0.18 |
| chrX_88114468_88114695 | 3.73 |
Il1rapl1 |
interleukin 1 receptor accessory protein-like 1 |
1064 |
0.64 |
| chr9_112229925_112230278 | 3.73 |
Arpp21 |
cyclic AMP-regulated phosphoprotein, 21 |
1801 |
0.31 |
| chrX_48519996_48520368 | 3.73 |
Rab33a |
RAB33A, member RAS oncogene family |
897 |
0.55 |
| chr3_17783692_17784517 | 3.71 |
Mir124-2hg |
Mir124-2 host gene (non-protein coding) |
5817 |
0.2 |
| chr8_12399326_12400483 | 3.71 |
Gm25239 |
predicted gene, 25239 |
3501 |
0.16 |
| chrX_166344665_166345995 | 3.68 |
Gpm6b |
glycoprotein m6b |
488 |
0.85 |
| chr7_92234907_92236280 | 3.67 |
Dlg2 |
discs large MAGUK scaffold protein 2 |
466 |
0.88 |
| chr1_177444257_177446079 | 3.65 |
Zbtb18 |
zinc finger and BTB domain containing 18 |
230 |
0.9 |
| chr5_103210548_103211780 | 3.64 |
Mapk10 |
mitogen-activated protein kinase 10 |
109 |
0.98 |
| chr3_13946382_13947629 | 3.64 |
Ralyl |
RALY RNA binding protein-like |
594 |
0.84 |
| chr2_97470615_97471412 | 3.64 |
Lrrc4c |
leucine rich repeat containing 4C |
2924 |
0.4 |
| chr1_42700819_42701404 | 3.64 |
Pou3f3 |
POU domain, class 3, transcription factor 3 |
5343 |
0.14 |
| chr7_60449424_60450258 | 3.63 |
Gm30196 |
predicted gene, 30196 |
156637 |
0.03 |
| chr1_81079064_81079824 | 3.62 |
Nyap2 |
neuronal tyrosine-phophorylated phosphoinositide 3-kinase adaptor 2 |
1861 |
0.49 |
| chr4_109342938_109343450 | 3.62 |
Eps15 |
epidermal growth factor receptor pathway substrate 15 |
59 |
0.97 |
| chr8_33747278_33748028 | 3.62 |
Smim18 |
small integral membrane protein 18 |
117 |
0.95 |
| chr3_34656299_34657721 | 3.60 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
974 |
0.4 |
| chr8_31915784_31916390 | 3.60 |
Nrg1 |
neuregulin 1 |
1563 |
0.42 |
| chr12_29528407_29529244 | 3.58 |
Myt1l |
myelin transcription factor 1-like |
424 |
0.85 |
| chr15_98986992_98987777 | 3.58 |
4930578M01Rik |
RIKEN cDNA 4930578M01 gene |
1490 |
0.22 |
| chr13_84056577_84057434 | 3.58 |
Gm17750 |
predicted gene, 17750 |
7767 |
0.22 |
| chr3_4796861_4798079 | 3.57 |
1110015O18Rik |
RIKEN cDNA 1110015O18 gene |
88 |
0.98 |
| chr4_116223265_116224130 | 3.57 |
Pik3r3 |
phosphoinositide-3-kinase regulatory subunit 3 |
2001 |
0.23 |
| chr10_49785211_49786117 | 3.57 |
Grik2 |
glutamate receptor, ionotropic, kainate 2 (beta 2) |
2255 |
0.25 |
| chr2_83814030_83814462 | 3.55 |
Fam171b |
family with sequence similarity 171, member B |
1610 |
0.34 |
| chr16_77598670_77599917 | 3.53 |
Mir99a |
microRNA 99a |
357 |
0.44 |
| chr8_45507516_45508498 | 3.52 |
Sorbs2 |
sorbin and SH3 domain containing 2 |
89 |
0.97 |
| chr7_51626624_51628140 | 3.52 |
Slc17a6 |
solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 |
2098 |
0.31 |
| chr12_71049290_71050137 | 3.52 |
Arid4a |
AT rich interactive domain 4A (RBP1-like) |
1372 |
0.4 |
| chr5_15939238_15939704 | 3.50 |
Cacna2d1 |
calcium channel, voltage-dependent, alpha2/delta subunit 1 |
4560 |
0.18 |
| chr11_87759834_87761999 | 3.50 |
Tspoap1 |
TSPO associated protein 1 |
329 |
0.75 |
| chr1_64116857_64117480 | 3.45 |
Klf7 |
Kruppel-like factor 7 (ubiquitous) |
4314 |
0.23 |
| chr2_105670737_105671693 | 3.42 |
Pax6 |
paired box 6 |
645 |
0.51 |
| chr5_150260534_150260992 | 3.41 |
Fry |
FRY microtubule binding protein |
996 |
0.55 |
| chr8_41052368_41053980 | 3.39 |
Gm16193 |
predicted gene 16193 |
64 |
0.96 |
| chr10_92161472_92161916 | 3.38 |
Rmst |
rhabdomyosarcoma 2 associated transcript (non-coding RNA) |
1067 |
0.55 |
| chr5_116589538_116590511 | 3.38 |
Srrm4 |
serine/arginine repetitive matrix 4 |
1793 |
0.34 |
| chr3_8512495_8512918 | 3.36 |
Stmn2 |
stathmin-like 2 |
3120 |
0.28 |
| chr1_25830818_25831017 | 3.35 |
Gm9884 |
predicted gene 9884 |
260 |
0.82 |
| chr2_181763361_181764530 | 3.34 |
Myt1 |
myelin transcription factor 1 |
613 |
0.66 |
| chr16_88287797_88287976 | 3.32 |
Grik1 |
glutamate receptor, ionotropic, kainate 1 |
1845 |
0.43 |
| chr3_102010214_102010759 | 3.32 |
Nhlh2 |
nescient helix loop helix 2 |
331 |
0.89 |
| chr12_117157079_117158175 | 3.32 |
Gm10421 |
predicted gene 10421 |
5976 |
0.31 |
| chr1_89071813_89072578 | 3.31 |
Sh3bp4 |
SH3-domain binding protein 4 |
1742 |
0.35 |
| chr9_112231189_112232055 | 3.29 |
Arpp21 |
cyclic AMP-regulated phosphoprotein, 21 |
280 |
0.9 |
| chr13_109442519_109443753 | 3.28 |
Pde4d |
phosphodiesterase 4D, cAMP specific |
953 |
0.73 |
| chr3_34651759_34652037 | 3.26 |
Sox2 |
SRY (sex determining region Y)-box 2 |
1493 |
0.26 |
| chr14_100374663_100375528 | 3.24 |
Gm26367 |
predicted gene, 26367 |
43388 |
0.15 |
| chr2_52557337_52558561 | 3.23 |
Cacnb4 |
calcium channel, voltage-dependent, beta 4 subunit |
611 |
0.74 |
| chr1_194622071_194623282 | 3.23 |
Plxna2 |
plexin A2 |
2851 |
0.26 |
| chr6_77246252_77246852 | 3.22 |
Lrrtm1 |
leucine rich repeat transmembrane neuronal 1 |
3630 |
0.32 |
| chr10_57787071_57787893 | 3.17 |
Fabp7 |
fatty acid binding protein 7, brain |
2559 |
0.25 |
| chr2_65932868_65933620 | 3.17 |
Csrnp3 |
cysteine-serine-rich nuclear protein 3 |
1379 |
0.47 |
| chr12_119235620_119236481 | 3.16 |
Itgb8 |
integrin beta 8 |
2720 |
0.31 |
| chr11_108606161_108607110 | 3.16 |
Cep112 |
centrosomal protein 112 |
1408 |
0.5 |
| chr18_35210523_35210985 | 3.15 |
Ctnna1 |
catenin (cadherin associated protein), alpha 1 |
4176 |
0.2 |
| chr4_110283765_110284001 | 3.12 |
Elavl4 |
ELAV like RNA binding protein 4 |
2733 |
0.4 |
| chr9_96731522_96733329 | 3.12 |
Zbtb38 |
zinc finger and BTB domain containing 38 |
244 |
0.91 |
| chr9_75683375_75684591 | 3.11 |
Scg3 |
secretogranin III |
8 |
0.97 |
| chrX_105390628_105392456 | 3.10 |
5330434G04Rik |
RIKEN cDNA 5330434G04 gene |
212 |
0.93 |
| chr14_93883900_93884713 | 3.10 |
Pcdh9 |
protocadherin 9 |
1442 |
0.55 |
| chr13_42709652_42710400 | 3.09 |
Phactr1 |
phosphatase and actin regulator 1 |
445 |
0.88 |
| chr11_80478619_80479391 | 3.09 |
Cdk5r1 |
cyclin-dependent kinase 5, regulatory subunit 1 (p35) |
1949 |
0.32 |
| chr10_29143400_29144848 | 3.09 |
Soga3 |
SOGA family member 3 |
65 |
0.5 |
| chr12_49386970_49387121 | 3.08 |
Gm43517 |
predicted gene 43517 |
1871 |
0.22 |
| chr18_72023211_72023790 | 3.07 |
Dcc |
deleted in colorectal carcinoma |
327517 |
0.01 |
| chr6_17750183_17750783 | 3.07 |
St7 |
suppression of tumorigenicity 7 |
1091 |
0.38 |
| chr16_16558986_16560577 | 3.06 |
Fgd4 |
FYVE, RhoGEF and PH domain containing 4 |
209 |
0.94 |
| chr7_90386713_90387752 | 3.05 |
Sytl2 |
synaptotagmin-like 2 |
127 |
0.96 |
| chr17_78508180_78509392 | 3.05 |
Vit |
vitrin |
614 |
0.7 |
| chr5_63651264_63652181 | 3.04 |
Gm9954 |
predicted gene 9954 |
828 |
0.61 |
| chr9_43070574_43070912 | 3.04 |
Arhgef12 |
Rho guanine nucleotide exchange factor (GEF) 12 |
28111 |
0.16 |
| chr3_34654574_34655689 | 3.03 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
905 |
0.42 |
| chr19_44757990_44758435 | 3.02 |
Pax2 |
paired box 2 |
818 |
0.54 |
| chr3_34562856_34563429 | 3.01 |
Sox2ot |
SOX2 overlapping transcript (non-protein coding) |
2750 |
0.22 |
| chr17_91092075_91093120 | 3.01 |
Nrxn1 |
neurexin I |
136 |
0.95 |
| chrX_9202282_9203095 | 3.01 |
Lancl3 |
LanC lantibiotic synthetase component C-like 3 (bacterial) |
2786 |
0.21 |
| chr4_82499658_82501360 | 3.01 |
Nfib |
nuclear factor I/B |
1193 |
0.5 |
| chr2_48538709_48539576 | 2.99 |
Gm13481 |
predicted gene 13481 |
81897 |
0.1 |
| chr7_87586513_87587584 | 2.98 |
Grm5 |
glutamate receptor, metabotropic 5 |
2650 |
0.4 |
| chr2_6881874_6882908 | 2.98 |
Gm13389 |
predicted gene 13389 |
1879 |
0.3 |
| chr9_96729464_96730774 | 2.97 |
Zbtb38 |
zinc finger and BTB domain containing 38 |
1083 |
0.47 |
| chr6_105680400_105681277 | 2.97 |
Cntn4 |
contactin 4 |
3063 |
0.29 |
| chr12_75175558_75175725 | 2.97 |
Kcnh5 |
potassium voltage-gated channel, subfamily H (eag-related), member 5 |
1691 |
0.52 |
| chr12_107997791_107998940 | 2.96 |
Bcl11b |
B cell leukemia/lymphoma 11B |
5049 |
0.31 |
| chr9_121299469_121300553 | 2.95 |
Trak1 |
trafficking protein, kinesin binding 1 |
2182 |
0.23 |
| chr7_51620788_51621096 | 2.94 |
Slc17a6 |
solute carrier family 17 (sodium-dependent inorganic phosphate cotransporter), member 6 |
1064 |
0.54 |
| chr10_102159422_102159718 | 2.93 |
Mgat4c |
MGAT4 family, member C |
491 |
0.89 |
| chr4_122998435_122998747 | 2.93 |
Mycl |
v-myc avian myelocytomatosis viral oncogene lung carcinoma derived |
289 |
0.87 |
| chr4_97582473_97584218 | 2.93 |
E130114P18Rik |
RIKEN cDNA E130114P18 gene |
1251 |
0.53 |
| chr2_116068937_116070512 | 2.91 |
G630016G05Rik |
RIKEN cDNA G630016G05 gene |
1756 |
0.28 |
| chr13_78446001_78446902 | 2.90 |
Gm31946 |
predicted gene, 31946 |
19203 |
0.18 |
| chr4_110290101_110291006 | 2.90 |
Elavl4 |
ELAV like RNA binding protein 4 |
281 |
0.95 |
| chr15_81936444_81938042 | 2.90 |
Csdc2 |
cold shock domain containing C2, RNA binding |
261 |
0.82 |
| chr13_83724722_83725570 | 2.90 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
2960 |
0.17 |
| chr10_36509078_36510052 | 2.88 |
Hs3st5 |
heparan sulfate (glucosamine) 3-O-sulfotransferase 5 |
2158 |
0.44 |
| chr4_33926104_33927188 | 2.88 |
Cnr1 |
cannabinoid receptor 1 (brain) |
444 |
0.88 |
| chr11_96301598_96303082 | 2.88 |
Hoxb5 |
homeobox B5 |
996 |
0.26 |
| chr10_109008310_109009456 | 2.88 |
Syt1 |
synaptotagmin I |
217 |
0.96 |
| chr7_99272446_99273539 | 2.88 |
Map6 |
microtubule-associated protein 6 |
3860 |
0.15 |
| chr7_62366760_62367184 | 2.87 |
Magel2 |
melanoma antigen, family L, 2 |
10038 |
0.18 |
| chr18_37216786_37217023 | 2.87 |
Gm10544 |
predicted gene 10544 |
38382 |
0.08 |
| chr5_107498769_107499247 | 2.87 |
Btbd8 |
BTB (POZ) domain containing 8 |
1229 |
0.33 |
| chr6_55680133_55680881 | 2.87 |
Neurod6 |
neurogenic differentiation 6 |
756 |
0.69 |
| chr4_78209143_78209724 | 2.87 |
Ptprd |
protein tyrosine phosphatase, receptor type, D |
2306 |
0.29 |
| chr2_77701567_77703605 | 2.85 |
Zfp385b |
zinc finger protein 385B |
686 |
0.8 |
| chr13_83719687_83720586 | 2.84 |
C130071C03Rik |
RIKEN cDNA C130071C03 gene |
1245 |
0.36 |
| chr2_137808388_137808680 | 2.84 |
Gm14062 |
predicted gene 14062 |
68297 |
0.14 |
| chr13_44946654_44947258 | 2.83 |
Dtnbp1 |
dystrobrevin binding protein 1 |
188 |
0.96 |
| chr7_16948034_16948663 | 2.83 |
Pnmal2 |
PNMA-like 2 |
3666 |
0.11 |
| chr13_83984481_83985348 | 2.83 |
Gm4241 |
predicted gene 4241 |
3077 |
0.26 |
| chr3_45384939_45386122 | 2.83 |
Pcdh10 |
protocadherin 10 |
2897 |
0.22 |
| chr6_25686466_25687390 | 2.82 |
Gpr37 |
G protein-coupled receptor 37 |
2864 |
0.38 |
| chr12_29534253_29535510 | 2.81 |
Gm20208 |
predicted gene, 20208 |
10 |
0.8 |
| chr5_43236846_43237650 | 2.81 |
Cpeb2 |
cytoplasmic polyadenylation element binding protein 2 |
67 |
0.96 |
| chr9_4794566_4795419 | 2.81 |
Gria4 |
glutamate receptor, ionotropic, AMPA4 (alpha 4) |
527 |
0.88 |
| chr2_158609356_158610186 | 2.80 |
Gm14204 |
predicted gene 14204 |
819 |
0.36 |
| chr3_63897818_63898430 | 2.80 |
Plch1 |
phospholipase C, eta 1 |
1249 |
0.38 |
| chr10_32887518_32887881 | 2.79 |
Nkain2 |
Na+/K+ transporting ATPase interacting 2 |
1997 |
0.43 |
| chr2_6883618_6884699 | 2.79 |
Gm13389 |
predicted gene 13389 |
112 |
0.85 |
| chr1_84693950_84694415 | 2.78 |
Mir5126 |
microRNA 5126 |
1657 |
0.28 |
| chr4_110281444_110282224 | 2.77 |
Elavl4 |
ELAV like RNA binding protein 4 |
4782 |
0.33 |
| chr3_5222045_5223052 | 2.76 |
Zfhx4 |
zinc finger homeodomain 4 |
1043 |
0.46 |
| chr14_75963198_75963625 | 2.75 |
Kctd4 |
potassium channel tetramerisation domain containing 4 |
8402 |
0.18 |
| chr4_11964013_11964651 | 2.70 |
Pdp1 |
pyruvate dehyrogenase phosphatase catalytic subunit 1 |
1489 |
0.31 |
| chr2_73773212_73773983 | 2.70 |
Chn1 |
chimerin 1 |
1446 |
0.44 |
| chr12_53249820_53250007 | 2.70 |
Npas3 |
neuronal PAS domain protein 3 |
1236 |
0.6 |
| chr10_70597271_70597504 | 2.70 |
Phyhipl |
phytanoyl-CoA hydroxylase interacting protein-like |
1695 |
0.45 |
| chr1_89582272_89582423 | 2.69 |
Agap1 |
ArfGAP with GTPase domain, ankyrin repeat and PH domain 1 |
2111 |
0.3 |
| chr2_136062179_136062393 | 2.67 |
Lamp5 |
lysosomal-associated membrane protein family, member 5 |
2855 |
0.3 |
| chr3_107039197_107039974 | 2.66 |
AI504432 |
expressed sequence AI504432 |
81 |
0.96 |
| chr1_62709496_62710533 | 2.64 |
Nrp2 |
neuropilin 2 |
6312 |
0.18 |
| chrX_165324069_165324869 | 2.63 |
Glra2 |
glycine receptor, alpha 2 subunit |
2924 |
0.41 |
| chr10_110453550_110454045 | 2.63 |
Nav3 |
neuron navigator 3 |
2407 |
0.34 |
| chr2_65929929_65930575 | 2.63 |
Csrnp3 |
cysteine-serine-rich nuclear protein 3 |
115 |
0.97 |
Network of associatons between targets according to the STRING database.

Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.9 | 11.8 | GO:0097118 | neuroligin clustering involved in postsynaptic membrane assembly(GO:0097118) |
| 2.4 | 2.4 | GO:2000807 | regulation of synaptic vesicle clustering(GO:2000807) |
| 2.3 | 6.8 | GO:0072221 | distal convoluted tubule development(GO:0072025) DCT cell differentiation(GO:0072069) metanephric distal convoluted tubule development(GO:0072221) metanephric distal tubule development(GO:0072235) metanephric DCT cell differentiation(GO:0072240) |
| 2.2 | 6.7 | GO:0021538 | epithalamus development(GO:0021538) habenula development(GO:0021986) |
| 2.1 | 6.2 | GO:0050923 | regulation of negative chemotaxis(GO:0050923) |
| 1.9 | 5.8 | GO:0046684 | response to pyrethroid(GO:0046684) |
| 1.8 | 7.3 | GO:0021615 | glossopharyngeal nerve morphogenesis(GO:0021615) |
| 1.8 | 5.4 | GO:1901078 | negative regulation of relaxation of muscle(GO:1901078) |
| 1.7 | 6.6 | GO:0051612 | negative regulation of neurotransmitter uptake(GO:0051581) regulation of serotonin uptake(GO:0051611) negative regulation of serotonin uptake(GO:0051612) |
| 1.6 | 4.8 | GO:0097155 | fasciculation of sensory neuron axon(GO:0097155) |
| 1.6 | 9.3 | GO:0045213 | neurotransmitter receptor metabolic process(GO:0045213) |
| 1.4 | 6.9 | GO:0060916 | mesenchymal cell proliferation involved in lung development(GO:0060916) |
| 1.3 | 16.2 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 1.3 | 3.8 | GO:0075509 | receptor-mediated endocytosis of virus by host cell(GO:0019065) endocytosis involved in viral entry into host cell(GO:0075509) |
| 1.3 | 5.0 | GO:0021698 | cerebellar cortex structural organization(GO:0021698) |
| 1.2 | 11.6 | GO:0045176 | apical protein localization(GO:0045176) |
| 1.1 | 4.6 | GO:0021869 | forebrain ventricular zone progenitor cell division(GO:0021869) |
| 1.1 | 12.4 | GO:0071625 | vocalization behavior(GO:0071625) |
| 1.1 | 2.2 | GO:2000611 | positive regulation of thyroid hormone generation(GO:2000611) |
| 1.1 | 2.2 | GO:0048880 | sensory system development(GO:0048880) |
| 1.1 | 3.2 | GO:0035887 | aortic smooth muscle cell differentiation(GO:0035887) |
| 1.1 | 3.2 | GO:0072318 | clathrin coat disassembly(GO:0072318) |
| 1.0 | 3.1 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 1.0 | 9.3 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 1.0 | 3.1 | GO:0072051 | juxtaglomerular apparatus development(GO:0072051) |
| 1.0 | 3.0 | GO:1900245 | positive regulation of MDA-5 signaling pathway(GO:1900245) |
| 1.0 | 7.0 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 1.0 | 2.0 | GO:0035262 | gonad morphogenesis(GO:0035262) |
| 1.0 | 2.9 | GO:0033563 | dorsal/ventral axon guidance(GO:0033563) |
| 0.9 | 3.7 | GO:0031117 | positive regulation of microtubule depolymerization(GO:0031117) |
| 0.9 | 3.6 | GO:0033602 | negative regulation of dopamine secretion(GO:0033602) |
| 0.9 | 8.2 | GO:0021859 | pyramidal neuron differentiation(GO:0021859) |
| 0.9 | 3.6 | GO:0007258 | JUN phosphorylation(GO:0007258) |
| 0.9 | 3.6 | GO:0071681 | response to indole-3-methanol(GO:0071680) cellular response to indole-3-methanol(GO:0071681) |
| 0.9 | 4.3 | GO:2000481 | positive regulation of cAMP-dependent protein kinase activity(GO:2000481) |
| 0.9 | 3.4 | GO:0060155 | platelet dense granule organization(GO:0060155) |
| 0.8 | 4.2 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.8 | 0.8 | GO:0035910 | ascending aorta development(GO:0035905) ascending aorta morphogenesis(GO:0035910) |
| 0.8 | 3.4 | GO:0045900 | negative regulation of translational elongation(GO:0045900) |
| 0.8 | 4.2 | GO:0097105 | presynaptic membrane assembly(GO:0097105) |
| 0.8 | 2.5 | GO:0021586 | pons maturation(GO:0021586) |
| 0.8 | 5.8 | GO:0002091 | negative regulation of receptor internalization(GO:0002091) |
| 0.8 | 3.3 | GO:0034773 | histone H4-K20 trimethylation(GO:0034773) |
| 0.8 | 6.9 | GO:0090129 | positive regulation of synapse maturation(GO:0090129) |
| 0.7 | 2.2 | GO:0072017 | distal tubule development(GO:0072017) |
| 0.7 | 4.4 | GO:0048852 | diencephalon morphogenesis(GO:0048852) |
| 0.7 | 3.6 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.7 | 2.9 | GO:0060244 | negative regulation of cell proliferation involved in contact inhibition(GO:0060244) |
| 0.7 | 5.7 | GO:0021796 | cerebral cortex regionalization(GO:0021796) |
| 0.7 | 3.4 | GO:0010891 | negative regulation of sequestering of triglyceride(GO:0010891) |
| 0.7 | 2.0 | GO:0097476 | motor neuron migration(GO:0097475) spinal cord motor neuron migration(GO:0097476) |
| 0.7 | 2.7 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.7 | 3.4 | GO:0031339 | negative regulation of vesicle fusion(GO:0031339) |
| 0.7 | 5.4 | GO:0051967 | negative regulation of synaptic transmission, glutamatergic(GO:0051967) |
| 0.7 | 0.7 | GO:0021563 | glossopharyngeal nerve development(GO:0021563) |
| 0.7 | 2.0 | GO:1903375 | cerebral cortex tangential migration using cell-axon interactions(GO:0021824) gonadotrophin-releasing hormone neuronal migration to the hypothalamus(GO:0021828) hypothalamic tangential migration using cell-axon interactions(GO:0021856) facioacoustic ganglion development(GO:1903375) |
| 0.7 | 2.0 | GO:0021570 | rhombomere 4 development(GO:0021570) |
| 0.7 | 0.7 | GO:0098598 | vocal learning(GO:0042297) imitative learning(GO:0098596) learned vocalization behavior or vocal learning(GO:0098598) |
| 0.7 | 2.0 | GO:0021902 | commitment of neuronal cell to specific neuron type in forebrain(GO:0021902) |
| 0.6 | 11.0 | GO:0071475 | cellular hyperosmotic salinity response(GO:0071475) |
| 0.6 | 1.3 | GO:0021773 | striatal medium spiny neuron differentiation(GO:0021773) |
| 0.6 | 1.9 | GO:1901529 | positive regulation of anion channel activity(GO:1901529) |
| 0.6 | 2.5 | GO:0090394 | negative regulation of excitatory postsynaptic potential(GO:0090394) |
| 0.6 | 2.5 | GO:0006651 | diacylglycerol biosynthetic process(GO:0006651) |
| 0.6 | 3.1 | GO:0048170 | positive regulation of long-term neuronal synaptic plasticity(GO:0048170) |
| 0.6 | 3.1 | GO:0014816 | skeletal muscle satellite cell differentiation(GO:0014816) |
| 0.6 | 0.6 | GO:0003358 | noradrenergic neuron development(GO:0003358) |
| 0.6 | 1.9 | GO:0016198 | axon choice point recognition(GO:0016198) |
| 0.6 | 4.3 | GO:0019227 | neuronal action potential propagation(GO:0019227) action potential propagation(GO:0098870) |
| 0.6 | 1.9 | GO:0021979 | hypothalamus cell differentiation(GO:0021979) |
| 0.6 | 3.1 | GO:0090273 | regulation of somatostatin secretion(GO:0090273) |
| 0.6 | 0.6 | GO:0022028 | tangential migration from the subventricular zone to the olfactory bulb(GO:0022028) |
| 0.6 | 0.6 | GO:0090494 | catecholamine uptake(GO:0090493) dopamine uptake(GO:0090494) |
| 0.6 | 1.8 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.6 | 6.5 | GO:0021542 | dentate gyrus development(GO:0021542) |
| 0.6 | 1.2 | GO:0036515 | serotonergic neuron axon guidance(GO:0036515) |
| 0.6 | 4.6 | GO:2000980 | regulation of auditory receptor cell differentiation(GO:0045607) regulation of mechanoreceptor differentiation(GO:0045631) regulation of inner ear receptor cell differentiation(GO:2000980) |
| 0.6 | 2.3 | GO:0060164 | regulation of timing of neuron differentiation(GO:0060164) |
| 0.6 | 1.1 | GO:0021853 | cerebral cortex GABAergic interneuron migration(GO:0021853) interneuron migration(GO:1904936) |
| 0.6 | 2.3 | GO:0061743 | motor learning(GO:0061743) |
| 0.5 | 6.0 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 0.5 | 1.6 | GO:2000211 | regulation of glutamate metabolic process(GO:2000211) |
| 0.5 | 2.1 | GO:0000160 | phosphorelay signal transduction system(GO:0000160) |
| 0.5 | 2.1 | GO:0097154 | GABAergic neuron differentiation(GO:0097154) |
| 0.5 | 5.9 | GO:0008038 | neuron recognition(GO:0008038) |
| 0.5 | 1.6 | GO:1904059 | regulation of locomotor rhythm(GO:1904059) |
| 0.5 | 6.8 | GO:0001504 | neurotransmitter uptake(GO:0001504) |
| 0.5 | 2.6 | GO:0098828 | modulation of inhibitory postsynaptic potential(GO:0098828) |
| 0.5 | 1.5 | GO:0035696 | monocyte extravasation(GO:0035696) regulation of monocyte extravasation(GO:2000437) |
| 0.5 | 0.5 | GO:0042420 | dopamine catabolic process(GO:0042420) |
| 0.5 | 1.5 | GO:0070777 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.5 | 4.6 | GO:0031536 | positive regulation of exit from mitosis(GO:0031536) |
| 0.5 | 2.0 | GO:0060836 | lymphatic endothelial cell differentiation(GO:0060836) |
| 0.5 | 4.0 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.5 | 1.5 | GO:0072137 | condensed mesenchymal cell proliferation(GO:0072137) |
| 0.5 | 9.0 | GO:0001964 | startle response(GO:0001964) |
| 0.5 | 1.4 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.5 | 1.4 | GO:0006532 | aspartate biosynthetic process(GO:0006532) |
| 0.4 | 3.1 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.4 | 0.4 | GO:0060666 | dichotomous subdivision of terminal units involved in salivary gland branching(GO:0060666) |
| 0.4 | 0.9 | GO:0003357 | noradrenergic neuron differentiation(GO:0003357) |
| 0.4 | 0.9 | GO:0007208 | phospholipase C-activating serotonin receptor signaling pathway(GO:0007208) |
| 0.4 | 3.9 | GO:0060539 | diaphragm development(GO:0060539) |
| 0.4 | 0.9 | GO:0060300 | regulation of cytokine activity(GO:0060300) |
| 0.4 | 2.2 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.4 | 2.1 | GO:0021957 | corticospinal tract morphogenesis(GO:0021957) |
| 0.4 | 2.5 | GO:2001054 | negative regulation of mesenchymal cell apoptotic process(GO:2001054) |
| 0.4 | 1.2 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.4 | 4.1 | GO:0047497 | establishment of mitochondrion localization, microtubule-mediated(GO:0034643) mitochondrion transport along microtubule(GO:0047497) |
| 0.4 | 3.3 | GO:0071420 | cellular response to histamine(GO:0071420) |
| 0.4 | 2.4 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.4 | 0.8 | GO:0007386 | compartment pattern specification(GO:0007386) |
| 0.4 | 2.4 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.4 | 3.6 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.4 | 1.2 | GO:0098903 | regulation of membrane repolarization during action potential(GO:0098903) |
| 0.4 | 2.8 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.4 | 1.2 | GO:0051654 | establishment of mitochondrion localization(GO:0051654) |
| 0.4 | 2.3 | GO:0021684 | cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.4 | 0.4 | GO:0072106 | regulation of ureteric bud formation(GO:0072106) positive regulation of ureteric bud formation(GO:0072107) |
| 0.4 | 1.9 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.4 | 1.1 | GO:2001016 | positive regulation of skeletal muscle cell differentiation(GO:2001016) |
| 0.4 | 3.0 | GO:2000463 | positive regulation of excitatory postsynaptic potential(GO:2000463) |
| 0.4 | 9.4 | GO:0021954 | central nervous system neuron development(GO:0021954) |
| 0.4 | 0.4 | GO:0014056 | acetylcholine secretion, neurotransmission(GO:0014055) regulation of acetylcholine secretion, neurotransmission(GO:0014056) |
| 0.4 | 1.5 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.4 | 1.5 | GO:0030035 | microspike assembly(GO:0030035) |
| 0.4 | 0.7 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.4 | 0.4 | GO:0072076 | nephrogenic mesenchyme development(GO:0072076) |
| 0.4 | 1.5 | GO:0090043 | regulation of tubulin deacetylation(GO:0090043) |
| 0.4 | 0.4 | GO:0014016 | neuroblast differentiation(GO:0014016) |
| 0.4 | 0.7 | GO:0048702 | embryonic neurocranium morphogenesis(GO:0048702) |
| 0.4 | 1.4 | GO:0045876 | positive regulation of sister chromatid cohesion(GO:0045876) |
| 0.4 | 0.4 | GO:0021604 | cranial nerve structural organization(GO:0021604) |
| 0.4 | 1.1 | GO:0033564 | anterior/posterior axon guidance(GO:0033564) |
| 0.4 | 0.4 | GO:0035564 | regulation of kidney size(GO:0035564) |
| 0.4 | 5.0 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.4 | 3.5 | GO:0051968 | positive regulation of synaptic transmission, glutamatergic(GO:0051968) |
| 0.3 | 2.4 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.3 | 1.0 | GO:0097119 | postsynaptic density protein 95 clustering(GO:0097119) |
| 0.3 | 2.4 | GO:2000035 | regulation of stem cell division(GO:2000035) |
| 0.3 | 0.7 | GO:0014012 | peripheral nervous system axon regeneration(GO:0014012) |
| 0.3 | 0.3 | GO:0097374 | sensory neuron axon guidance(GO:0097374) |
| 0.3 | 0.7 | GO:0007196 | adenylate cyclase-inhibiting G-protein coupled glutamate receptor signaling pathway(GO:0007196) |
| 0.3 | 0.7 | GO:0097195 | pilomotor reflex(GO:0097195) |
| 0.3 | 15.0 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.3 | 2.0 | GO:0042473 | outer ear morphogenesis(GO:0042473) |
| 0.3 | 0.6 | GO:0070253 | somatostatin secretion(GO:0070253) |
| 0.3 | 8.1 | GO:0045773 | positive regulation of axon extension(GO:0045773) |
| 0.3 | 1.3 | GO:2000650 | negative regulation of sodium ion transmembrane transporter activity(GO:2000650) |
| 0.3 | 0.3 | GO:0060523 | prostate epithelial cord elongation(GO:0060523) |
| 0.3 | 3.2 | GO:0043374 | CD8-positive, alpha-beta T cell differentiation(GO:0043374) |
| 0.3 | 1.9 | GO:0030432 | peristalsis(GO:0030432) |
| 0.3 | 2.2 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.3 | 3.2 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.3 | 1.9 | GO:0060052 | neurofilament cytoskeleton organization(GO:0060052) |
| 0.3 | 1.9 | GO:0015816 | glycine transport(GO:0015816) |
| 0.3 | 0.9 | GO:0043587 | tongue morphogenesis(GO:0043587) |
| 0.3 | 1.2 | GO:0006538 | glutamate catabolic process(GO:0006538) |
| 0.3 | 0.9 | GO:0070634 | transepithelial ammonium transport(GO:0070634) |
| 0.3 | 0.6 | GO:0060406 | positive regulation of penile erection(GO:0060406) |
| 0.3 | 0.9 | GO:0035641 | locomotory exploration behavior(GO:0035641) |
| 0.3 | 0.9 | GO:0019732 | antifungal humoral response(GO:0019732) |
| 0.3 | 0.6 | GO:0060166 | olfactory pit development(GO:0060166) |
| 0.3 | 0.6 | GO:0016080 | synaptic vesicle targeting(GO:0016080) |
| 0.3 | 1.2 | GO:1904322 | response to forskolin(GO:1904321) cellular response to forskolin(GO:1904322) |
| 0.3 | 2.7 | GO:0070257 | positive regulation of mucus secretion(GO:0070257) |
| 0.3 | 1.8 | GO:0070571 | negative regulation of neuron projection regeneration(GO:0070571) |
| 0.3 | 1.7 | GO:0060371 | regulation of atrial cardiac muscle cell membrane depolarization(GO:0060371) |
| 0.3 | 0.9 | GO:0002087 | regulation of respiratory gaseous exchange by neurological system process(GO:0002087) |
| 0.3 | 0.6 | GO:0070100 | negative regulation of chemokine-mediated signaling pathway(GO:0070100) |
| 0.3 | 0.6 | GO:0021530 | spinal cord oligodendrocyte cell differentiation(GO:0021529) spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.3 | 4.6 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.3 | 0.9 | GO:0071492 | cellular response to UV-A(GO:0071492) |
| 0.3 | 4.7 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.3 | 0.8 | GO:0097503 | sialylation(GO:0097503) |
| 0.3 | 0.8 | GO:0042713 | sperm ejaculation(GO:0042713) |
| 0.3 | 0.8 | GO:0021978 | telencephalon regionalization(GO:0021978) |
| 0.3 | 19.2 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.3 | 24.0 | GO:0007612 | learning(GO:0007612) |
| 0.3 | 0.5 | GO:0051964 | negative regulation of synapse assembly(GO:0051964) |
| 0.3 | 1.1 | GO:0060480 | lung goblet cell differentiation(GO:0060480) |
| 0.3 | 0.8 | GO:1904491 | protein localization to ciliary transition zone(GO:1904491) |
| 0.3 | 0.5 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.3 | 0.8 | GO:1900086 | regulation of peptidyl-tyrosine autophosphorylation(GO:1900084) positive regulation of peptidyl-tyrosine autophosphorylation(GO:1900086) |
| 0.3 | 0.8 | GO:0060279 | positive regulation of ovulation(GO:0060279) |
| 0.3 | 2.1 | GO:0060732 | positive regulation of inositol phosphate biosynthetic process(GO:0060732) |
| 0.3 | 0.8 | GO:0034628 | nicotinamide nucleotide biosynthetic process from aspartate(GO:0019355) 'de novo' NAD biosynthetic process from aspartate(GO:0034628) |
| 0.3 | 1.6 | GO:2000480 | negative regulation of cAMP-dependent protein kinase activity(GO:2000480) |
| 0.3 | 0.5 | GO:0060648 | mammary gland bud morphogenesis(GO:0060648) |
| 0.3 | 0.8 | GO:0030167 | proteoglycan catabolic process(GO:0030167) |
| 0.3 | 1.0 | GO:0071276 | cellular response to cadmium ion(GO:0071276) |
| 0.3 | 5.0 | GO:0051491 | positive regulation of filopodium assembly(GO:0051491) |
| 0.3 | 1.0 | GO:1903966 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.2 | 0.7 | GO:0003139 | secondary heart field specification(GO:0003139) |
| 0.2 | 2.5 | GO:0033523 | histone H2B ubiquitination(GO:0033523) |
| 0.2 | 1.7 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 0.2 | 0.7 | GO:1902177 | positive regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902177) |
| 0.2 | 0.2 | GO:0061642 | chemoattraction of axon(GO:0061642) |
| 0.2 | 0.5 | GO:0021830 | interneuron migration from the subpallium to the cortex(GO:0021830) |
| 0.2 | 0.2 | GO:1904304 | regulation of gastro-intestinal system smooth muscle contraction(GO:1904304) |
| 0.2 | 0.7 | GO:2000327 | regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000325) positive regulation of ligand-dependent nuclear receptor transcription coactivator activity(GO:2000327) |
| 0.2 | 1.4 | GO:0060179 | male mating behavior(GO:0060179) |
| 0.2 | 0.5 | GO:0003150 | muscular septum morphogenesis(GO:0003150) |
| 0.2 | 1.2 | GO:0034638 | phosphatidylcholine catabolic process(GO:0034638) |
| 0.2 | 0.5 | GO:0021912 | regulation of transcription from RNA polymerase II promoter involved in spinal cord motor neuron fate specification(GO:0021912) |
| 0.2 | 1.4 | GO:0043615 | astrocyte cell migration(GO:0043615) |
| 0.2 | 0.7 | GO:0015808 | L-alanine transport(GO:0015808) |
| 0.2 | 0.4 | GO:0070858 | negative regulation of bile acid biosynthetic process(GO:0070858) negative regulation of bile acid metabolic process(GO:1904252) |
| 0.2 | 0.4 | GO:1902498 | regulation of protein autoubiquitination(GO:1902498) |
| 0.2 | 0.7 | GO:0048505 | regulation of timing of cell differentiation(GO:0048505) |
| 0.2 | 0.4 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.2 | 1.3 | GO:1901621 | negative regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901621) |
| 0.2 | 0.4 | GO:0034182 | regulation of maintenance of sister chromatid cohesion(GO:0034091) regulation of maintenance of mitotic sister chromatid cohesion(GO:0034182) |
| 0.2 | 0.4 | GO:2000049 | positive regulation of cell-cell adhesion mediated by cadherin(GO:2000049) |
| 0.2 | 0.6 | GO:1902775 | mitochondrial large ribosomal subunit assembly(GO:1902775) |
| 0.2 | 0.8 | GO:0060437 | lung growth(GO:0060437) |
| 0.2 | 2.3 | GO:0048714 | positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.2 | 0.8 | GO:0001957 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.2 | 0.4 | GO:0060687 | regulation of branching involved in prostate gland morphogenesis(GO:0060687) |
| 0.2 | 0.6 | GO:0051684 | maintenance of Golgi location(GO:0051684) |
| 0.2 | 4.5 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.2 | 0.6 | GO:0045964 | positive regulation of catecholamine metabolic process(GO:0045915) positive regulation of dopamine metabolic process(GO:0045964) |
| 0.2 | 1.8 | GO:0061213 | positive regulation of mesonephros development(GO:0061213) |
| 0.2 | 1.0 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.2 | 0.4 | GO:0022038 | corpus callosum development(GO:0022038) |
| 0.2 | 0.6 | GO:0010766 | negative regulation of sodium ion transport(GO:0010766) |
| 0.2 | 3.1 | GO:0010758 | regulation of macrophage chemotaxis(GO:0010758) |
| 0.2 | 0.6 | GO:0032474 | otolith morphogenesis(GO:0032474) |
| 0.2 | 0.4 | GO:0031915 | positive regulation of synaptic plasticity(GO:0031915) |
| 0.2 | 0.8 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 0.2 | 0.4 | GO:0035609 | C-terminal protein deglutamylation(GO:0035609) |
| 0.2 | 0.7 | GO:0021895 | cerebral cortex neuron differentiation(GO:0021895) |
| 0.2 | 0.6 | GO:2000850 | negative regulation of steroid hormone secretion(GO:2000832) negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.2 | 0.4 | GO:0090361 | platelet-derived growth factor production(GO:0090360) regulation of platelet-derived growth factor production(GO:0090361) |
| 0.2 | 0.2 | GO:0016539 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.2 | 0.5 | GO:0060020 | Bergmann glial cell differentiation(GO:0060020) |
| 0.2 | 1.5 | GO:0021702 | cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.2 | 0.7 | GO:0021871 | forebrain regionalization(GO:0021871) |
| 0.2 | 0.4 | GO:0021692 | cerebellar Purkinje cell layer morphogenesis(GO:0021692) |
| 0.2 | 0.7 | GO:1903215 | negative regulation of protein targeting to mitochondrion(GO:1903215) |
| 0.2 | 0.7 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.2 | 1.4 | GO:0045075 | interleukin-12 biosynthetic process(GO:0042090) regulation of interleukin-12 biosynthetic process(GO:0045075) |
| 0.2 | 0.2 | GO:0035022 | positive regulation of Rac protein signal transduction(GO:0035022) |
| 0.2 | 0.3 | GO:2000173 | negative regulation of branching morphogenesis of a nerve(GO:2000173) |
| 0.2 | 1.3 | GO:0007625 | grooming behavior(GO:0007625) |
| 0.2 | 0.2 | GO:2000828 | regulation of parathyroid hormone secretion(GO:2000828) |
| 0.2 | 0.5 | GO:1903421 | regulation of synaptic vesicle recycling(GO:1903421) |
| 0.2 | 0.3 | GO:0045113 | regulation of integrin biosynthetic process(GO:0045113) |
| 0.2 | 0.5 | GO:0021794 | thalamus development(GO:0021794) |
| 0.2 | 0.5 | GO:1902309 | negative regulation of peptidyl-serine dephosphorylation(GO:1902309) |
| 0.2 | 0.8 | GO:0031290 | retinal ganglion cell axon guidance(GO:0031290) |
| 0.2 | 0.3 | GO:0061470 | T follicular helper cell differentiation(GO:0061470) |
| 0.2 | 0.2 | GO:0014042 | positive regulation of neuron maturation(GO:0014042) |
| 0.2 | 0.5 | GO:0015755 | fructose transport(GO:0015755) |
| 0.2 | 0.6 | GO:0031937 | positive regulation of chromatin silencing(GO:0031937) |
| 0.2 | 0.9 | GO:0060850 | regulation of transcription involved in cell fate commitment(GO:0060850) |
| 0.2 | 0.5 | GO:0070649 | formin-nucleated actin cable assembly(GO:0070649) |
| 0.1 | 0.7 | GO:0010739 | positive regulation of protein kinase A signaling(GO:0010739) |
| 0.1 | 0.1 | GO:0070640 | vitamin D3 metabolic process(GO:0070640) |
| 0.1 | 0.4 | GO:0030327 | prenylated protein catabolic process(GO:0030327) |
| 0.1 | 0.1 | GO:1903261 | regulation of serine phosphorylation of STAT3 protein(GO:1903261) |
| 0.1 | 2.0 | GO:0007617 | mating behavior(GO:0007617) multi-organism reproductive behavior(GO:0044705) |
| 0.1 | 0.1 | GO:1902308 | regulation of peptidyl-serine dephosphorylation(GO:1902308) |
| 0.1 | 0.4 | GO:0003419 | growth plate cartilage chondrocyte proliferation(GO:0003419) |
| 0.1 | 1.0 | GO:1902855 | regulation of nonmotile primary cilium assembly(GO:1902855) |
| 0.1 | 0.3 | GO:0051562 | negative regulation of mitochondrial calcium ion concentration(GO:0051562) |
| 0.1 | 0.4 | GO:0060178 | regulation of exocyst localization(GO:0060178) |
| 0.1 | 0.6 | GO:0030091 | protein repair(GO:0030091) |
| 0.1 | 4.2 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.1 | 0.3 | GO:0009629 | response to gravity(GO:0009629) |
| 0.1 | 0.7 | GO:0032099 | negative regulation of appetite(GO:0032099) |
| 0.1 | 0.3 | GO:0042891 | antibiotic transport(GO:0042891) |
| 0.1 | 0.4 | GO:0031585 | regulation of inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0031585) |
| 0.1 | 0.4 | GO:0034983 | peptidyl-lysine deacetylation(GO:0034983) |
| 0.1 | 0.1 | GO:0048669 | collateral sprouting in absence of injury(GO:0048669) |
| 0.1 | 0.3 | GO:2001013 | epithelial cell proliferation involved in renal tubule morphogenesis(GO:2001013) |
| 0.1 | 1.0 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.1 | 0.3 | GO:0039663 | fusion of virus membrane with host plasma membrane(GO:0019064) membrane fusion involved in viral entry into host cell(GO:0039663) multi-organism membrane fusion(GO:0044800) |
| 0.1 | 0.4 | GO:2000686 | regulation of rubidium ion transmembrane transporter activity(GO:2000686) |
| 0.1 | 0.5 | GO:0090037 | positive regulation of protein kinase C signaling(GO:0090037) |
| 0.1 | 0.9 | GO:1901538 | DNA methylation involved in embryo development(GO:0043045) changes to DNA methylation involved in embryo development(GO:1901538) |
| 0.1 | 0.3 | GO:0071798 | response to prostaglandin D(GO:0071798) cellular response to prostaglandin D stimulus(GO:0071799) |
| 0.1 | 2.1 | GO:0019228 | neuronal action potential(GO:0019228) |
| 0.1 | 1.3 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.1 | 1.4 | GO:0007616 | long-term memory(GO:0007616) |
| 0.1 | 0.9 | GO:0048245 | eosinophil chemotaxis(GO:0048245) |
| 0.1 | 0.5 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.1 | 0.4 | GO:0006177 | GMP biosynthetic process(GO:0006177) |
| 0.1 | 0.5 | GO:0072393 | microtubule anchoring at microtubule organizing center(GO:0072393) |
| 0.1 | 0.5 | GO:0072697 | protein localization to cell cortex(GO:0072697) |
| 0.1 | 0.6 | GO:0009256 | 10-formyltetrahydrofolate metabolic process(GO:0009256) |
| 0.1 | 0.4 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.1 | 0.2 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.1 | 1.8 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.1 | 0.4 | GO:0045074 | interleukin-10 biosynthetic process(GO:0042091) regulation of interleukin-10 biosynthetic process(GO:0045074) |
| 0.1 | 0.5 | GO:0010572 | positive regulation of platelet activation(GO:0010572) |
| 0.1 | 0.2 | GO:0018197 | peptidyl-aspartic acid modification(GO:0018197) |
| 0.1 | 0.6 | GO:0061590 | calcium activated phospholipid scrambling(GO:0061588) calcium activated phosphatidylcholine scrambling(GO:0061590) calcium activated galactosylceramide scrambling(GO:0061591) |
| 0.1 | 2.2 | GO:0030901 | midbrain development(GO:0030901) |
| 0.1 | 0.6 | GO:0021670 | lateral ventricle development(GO:0021670) |
| 0.1 | 0.4 | GO:0038128 | ERBB2 signaling pathway(GO:0038128) |
| 0.1 | 0.1 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) |
| 0.1 | 0.2 | GO:0071698 | olfactory placode formation(GO:0030910) olfactory placode development(GO:0071698) olfactory placode morphogenesis(GO:0071699) |
| 0.1 | 0.1 | GO:0046881 | positive regulation of follicle-stimulating hormone secretion(GO:0046881) |
| 0.1 | 0.8 | GO:0042760 | very long-chain fatty acid catabolic process(GO:0042760) |
| 0.1 | 0.1 | GO:0060762 | regulation of branching involved in mammary gland duct morphogenesis(GO:0060762) |
| 0.1 | 3.2 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 0.6 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.1 | 0.7 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 1.4 | GO:0021766 | hippocampus development(GO:0021766) |
| 0.1 | 0.6 | GO:0097343 | ripoptosome assembly(GO:0097343) ripoptosome assembly involved in necroptotic process(GO:1901026) |
| 0.1 | 0.3 | GO:0032066 | nucleolus to nucleoplasm transport(GO:0032066) |
| 0.1 | 0.2 | GO:0038026 | reelin-mediated signaling pathway(GO:0038026) |
| 0.1 | 4.7 | GO:0050905 | neuromuscular process(GO:0050905) |
| 0.1 | 0.2 | GO:1900164 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of transcription from RNA polymerase II promoter involved in determination of left/right symmetry(GO:1900094) nodal signaling pathway involved in determination of lateral mesoderm left/right asymmetry(GO:1900164) |
| 0.1 | 8.4 | GO:0097485 | neuron projection guidance(GO:0097485) |
| 0.1 | 0.2 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.1 | 0.1 | GO:1900619 | acetylcholine metabolic process(GO:0008291) acetate ester metabolic process(GO:1900619) |
| 0.1 | 0.3 | GO:0016560 | protein import into peroxisome matrix, docking(GO:0016560) |
| 0.1 | 1.5 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.1 | 0.5 | GO:0030205 | dermatan sulfate metabolic process(GO:0030205) |
| 0.1 | 2.2 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.1 | 1.4 | GO:1903831 | acetylcholine receptor signaling pathway(GO:0095500) postsynaptic signal transduction(GO:0098926) signal transduction involved in cellular response to ammonium ion(GO:1903831) response to acetylcholine(GO:1905144) cellular response to acetylcholine(GO:1905145) |
| 0.1 | 0.1 | GO:0043553 | negative regulation of phosphatidylinositol 3-kinase activity(GO:0043553) |
| 0.1 | 0.2 | GO:0031666 | positive regulation of lipopolysaccharide-mediated signaling pathway(GO:0031666) |
| 0.1 | 0.3 | GO:0071242 | cellular response to ammonium ion(GO:0071242) |
| 0.1 | 0.2 | GO:1990000 | amyloid fibril formation(GO:1990000) |
| 0.1 | 0.2 | GO:0001922 | B-1 B cell homeostasis(GO:0001922) |
| 0.1 | 0.1 | GO:2000172 | regulation of branching morphogenesis of a nerve(GO:2000172) |
| 0.1 | 0.2 | GO:0034047 | regulation of protein phosphatase type 2A activity(GO:0034047) |
| 0.1 | 0.2 | GO:2000074 | regulation of type B pancreatic cell development(GO:2000074) |
| 0.1 | 0.2 | GO:0034729 | histone H3-K79 methylation(GO:0034729) |
| 0.1 | 0.3 | GO:0010986 | positive regulation of lipoprotein particle clearance(GO:0010986) |
| 0.1 | 0.2 | GO:0032058 | positive regulation of translational initiation in response to stress(GO:0032058) |
| 0.1 | 0.2 | GO:0010360 | negative regulation of anion channel activity(GO:0010360) |
| 0.1 | 0.3 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.1 | 0.7 | GO:0072010 | renal filtration cell differentiation(GO:0061318) glomerular epithelium development(GO:0072010) glomerular visceral epithelial cell differentiation(GO:0072112) glomerular epithelial cell differentiation(GO:0072311) |
| 0.1 | 0.1 | GO:1902866 | regulation of retina development in camera-type eye(GO:1902866) |
| 0.1 | 0.3 | GO:0015889 | cobalamin transport(GO:0015889) |
| 0.1 | 0.3 | GO:0046167 | glycerol-3-phosphate biosynthetic process(GO:0046167) |
| 0.1 | 0.2 | GO:0002572 | pro-T cell differentiation(GO:0002572) |
| 0.1 | 0.4 | GO:0050932 | regulation of pigment cell differentiation(GO:0050932) positive regulation of pigment cell differentiation(GO:0050942) |
| 0.1 | 0.9 | GO:0055062 | phosphate ion homeostasis(GO:0055062) trivalent inorganic anion homeostasis(GO:0072506) |
| 0.1 | 0.3 | GO:0030210 | heparin biosynthetic process(GO:0030210) |
| 0.1 | 0.7 | GO:0045603 | positive regulation of endothelial cell differentiation(GO:0045603) |
| 0.1 | 0.1 | GO:2000618 | regulation of histone H4-K16 acetylation(GO:2000618) |
| 0.1 | 0.2 | GO:0098904 | regulation of AV node cell action potential(GO:0098904) |
| 0.1 | 0.4 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.1 | 0.4 | GO:1904659 | glucose transmembrane transport(GO:1904659) |
| 0.1 | 0.2 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 0.2 | GO:0046985 | positive regulation of hemoglobin biosynthetic process(GO:0046985) |
| 0.1 | 0.7 | GO:0098868 | endochondral bone growth(GO:0003416) bone growth(GO:0098868) |
| 0.1 | 0.1 | GO:1902996 | neurofibrillary tangle assembly(GO:1902988) regulation of neurofibrillary tangle assembly(GO:1902996) |
| 0.1 | 0.4 | GO:1901491 | negative regulation of lymphangiogenesis(GO:1901491) |
| 0.1 | 0.3 | GO:0036315 | cellular response to sterol(GO:0036315) |
| 0.1 | 0.3 | GO:0032484 | Ral protein signal transduction(GO:0032484) |
| 0.1 | 0.1 | GO:0060478 | acrosomal vesicle exocytosis(GO:0060478) |
| 0.1 | 0.1 | GO:1904139 | microglial cell migration(GO:1904124) regulation of microglial cell migration(GO:1904139) |
| 0.1 | 0.9 | GO:0007342 | fusion of sperm to egg plasma membrane(GO:0007342) |
| 0.1 | 0.1 | GO:0048715 | negative regulation of oligodendrocyte differentiation(GO:0048715) |
| 0.1 | 0.3 | GO:0014051 | gamma-aminobutyric acid secretion(GO:0014051) |
| 0.1 | 0.3 | GO:0007296 | vitellogenesis(GO:0007296) |
| 0.1 | 0.3 | GO:0051823 | regulation of synapse structural plasticity(GO:0051823) |
| 0.1 | 0.5 | GO:0045086 | positive regulation of interleukin-2 biosynthetic process(GO:0045086) |
| 0.1 | 0.4 | GO:0043951 | negative regulation of cAMP-mediated signaling(GO:0043951) |
| 0.1 | 0.1 | GO:0043465 | regulation of fermentation(GO:0043465) regulation of NAD metabolic process(GO:1902688) regulation of glucose catabolic process to lactate via pyruvate(GO:1904023) |
| 0.1 | 0.1 | GO:2000286 | receptor internalization involved in canonical Wnt signaling pathway(GO:2000286) |
| 0.1 | 0.1 | GO:0050968 | detection of chemical stimulus involved in sensory perception of pain(GO:0050968) |
| 0.1 | 0.4 | GO:0032596 | protein transport into membrane raft(GO:0032596) |
| 0.1 | 0.6 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.1 | 0.2 | GO:0051365 | cellular response to potassium ion starvation(GO:0051365) |
| 0.1 | 0.2 | GO:0002315 | marginal zone B cell differentiation(GO:0002315) |
| 0.1 | 0.4 | GO:0032291 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.1 | 0.2 | GO:0007290 | spermatid nucleus elongation(GO:0007290) |
| 0.1 | 0.2 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 0.1 | 0.2 | GO:0060075 | regulation of resting membrane potential(GO:0060075) |
| 0.1 | 0.2 | GO:0036118 | hyaluranon cable assembly(GO:0036118) regulation of hyaluranon cable assembly(GO:1900104) positive regulation of hyaluranon cable assembly(GO:1900106) |
| 0.1 | 0.2 | GO:0090310 | negative regulation of methylation-dependent chromatin silencing(GO:0090310) |
| 0.1 | 1.0 | GO:0030033 | microvillus assembly(GO:0030033) microvillus organization(GO:0032528) |
| 0.1 | 0.2 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.1 | 1.1 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) |
| 0.1 | 0.5 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.1 | 0.6 | GO:0007288 | sperm axoneme assembly(GO:0007288) |
| 0.1 | 0.2 | GO:0002322 | B cell proliferation involved in immune response(GO:0002322) |
| 0.1 | 0.5 | GO:0070995 | NADPH oxidation(GO:0070995) |
| 0.1 | 0.3 | GO:0035428 | hexose transmembrane transport(GO:0035428) |
| 0.1 | 0.3 | GO:0009838 | abscission(GO:0009838) |
| 0.1 | 0.4 | GO:0007216 | G-protein coupled glutamate receptor signaling pathway(GO:0007216) |
| 0.1 | 0.2 | GO:0060355 | positive regulation of cell adhesion molecule production(GO:0060355) |
| 0.1 | 0.3 | GO:2001206 | positive regulation of osteoclast development(GO:2001206) |
| 0.1 | 4.8 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 0.7 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.1 | 0.3 | GO:0045076 | regulation of interleukin-2 biosynthetic process(GO:0045076) |
| 0.1 | 0.3 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.1 | 0.1 | GO:2000599 | regulation of cyclin catabolic process(GO:2000598) negative regulation of cyclin catabolic process(GO:2000599) |
| 0.1 | 0.3 | GO:0015822 | ornithine transport(GO:0015822) L-ornithine transmembrane transport(GO:1903352) |
| 0.1 | 0.9 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
| 0.1 | 0.3 | GO:0046541 | saliva secretion(GO:0046541) |
| 0.1 | 0.3 | GO:0035989 | tendon development(GO:0035989) |
| 0.1 | 0.1 | GO:0060684 | epithelial-mesenchymal cell signaling(GO:0060684) |
| 0.1 | 0.1 | GO:0035483 | gastric motility(GO:0035482) gastric emptying(GO:0035483) |
| 0.1 | 0.2 | GO:0061154 | endothelial tube morphogenesis(GO:0061154) |
| 0.1 | 0.3 | GO:0014744 | positive regulation of muscle adaptation(GO:0014744) |
| 0.1 | 0.3 | GO:0034969 | histone arginine methylation(GO:0034969) |
| 0.1 | 0.1 | GO:0035964 | COPI-coated vesicle budding(GO:0035964) |
| 0.1 | 0.1 | GO:0018992 | germ-line sex determination(GO:0018992) |
| 0.1 | 0.2 | GO:0045657 | positive regulation of monocyte differentiation(GO:0045657) |
| 0.1 | 0.1 | GO:0072553 | terminal button organization(GO:0072553) |
| 0.1 | 0.1 | GO:1903054 | negative regulation of extracellular matrix organization(GO:1903054) |
| 0.1 | 0.1 | GO:0014873 | response to muscle activity involved in regulation of muscle adaptation(GO:0014873) |
| 0.1 | 0.2 | GO:1904217 | regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904217) positive regulation of CDP-diacylglycerol-serine O-phosphatidyltransferase activity(GO:1904219) regulation of serine C-palmitoyltransferase activity(GO:1904220) positive regulation of serine C-palmitoyltransferase activity(GO:1904222) |
| 0.1 | 0.3 | GO:0070940 | dephosphorylation of RNA polymerase II C-terminal domain(GO:0070940) |
| 0.1 | 0.3 | GO:0080182 | histone H3-K4 trimethylation(GO:0080182) |
| 0.1 | 0.2 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.1 | GO:0021546 | rhombomere development(GO:0021546) |
| 0.1 | 0.1 | GO:0071675 | regulation of mononuclear cell migration(GO:0071675) |
| 0.1 | 0.8 | GO:0061462 | protein localization to lysosome(GO:0061462) |
| 0.1 | 0.1 | GO:0042053 | regulation of dopamine metabolic process(GO:0042053) |
| 0.1 | 0.1 | GO:0071502 | cellular response to temperature stimulus(GO:0071502) |
| 0.1 | 0.1 | GO:1904587 | glycoprotein ERAD pathway(GO:0097466) response to glycoprotein(GO:1904587) |
| 0.1 | 0.4 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 0.4 | GO:0033148 | positive regulation of intracellular estrogen receptor signaling pathway(GO:0033148) |
| 0.1 | 0.6 | GO:2000300 | regulation of synaptic vesicle transport(GO:1902803) regulation of synaptic vesicle exocytosis(GO:2000300) |
| 0.1 | 0.1 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 0.1 | GO:0055099 | response to high density lipoprotein particle(GO:0055099) |
| 0.1 | 0.2 | GO:0045332 | phospholipid translocation(GO:0045332) |
| 0.1 | 0.4 | GO:0019800 | peptide cross-linking via chondroitin 4-sulfate glycosaminoglycan(GO:0019800) |
| 0.1 | 0.4 | GO:0032094 | response to food(GO:0032094) |
| 0.1 | 0.7 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.1 | 0.1 | GO:0021938 | smoothened signaling pathway involved in regulation of cerebellar granule cell precursor cell proliferation(GO:0021938) |
| 0.1 | 0.2 | GO:0048570 | notochord morphogenesis(GO:0048570) |
| 0.1 | 0.1 | GO:0090240 | positive regulation of histone H4 acetylation(GO:0090240) |
| 0.1 | 0.2 | GO:0019859 | pyrimidine nucleobase catabolic process(GO:0006208) thymine catabolic process(GO:0006210) thymine metabolic process(GO:0019859) |
| 0.1 | 0.2 | GO:0015812 | gamma-aminobutyric acid transport(GO:0015812) |
| 0.1 | 0.2 | GO:0032687 | negative regulation of interferon-alpha production(GO:0032687) |
| 0.1 | 0.1 | GO:0043490 | malate-aspartate shuttle(GO:0043490) |
| 0.1 | 0.1 | GO:0070666 | mast cell proliferation(GO:0070662) regulation of mast cell proliferation(GO:0070666) positive regulation of mast cell proliferation(GO:0070668) |
| 0.1 | 0.1 | GO:2000680 | regulation of rubidium ion transport(GO:2000680) |
| 0.1 | 0.3 | GO:0090209 | negative regulation of triglyceride metabolic process(GO:0090209) |
| 0.1 | 0.1 | GO:0097011 | cellular response to granulocyte macrophage colony-stimulating factor stimulus(GO:0097011) |
| 0.1 | 0.3 | GO:0035418 | protein localization to synapse(GO:0035418) |
| 0.1 | 0.1 | GO:1905098 | negative regulation of guanyl-nucleotide exchange factor activity(GO:1905098) |
| 0.1 | 0.1 | GO:0001831 | trophectodermal cellular morphogenesis(GO:0001831) |
| 0.1 | 0.5 | GO:0006228 | UTP biosynthetic process(GO:0006228) |
| 0.1 | 0.1 | GO:2000418 | positive regulation of eosinophil migration(GO:2000418) |
| 0.1 | 0.4 | GO:0090051 | negative regulation of cell migration involved in sprouting angiogenesis(GO:0090051) |
| 0.1 | 0.1 | GO:0048861 | leukemia inhibitory factor signaling pathway(GO:0048861) |
| 0.1 | 0.1 | GO:0038095 | Fc-epsilon receptor signaling pathway(GO:0038095) |
| 0.1 | 0.3 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.1 | 0.2 | GO:1901301 | regulation of cargo loading into COPII-coated vesicle(GO:1901301) |
| 0.1 | 0.7 | GO:0045956 | positive regulation of calcium ion-dependent exocytosis(GO:0045956) |
| 0.1 | 0.9 | GO:0009649 | entrainment of circadian clock(GO:0009649) |
| 0.1 | 0.4 | GO:0048490 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.1 | 0.2 | GO:2000019 | negative regulation of male gonad development(GO:2000019) |
| 0.1 | 0.1 | GO:0098902 | regulation of membrane depolarization during action potential(GO:0098902) |
| 0.1 | 0.6 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 0.2 | GO:0035766 | cell chemotaxis to fibroblast growth factor(GO:0035766) endothelial cell chemotaxis to fibroblast growth factor(GO:0035768) regulation of cell chemotaxis to fibroblast growth factor(GO:1904847) regulation of endothelial cell chemotaxis to fibroblast growth factor(GO:2000544) |
| 0.1 | 0.1 | GO:2001170 | negative regulation of ATP biosynthetic process(GO:2001170) |
| 0.1 | 0.1 | GO:2000047 | regulation of cell-cell adhesion mediated by cadherin(GO:2000047) |
| 0.1 | 0.2 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.1 | 0.4 | GO:0032986 | nucleosome disassembly(GO:0006337) protein-DNA complex disassembly(GO:0032986) |
| 0.1 | 0.7 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 0.2 | GO:0038165 | oncostatin-M-mediated signaling pathway(GO:0038165) |
| 0.1 | 0.1 | GO:0061196 | fungiform papilla development(GO:0061196) |
| 0.1 | 0.2 | GO:0002913 | positive regulation of T cell anergy(GO:0002669) positive regulation of lymphocyte anergy(GO:0002913) |
| 0.1 | 0.2 | GO:0070814 | hydrogen sulfide biosynthetic process(GO:0070814) |
| 0.1 | 0.2 | GO:0006362 | transcription elongation from RNA polymerase I promoter(GO:0006362) |
| 0.1 | 0.1 | GO:0055118 | negative regulation of cardiac muscle contraction(GO:0055118) |
| 0.1 | 0.3 | GO:0048539 | bone marrow development(GO:0048539) |
| 0.1 | 0.3 | GO:0006528 | asparagine metabolic process(GO:0006528) |
| 0.1 | 0.1 | GO:0090135 | actin filament branching(GO:0090135) |
| 0.0 | 0.1 | GO:0035434 | copper ion transmembrane transport(GO:0035434) |
| 0.0 | 0.3 | GO:0001759 | organ induction(GO:0001759) |
| 0.0 | 1.6 | GO:0007040 | lysosome organization(GO:0007040) lytic vacuole organization(GO:0080171) |
| 0.0 | 0.2 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.0 | 0.1 | GO:0038044 | transforming growth factor-beta secretion(GO:0038044) regulation of transforming growth factor-beta secretion(GO:2001201) |
| 0.0 | 0.1 | GO:0016081 | synaptic vesicle docking(GO:0016081) |
| 0.0 | 0.3 | GO:0051189 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.0 | 0.0 | GO:1903795 | regulation of inorganic anion transmembrane transport(GO:1903795) |
| 0.0 | 0.2 | GO:0043415 | positive regulation of skeletal muscle tissue regeneration(GO:0043415) |
| 0.0 | 0.1 | GO:0010046 | response to mycotoxin(GO:0010046) |
| 0.0 | 0.1 | GO:0035414 | negative regulation of catenin import into nucleus(GO:0035414) |
| 0.0 | 0.0 | GO:1990034 | calcium ion export from cell(GO:1990034) |
| 0.0 | 0.0 | GO:0071332 | cellular response to fructose stimulus(GO:0071332) |
| 0.0 | 0.2 | GO:0008626 | granzyme-mediated apoptotic signaling pathway(GO:0008626) |
| 0.0 | 0.2 | GO:0031441 | negative regulation of mRNA 3'-end processing(GO:0031441) |
| 0.0 | 0.2 | GO:0046487 | glyoxylate metabolic process(GO:0046487) |
| 0.0 | 0.0 | GO:0032808 | lacrimal gland development(GO:0032808) |
| 0.0 | 0.5 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.0 | 0.1 | GO:0002051 | osteoblast fate commitment(GO:0002051) |
| 0.0 | 0.1 | GO:0090074 | negative regulation of protein homodimerization activity(GO:0090074) |
| 0.0 | 0.6 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.1 | GO:0045590 | negative regulation of regulatory T cell differentiation(GO:0045590) |
| 0.0 | 0.2 | GO:0070314 | G1 to G0 transition(GO:0070314) |
| 0.0 | 0.0 | GO:0090271 | positive regulation of fibroblast growth factor production(GO:0090271) |
| 0.0 | 0.0 | GO:0060080 | inhibitory postsynaptic potential(GO:0060080) |
| 0.0 | 0.2 | GO:0021514 | ventral spinal cord interneuron differentiation(GO:0021514) |
| 0.0 | 0.3 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 0.4 | GO:0030828 | positive regulation of cGMP biosynthetic process(GO:0030828) |
| 0.0 | 0.1 | GO:1903026 | negative regulation of RNA polymerase II regulatory region sequence-specific DNA binding(GO:1903026) |
| 0.0 | 0.1 | GO:0038031 | non-canonical Wnt signaling pathway via JNK cascade(GO:0038031) |
| 0.0 | 0.0 | GO:0006537 | glutamate biosynthetic process(GO:0006537) |
| 0.0 | 0.4 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.0 | 0.5 | GO:0001658 | branching involved in ureteric bud morphogenesis(GO:0001658) |
| 0.0 | 0.0 | GO:0021819 | layer formation in cerebral cortex(GO:0021819) |
| 0.0 | 0.0 | GO:0072217 | negative regulation of metanephros development(GO:0072217) |
| 0.0 | 0.2 | GO:0051386 | regulation of neurotrophin TRK receptor signaling pathway(GO:0051386) |
| 0.0 | 0.0 | GO:0071288 | cellular response to mercury ion(GO:0071288) |
| 0.0 | 0.1 | GO:0010025 | wax biosynthetic process(GO:0010025) wax metabolic process(GO:0010166) |
| 0.0 | 0.1 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.0 | 1.7 | GO:0050808 | synapse organization(GO:0050808) |
| 0.0 | 0.0 | GO:0032741 | positive regulation of interleukin-18 production(GO:0032741) |
| 0.0 | 0.0 | GO:0045875 | negative regulation of sister chromatid cohesion(GO:0045875) |
| 0.0 | 0.3 | GO:0021889 | olfactory bulb interneuron differentiation(GO:0021889) |
| 0.0 | 0.0 | GO:1901490 | regulation of lymphangiogenesis(GO:1901490) |
| 0.0 | 0.1 | GO:0019336 | phenol-containing compound catabolic process(GO:0019336) |
| 0.0 | 0.2 | GO:0042256 | mature ribosome assembly(GO:0042256) |
| 0.0 | 0.4 | GO:0001919 | regulation of receptor recycling(GO:0001919) |
| 0.0 | 0.0 | GO:0045578 | negative regulation of B cell differentiation(GO:0045578) |
| 0.0 | 0.1 | GO:0001710 | mesodermal cell fate commitment(GO:0001710) |
| 0.0 | 0.0 | GO:0071871 | response to epinephrine(GO:0071871) cellular response to epinephrine stimulus(GO:0071872) |
| 0.0 | 1.2 | GO:0001505 | regulation of neurotransmitter levels(GO:0001505) |
| 0.0 | 0.1 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.0 | 1.5 | GO:0050911 | detection of chemical stimulus involved in sensory perception of smell(GO:0050911) |
| 0.0 | 0.3 | GO:0043651 | linoleic acid metabolic process(GO:0043651) |
| 0.0 | 0.5 | GO:1902187 | negative regulation of viral release from host cell(GO:1902187) |
| 0.0 | 0.1 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.0 | 0.2 | GO:0042487 | regulation of odontogenesis of dentin-containing tooth(GO:0042487) |
| 0.0 | 0.1 | GO:0021984 | adenohypophysis development(GO:0021984) |
| 0.0 | 0.1 | GO:0045760 | positive regulation of action potential(GO:0045760) |
| 0.0 | 0.5 | GO:0035088 | establishment or maintenance of apical/basal cell polarity(GO:0035088) establishment or maintenance of bipolar cell polarity(GO:0061245) |
| 0.0 | 0.0 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.0 | 0.0 | GO:0010579 | regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010578) positive regulation of adenylate cyclase activity involved in G-protein coupled receptor signaling pathway(GO:0010579) |
| 0.0 | 0.2 | GO:0042921 | glucocorticoid receptor signaling pathway(GO:0042921) |
| 0.0 | 0.1 | GO:0006295 | nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) |
| 0.0 | 1.0 | GO:0046426 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.0 | 0.2 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.0 | GO:0060447 | bud outgrowth involved in lung branching(GO:0060447) |
| 0.0 | 0.1 | GO:0000733 | DNA strand renaturation(GO:0000733) |
| 0.0 | 0.2 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.0 | 0.3 | GO:0016242 | negative regulation of macroautophagy(GO:0016242) |
| 0.0 | 0.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.0 | 0.1 | GO:0090370 | negative regulation of cholesterol efflux(GO:0090370) |
| 0.0 | 0.2 | GO:0006013 | mannose metabolic process(GO:0006013) |
| 0.0 | 0.0 | GO:0071873 | response to norepinephrine(GO:0071873) |
| 0.0 | 0.1 | GO:0010940 | positive regulation of necrotic cell death(GO:0010940) |
| 0.0 | 0.3 | GO:0010640 | regulation of platelet-derived growth factor receptor signaling pathway(GO:0010640) |
| 0.0 | 0.1 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.0 | 0.1 | GO:0060700 | regulation of ribonuclease activity(GO:0060700) |
| 0.0 | 0.0 | GO:0070428 | regulation of nucleotide-binding oligomerization domain containing 1 signaling pathway(GO:0070428) |
| 0.0 | 0.1 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.0 | 0.1 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.0 | 0.1 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.0 | 0.0 | GO:0070343 | white fat cell proliferation(GO:0070343) regulation of white fat cell proliferation(GO:0070350) |
| 0.0 | 0.1 | GO:0060967 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.0 | 0.1 | GO:0098915 | membrane repolarization during ventricular cardiac muscle cell action potential(GO:0098915) |
| 0.0 | 0.1 | GO:0070366 | regulation of hepatocyte differentiation(GO:0070366) |
| 0.0 | 0.2 | GO:0001696 | gastric acid secretion(GO:0001696) |
| 0.0 | 0.3 | GO:0090004 | positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.0 | 0.1 | GO:0002291 | T cell activation via T cell receptor contact with antigen bound to MHC molecule on antigen presenting cell(GO:0002291) |
| 0.0 | 0.2 | GO:0050667 | homocysteine metabolic process(GO:0050667) |
| 0.0 | 0.1 | GO:0030242 | pexophagy(GO:0030242) |
| 0.0 | 0.5 | GO:0007405 | neuroblast proliferation(GO:0007405) |
| 0.0 | 0.0 | GO:0045630 | positive regulation of T-helper 2 cell differentiation(GO:0045630) |
| 0.0 | 0.1 | GO:0002666 | positive regulation of T cell tolerance induction(GO:0002666) |
| 0.0 | 0.1 | GO:0014002 | astrocyte development(GO:0014002) |
| 0.0 | 0.1 | GO:1902714 | negative regulation of interferon-gamma secretion(GO:1902714) |
| 0.0 | 0.1 | GO:1901165 | positive regulation of trophoblast cell migration(GO:1901165) |
| 0.0 | 0.1 | GO:0006361 | transcription initiation from RNA polymerase I promoter(GO:0006361) |
| 0.0 | 0.1 | GO:1901421 | positive regulation of response to alcohol(GO:1901421) |
| 0.0 | 0.3 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.0 | 0.1 | GO:0071898 | regulation of estrogen receptor binding(GO:0071898) negative regulation of estrogen receptor binding(GO:0071899) |
| 0.0 | 0.0 | GO:0035878 | nail development(GO:0035878) |
| 0.0 | 0.1 | GO:0071732 | cellular response to nitric oxide(GO:0071732) |
| 0.0 | 0.1 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.0 | 0.0 | GO:0032226 | positive regulation of synaptic transmission, dopaminergic(GO:0032226) |
| 0.0 | 0.0 | GO:0060763 | mammary duct terminal end bud growth(GO:0060763) |
| 0.0 | 0.3 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.0 | 0.1 | GO:0015888 | thiamine transport(GO:0015888) |
| 0.0 | 0.1 | GO:0016480 | negative regulation of transcription from RNA polymerase III promoter(GO:0016480) |
| 0.0 | 0.0 | GO:0043416 | regulation of skeletal muscle tissue regeneration(GO:0043416) |
| 0.0 | 0.0 | GO:0070472 | regulation of uterine smooth muscle contraction(GO:0070472) positive regulation of uterine smooth muscle contraction(GO:0070474) |
| 0.0 | 0.2 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.0 | 0.3 | GO:0043113 | receptor clustering(GO:0043113) |
| 0.0 | 0.0 | GO:0015810 | aspartate transport(GO:0015810) |
| 0.0 | 0.3 | GO:0010839 | negative regulation of keratinocyte proliferation(GO:0010839) |
| 0.0 | 0.1 | GO:0071351 | cellular response to interleukin-18(GO:0071351) |
| 0.0 | 0.2 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.0 | 0.5 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.0 | 0.1 | GO:2001286 | regulation of caveolin-mediated endocytosis(GO:2001286) |
| 0.0 | 0.1 | GO:0070948 | regulation of neutrophil mediated cytotoxicity(GO:0070948) regulation of neutrophil mediated killing of symbiont cell(GO:0070949) |
| 0.0 | 0.9 | GO:0007566 | embryo implantation(GO:0007566) |
| 0.0 | 0.1 | GO:0002759 | regulation of antimicrobial humoral response(GO:0002759) |
| 0.0 | 0.2 | GO:0036120 | response to platelet-derived growth factor(GO:0036119) cellular response to platelet-derived growth factor stimulus(GO:0036120) |
| 0.0 | 0.1 | GO:0051342 | regulation of cyclic-nucleotide phosphodiesterase activity(GO:0051342) |
| 0.0 | 0.0 | GO:0033240 | positive regulation of cellular amine metabolic process(GO:0033240) |
| 0.0 | 0.0 | GO:0060084 | synaptic transmission involved in micturition(GO:0060084) |
| 0.0 | 0.0 | GO:2000321 | positive regulation of T-helper 17 cell differentiation(GO:2000321) |
| 0.0 | 0.0 | GO:0090598 | male genitalia morphogenesis(GO:0048808) male anatomical structure morphogenesis(GO:0090598) |
| 0.0 | 0.2 | GO:1903624 | regulation of DNA catabolic process(GO:1903624) |
| 0.0 | 0.0 | GO:0009912 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.0 | 0.1 | GO:0010637 | negative regulation of mitochondrial fusion(GO:0010637) |
| 0.0 | 0.0 | GO:0002378 | immunoglobulin biosynthetic process(GO:0002378) |
| 0.0 | 0.2 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.0 | 0.5 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.0 | 0.2 | GO:0006541 | glutamine metabolic process(GO:0006541) |
| 0.0 | 0.1 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.0 | 0.1 | GO:0071257 | cellular response to electrical stimulus(GO:0071257) |
| 0.0 | 0.0 | GO:0001880 | Mullerian duct regression(GO:0001880) |
| 0.0 | 0.1 | GO:0045717 | negative regulation of fatty acid biosynthetic process(GO:0045717) |
| 0.0 | 0.1 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.0 | 0.0 | GO:0061055 | myotome development(GO:0061055) |
| 0.0 | 0.0 | GO:0001827 | inner cell mass cell fate commitment(GO:0001827) |
| 0.0 | 0.0 | GO:1903671 | negative regulation of sprouting angiogenesis(GO:1903671) |
| 0.0 | 0.3 | GO:0043368 | positive T cell selection(GO:0043368) |
| 0.0 | 0.1 | GO:0003433 | chondrocyte development involved in endochondral bone morphogenesis(GO:0003433) |
| 0.0 | 0.0 | GO:0060689 | cell differentiation involved in salivary gland development(GO:0060689) |
| 0.0 | 0.0 | GO:2000646 | positive regulation of receptor catabolic process(GO:2000646) |
| 0.0 | 0.1 | GO:0035927 | RNA import into mitochondrion(GO:0035927) |
| 0.0 | 0.1 | GO:0032511 | late endosome to vacuole transport via multivesicular body sorting pathway(GO:0032511) protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 0.0 | 0.0 | GO:0043397 | corticotropin-releasing hormone secretion(GO:0043396) regulation of corticotropin-releasing hormone secretion(GO:0043397) |
| 0.0 | 0.0 | GO:1904528 | regulation of microtubule binding(GO:1904526) positive regulation of microtubule binding(GO:1904528) |
| 0.0 | 0.0 | GO:1900020 | regulation of protein kinase C activity(GO:1900019) positive regulation of protein kinase C activity(GO:1900020) |
| 0.0 | 0.0 | GO:0045988 | negative regulation of striated muscle contraction(GO:0045988) |
| 0.0 | 0.1 | GO:0072321 | chaperone-mediated protein transport(GO:0072321) |
| 0.0 | 0.2 | GO:0035774 | positive regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0035774) |
| 0.0 | 0.1 | GO:0016321 | female meiosis chromosome segregation(GO:0016321) |
| 0.0 | 0.2 | GO:0072600 | establishment of protein localization to Golgi(GO:0072600) |
| 0.0 | 0.0 | GO:0036476 | neuron death in response to hydrogen peroxide(GO:0036476) regulation of hydrogen peroxide-induced neuron death(GO:1903207) negative regulation of hydrogen peroxide-induced neuron death(GO:1903208) |
| 0.0 | 0.5 | GO:0031424 | keratinization(GO:0031424) |
| 0.0 | 0.1 | GO:0060023 | soft palate development(GO:0060023) |
| 0.0 | 0.1 | GO:0034227 | tRNA thio-modification(GO:0034227) |
| 0.0 | 0.0 | GO:0014827 | intestine smooth muscle contraction(GO:0014827) |
| 0.0 | 0.0 | GO:0034499 | late endosome to Golgi transport(GO:0034499) |
| 0.0 | 0.1 | GO:0042640 | anagen(GO:0042640) |
| 0.0 | 0.0 | GO:0007632 | visual behavior(GO:0007632) |
| 0.0 | 0.1 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.0 | 0.0 | GO:0035095 | behavioral response to nicotine(GO:0035095) |
| 0.0 | 0.0 | GO:0002277 | myeloid dendritic cell activation involved in immune response(GO:0002277) |
| 0.0 | 0.0 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) |
| 0.0 | 0.0 | GO:0021696 | cerebellar cortex morphogenesis(GO:0021696) |
| 0.0 | 0.0 | GO:0006681 | galactosylceramide metabolic process(GO:0006681) |
| 0.0 | 0.1 | GO:0043537 | negative regulation of blood vessel endothelial cell migration(GO:0043537) |
| 0.0 | 0.0 | GO:0042851 | L-alanine metabolic process(GO:0042851) |
| 0.0 | 0.0 | GO:1901620 | regulation of smoothened signaling pathway involved in dorsal/ventral neural tube patterning(GO:1901620) |
| 0.0 | 0.0 | GO:0010840 | regulation of circadian sleep/wake cycle, wakefulness(GO:0010840) circadian sleep/wake cycle, wakefulness(GO:0042746) |
| 0.0 | 0.1 | GO:0002325 | natural killer cell differentiation involved in immune response(GO:0002325) negative regulation of natural killer cell differentiation(GO:0032824) regulation of natural killer cell differentiation involved in immune response(GO:0032826) negative regulation of natural killer cell differentiation involved in immune response(GO:0032827) |
| 0.0 | 0.0 | GO:0032802 | low-density lipoprotein particle receptor catabolic process(GO:0032802) regulation of low-density lipoprotein particle receptor catabolic process(GO:0032803) |
| 0.0 | 0.2 | GO:0030204 | chondroitin sulfate metabolic process(GO:0030204) |
| 0.0 | 0.0 | GO:0035542 | regulation of SNARE complex assembly(GO:0035542) |
| 0.0 | 0.2 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.0 | 0.0 | GO:0046532 | regulation of photoreceptor cell differentiation(GO:0046532) |
| 0.0 | 0.1 | GO:0046116 | queuosine biosynthetic process(GO:0008616) queuosine metabolic process(GO:0046116) |
| 0.0 | 0.2 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.0 | 0.0 | GO:0035754 | B cell chemotaxis(GO:0035754) |
| 0.0 | 0.2 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.0 | 0.1 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.0 | 0.0 | GO:0035582 | sequestering of BMP in extracellular matrix(GO:0035582) |
| 0.0 | 0.7 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 0.0 | GO:0071985 | multivesicular body sorting pathway(GO:0071985) |
| 0.0 | 0.0 | GO:1901856 | negative regulation of cellular respiration(GO:1901856) |
| 0.0 | 0.0 | GO:0042231 | interleukin-13 biosynthetic process(GO:0042231) |
| 0.0 | 0.1 | GO:0002591 | positive regulation of antigen processing and presentation of peptide antigen via MHC class I(GO:0002591) |
| 0.0 | 0.0 | GO:1902445 | regulation of mitochondrial membrane permeability involved in programmed necrotic cell death(GO:1902445) |
| 0.0 | 0.1 | GO:1904294 | positive regulation of ERAD pathway(GO:1904294) |
| 0.0 | 0.0 | GO:0046499 | S-adenosylmethioninamine metabolic process(GO:0046499) |
| 0.0 | 0.0 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.0 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.0 | 0.0 | GO:0061668 | mitochondrial ribosome assembly(GO:0061668) |
| 0.0 | 0.1 | GO:0061099 | negative regulation of protein tyrosine kinase activity(GO:0061099) |
| 0.0 | 0.0 | GO:0016103 | diterpenoid catabolic process(GO:0016103) retinoic acid catabolic process(GO:0034653) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 7.4 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 1.3 | 10.5 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 1.0 | 7.2 | GO:0044300 | cerebellar mossy fiber(GO:0044300) |
| 0.9 | 8.5 | GO:0005883 | neurofilament(GO:0005883) |
| 0.9 | 7.6 | GO:0071437 | invadopodium(GO:0071437) |
| 0.9 | 0.9 | GO:0044294 | dendritic growth cone(GO:0044294) |
| 0.8 | 2.5 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.8 | 7.5 | GO:0044224 | juxtaparanode region of axon(GO:0044224) |
| 0.8 | 2.5 | GO:0014701 | junctional sarcoplasmic reticulum membrane(GO:0014701) |
| 0.8 | 9.1 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) |
| 0.7 | 5.0 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.7 | 4.1 | GO:0005915 | zonula adherens(GO:0005915) |
| 0.7 | 6.7 | GO:0030673 | axolemma(GO:0030673) |
| 0.7 | 5.3 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.6 | 12.3 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.6 | 3.2 | GO:0097433 | dense body(GO:0097433) |
| 0.6 | 29.4 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.6 | 1.8 | GO:0034750 | Scrib-APC-beta-catenin complex(GO:0034750) |
| 0.6 | 4.3 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.5 | 2.1 | GO:0032541 | cortical endoplasmic reticulum(GO:0032541) |
| 0.5 | 2.4 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.5 | 3.3 | GO:0036057 | filtration diaphragm(GO:0036056) slit diaphragm(GO:0036057) |
| 0.5 | 1.4 | GO:0072534 | perineuronal net(GO:0072534) |
| 0.4 | 10.5 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.4 | 3.0 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.4 | 12.6 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.4 | 12.5 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.4 | 6.0 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.4 | 5.5 | GO:0031527 | filopodium membrane(GO:0031527) |
| 0.4 | 12.1 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.4 | 5.4 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.4 | 3.5 | GO:0097449 | astrocyte projection(GO:0097449) |
| 0.3 | 3.2 | GO:0031235 | intrinsic component of the cytoplasmic side of the plasma membrane(GO:0031235) |
| 0.3 | 2.9 | GO:0043194 | axon initial segment(GO:0043194) |
| 0.3 | 7.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.3 | 3.8 | GO:0031045 | dense core granule(GO:0031045) |
| 0.3 | 0.3 | GO:0070033 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin II complex(GO:0070033) |
| 0.3 | 0.6 | GO:0097441 | basilar dendrite(GO:0097441) |
| 0.3 | 0.3 | GO:0035838 | growing cell tip(GO:0035838) |
| 0.3 | 43.3 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.3 | 3.8 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.3 | 2.3 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.3 | 2.5 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.3 | 1.1 | GO:1990696 | USH2 complex(GO:1990696) |
| 0.3 | 23.2 | GO:0030427 | site of polarized growth(GO:0030427) |
| 0.3 | 8.0 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.2 | 2.2 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.2 | 0.5 | GO:0000836 | ER ubiquitin ligase complex(GO:0000835) Hrd1p ubiquitin ligase complex(GO:0000836) |
| 0.2 | 2.7 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.2 | 1.6 | GO:0032590 | dendrite membrane(GO:0032590) |
| 0.2 | 1.4 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.2 | 0.9 | GO:0061574 | ASAP complex(GO:0061574) |
| 0.2 | 0.7 | GO:0051286 | cell tip(GO:0051286) |
| 0.2 | 17.3 | GO:0043204 | perikaryon(GO:0043204) |
| 0.2 | 3.7 | GO:0032809 | neuronal cell body membrane(GO:0032809) |
| 0.2 | 0.6 | GO:0032937 | SREBP-SCAP-Insig complex(GO:0032937) |
| 0.2 | 0.4 | GO:0033647 | host intracellular organelle(GO:0033647) host intracellular membrane-bounded organelle(GO:0033648) |
| 0.2 | 1.4 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.2 | 0.2 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.2 | 0.6 | GO:0034706 | sodium channel complex(GO:0034706) |
| 0.2 | 1.1 | GO:0070688 | MLL5-L complex(GO:0070688) |
| 0.2 | 0.7 | GO:0070852 | cell body fiber(GO:0070852) |
| 0.2 | 1.2 | GO:0042622 | photoreceptor outer segment membrane(GO:0042622) |
| 0.1 | 0.9 | GO:0001674 | female germ cell nucleus(GO:0001674) |
| 0.1 | 1.3 | GO:0045334 | clathrin-coated endocytic vesicle(GO:0045334) |
| 0.1 | 6.2 | GO:0008076 | voltage-gated potassium channel complex(GO:0008076) |
| 0.1 | 0.8 | GO:0019907 | cyclin-dependent protein kinase activating kinase holoenzyme complex(GO:0019907) |
| 0.1 | 5.3 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.1 | 0.5 | GO:1990716 | axonemal central apparatus(GO:1990716) |
| 0.1 | 0.6 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 7.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.1 | 0.4 | GO:0031983 | vesicle lumen(GO:0031983) |
| 0.1 | 1.5 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.1 | 0.8 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.1 | 0.1 | GO:0044393 | microspike(GO:0044393) |
| 0.1 | 0.3 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 0.2 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.1 | 0.5 | GO:0044292 | dendrite terminus(GO:0044292) |
| 0.1 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.1 | 0.3 | GO:0044530 | supraspliceosomal complex(GO:0044530) |
| 0.1 | 0.3 | GO:1990393 | 3M complex(GO:1990393) |
| 0.1 | 0.5 | GO:0002177 | manchette(GO:0002177) |
| 0.1 | 0.7 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.1 | 0.5 | GO:0001940 | male pronucleus(GO:0001940) |
| 0.1 | 0.3 | GO:0043203 | axon hillock(GO:0043203) |
| 0.1 | 0.4 | GO:0033270 | paranode region of axon(GO:0033270) |
| 0.1 | 0.7 | GO:0005687 | U4 snRNP(GO:0005687) |
| 0.1 | 0.7 | GO:0031011 | Ino80 complex(GO:0031011) |
| 0.1 | 0.1 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.1 | 6.3 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.1 | 0.3 | GO:1990851 | Wnt-Frizzled-LRP5/6 complex(GO:1990851) |
| 0.1 | 4.9 | GO:0070382 | exocytic vesicle(GO:0070382) |
| 0.1 | 1.4 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 7.6 | GO:0030425 | dendrite(GO:0030425) |
| 0.1 | 0.5 | GO:0008074 | guanylate cyclase complex, soluble(GO:0008074) |
| 0.1 | 0.7 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 0.7 | GO:0030123 | AP-3 adaptor complex(GO:0030123) |
| 0.1 | 0.8 | GO:0043205 | fibril(GO:0043205) |
| 0.1 | 1.0 | GO:0030904 | retromer complex(GO:0030904) |
| 0.1 | 0.1 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.0 | 2.3 | GO:0035097 | histone methyltransferase complex(GO:0035097) |
| 0.0 | 0.1 | GO:0035985 | senescence-associated heterochromatin focus(GO:0035985) |
| 0.0 | 1.7 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 0.1 | GO:0016589 | NURF complex(GO:0016589) |
| 0.0 | 0.1 | GO:0043527 | tRNA methyltransferase complex(GO:0043527) |
| 0.0 | 0.3 | GO:0034464 | BBSome(GO:0034464) |
| 0.0 | 1.1 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.5 | GO:0042405 | nuclear inclusion body(GO:0042405) |
| 0.0 | 0.1 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.0 | 0.4 | GO:0071006 | U2-type catalytic step 1 spliceosome(GO:0071006) |
| 0.0 | 0.1 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.1 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.0 | 0.2 | GO:0032389 | MutLalpha complex(GO:0032389) |
| 0.0 | 0.1 | GO:0044326 | dendritic spine neck(GO:0044326) |
| 0.0 | 0.4 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.0 | 0.1 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.0 | 0.1 | GO:0032585 | multivesicular body membrane(GO:0032585) |
| 0.0 | 0.5 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 0.3 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.0 | 0.8 | GO:0005720 | nuclear heterochromatin(GO:0005720) |
| 0.0 | 0.1 | GO:0005767 | secondary lysosome(GO:0005767) |
| 0.0 | 0.1 | GO:0045098 | type III intermediate filament(GO:0045098) |
| 0.0 | 0.2 | GO:0071546 | pi-body(GO:0071546) |
| 0.0 | 0.1 | GO:0044352 | pinosome(GO:0044352) macropinosome(GO:0044354) |
| 0.0 | 0.3 | GO:0042613 | MHC class II protein complex(GO:0042613) |
| 0.0 | 0.1 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.0 | 0.1 | GO:0016012 | dystroglycan complex(GO:0016011) sarcoglycan complex(GO:0016012) |
| 0.0 | 0.9 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.2 | GO:0005677 | chromatin silencing complex(GO:0005677) |
| 0.0 | 0.1 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.0 | 0.2 | GO:0030008 | TRAPP complex(GO:0030008) |
| 0.0 | 0.1 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 0.1 | GO:0043202 | lysosomal lumen(GO:0043202) |
| 0.0 | 0.2 | GO:0043198 | dendritic shaft(GO:0043198) |
| 0.0 | 0.1 | GO:0045293 | mRNA editing complex(GO:0045293) |
| 0.0 | 5.1 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.0 | GO:0071598 | neuronal ribonucleoprotein granule(GO:0071598) |
| 0.0 | 0.0 | GO:0042827 | platelet dense granule(GO:0042827) |
| 0.0 | 0.1 | GO:0005594 | collagen type IX trimer(GO:0005594) |
| 0.0 | 0.1 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.0 | 0.1 | GO:0097629 | extrinsic component of omegasome membrane(GO:0097629) |
| 0.0 | 0.0 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 0.3 | GO:0005797 | Golgi medial cisterna(GO:0005797) |
| 0.0 | 0.1 | GO:0043293 | apoptosome(GO:0043293) |
| 0.0 | 0.0 | GO:0044298 | cell body membrane(GO:0044298) |
| 0.0 | 0.3 | GO:0005922 | connexon complex(GO:0005922) |
| 0.0 | 0.0 | GO:0000125 | PCAF complex(GO:0000125) |
| 0.0 | 0.0 | GO:0000235 | astral microtubule(GO:0000235) |
| 0.0 | 0.0 | GO:0034705 | potassium channel complex(GO:0034705) |
| 0.0 | 0.1 | GO:0042571 | immunoglobulin complex, circulating(GO:0042571) |
| 0.0 | 0.0 | GO:0097386 | glial cell projection(GO:0097386) |
| 0.0 | 0.1 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 1.3 | GO:0035068 | micro-ribonucleoprotein complex(GO:0035068) |
| 0.0 | 0.2 | GO:0016581 | NuRD complex(GO:0016581) CHD-type complex(GO:0090545) |
| 0.0 | 0.1 | GO:0035631 | CD40 receptor complex(GO:0035631) |
| 0.0 | 0.1 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.0 | 0.0 | GO:0000802 | transverse filament(GO:0000802) |
| 0.0 | 0.0 | GO:0031084 | BLOC-2 complex(GO:0031084) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 10.7 | GO:0005519 | cytoskeletal regulatory protein binding(GO:0005519) |
| 2.9 | 5.8 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) |
| 2.7 | 8.1 | GO:0097109 | neuroligin family protein binding(GO:0097109) |
| 2.5 | 12.7 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 1.8 | 7.0 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 1.7 | 5.0 | GO:0005176 | ErbB-2 class receptor binding(GO:0005176) |
| 1.4 | 6.8 | GO:0005095 | GTPase inhibitor activity(GO:0005095) |
| 1.3 | 3.8 | GO:0001075 | transcription factor activity, RNA polymerase II core promoter sequence-specific binding involved in preinitiation complex assembly(GO:0001075) |
| 1.3 | 5.1 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 1.2 | 7.3 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 1.2 | 7.1 | GO:0016933 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 1.1 | 3.2 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 1.0 | 5.1 | GO:0008454 | alpha-1,3-mannosylglycoprotein 4-beta-N-acetylglucosaminyltransferase activity(GO:0008454) |
| 1.0 | 6.8 | GO:0005004 | GPI-linked ephrin receptor activity(GO:0005004) |
| 1.0 | 2.9 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.9 | 21.6 | GO:0071837 | HMG box domain binding(GO:0071837) |
| 0.9 | 3.7 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.9 | 10.7 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) |
| 0.9 | 3.5 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.8 | 3.3 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.8 | 8.5 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.8 | 6.9 | GO:0001091 | RNA polymerase II basal transcription factor binding(GO:0001091) |
| 0.8 | 2.3 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.7 | 2.9 | GO:0003846 | 2-acylglycerol O-acyltransferase activity(GO:0003846) |
| 0.7 | 3.6 | GO:0008499 | UDP-galactose:beta-N-acetylglucosamine beta-1,3-galactosyltransferase activity(GO:0008499) |
| 0.6 | 1.8 | GO:0050816 | phosphothreonine binding(GO:0050816) |
| 0.6 | 2.3 | GO:0034056 | estrogen response element binding(GO:0034056) |
| 0.6 | 1.1 | GO:0005237 | inhibitory extracellular ligand-gated ion channel activity(GO:0005237) |
| 0.6 | 1.7 | GO:1990430 | extracellular matrix protein binding(GO:1990430) |
| 0.5 | 3.8 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.5 | 2.2 | GO:0005294 | neutral L-amino acid secondary active transmembrane transporter activity(GO:0005294) |
| 0.5 | 4.3 | GO:0098988 | G-protein coupled glutamate receptor activity(GO:0098988) |
| 0.5 | 2.2 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.5 | 10.2 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.5 | 0.5 | GO:0031711 | bradykinin receptor binding(GO:0031711) |
| 0.5 | 1.5 | GO:0043423 | 3-phosphoinositide-dependent protein kinase binding(GO:0043423) |
| 0.5 | 3.0 | GO:0001588 | dopamine neurotransmitter receptor activity, coupled via Gs(GO:0001588) |
| 0.5 | 2.0 | GO:0005280 | hydrogen:amino acid symporter activity(GO:0005280) |
| 0.5 | 3.8 | GO:0002162 | dystroglycan binding(GO:0002162) |
| 0.5 | 1.4 | GO:0004691 | cAMP-dependent protein kinase activity(GO:0004691) |
| 0.5 | 2.3 | GO:0000155 | phosphorelay sensor kinase activity(GO:0000155) |
| 0.5 | 1.4 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) |
| 0.4 | 10.7 | GO:0017091 | AU-rich element binding(GO:0017091) |
| 0.4 | 3.9 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.4 | 0.4 | GO:0031871 | proteinase activated receptor binding(GO:0031871) |
| 0.4 | 8.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.4 | 1.3 | GO:0030144 | alpha-1,6-mannosylglycoprotein 6-beta-N-acetylglucosaminyltransferase activity(GO:0030144) |
| 0.4 | 2.5 | GO:0031749 | D2 dopamine receptor binding(GO:0031749) |
| 0.4 | 1.3 | GO:0005502 | 11-cis retinal binding(GO:0005502) |
| 0.4 | 7.1 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.4 | 0.8 | GO:0005167 | neurotrophin TRK receptor binding(GO:0005167) |
| 0.4 | 1.2 | GO:0048101 | calcium- and calmodulin-regulated 3',5'-cyclic-GMP phosphodiesterase activity(GO:0048101) |
| 0.4 | 3.0 | GO:0008599 | protein phosphatase type 1 regulator activity(GO:0008599) |
| 0.4 | 1.5 | GO:0035727 | lysophosphatidic acid binding(GO:0035727) |
| 0.4 | 0.7 | GO:0035939 | microsatellite binding(GO:0035939) |
| 0.4 | 6.1 | GO:0004435 | phosphatidylinositol phospholipase C activity(GO:0004435) |
| 0.4 | 8.9 | GO:0001106 | RNA polymerase II transcription corepressor activity(GO:0001106) |
| 0.4 | 3.5 | GO:0051378 | serotonin binding(GO:0051378) |
| 0.4 | 1.1 | GO:0097108 | hedgehog family protein binding(GO:0097108) |
| 0.3 | 1.4 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.3 | 2.6 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) |
| 0.3 | 3.5 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.3 | 1.6 | GO:0086080 | protein binding involved in heterotypic cell-cell adhesion(GO:0086080) |
| 0.3 | 7.8 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.3 | 5.3 | GO:0008327 | methyl-CpG binding(GO:0008327) |
| 0.3 | 4.2 | GO:0045295 | gamma-catenin binding(GO:0045295) |
| 0.3 | 1.2 | GO:0019871 | sodium channel inhibitor activity(GO:0019871) |
| 0.3 | 2.9 | GO:0070300 | phosphatidic acid binding(GO:0070300) |
| 0.3 | 4.8 | GO:0035255 | ionotropic glutamate receptor binding(GO:0035255) |
| 0.3 | 0.8 | GO:0098821 | BMP receptor activity(GO:0098821) |
| 0.3 | 0.3 | GO:0004517 | nitric-oxide synthase activity(GO:0004517) |
| 0.3 | 3.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.3 | 2.9 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.3 | 1.8 | GO:0043495 | protein anchor(GO:0043495) |
| 0.3 | 0.8 | GO:0000309 | nicotinamide-nucleotide adenylyltransferase activity(GO:0000309) nicotinate-nucleotide adenylyltransferase activity(GO:0004515) |
| 0.3 | 3.1 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.3 | 5.4 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.3 | 2.6 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.3 | 5.9 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.3 | 0.8 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.3 | 1.0 | GO:0032896 | palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 0.3 | 0.8 | GO:0004937 | alpha1-adrenergic receptor activity(GO:0004937) |
| 0.2 | 1.5 | GO:0015198 | oligopeptide transporter activity(GO:0015198) |
| 0.2 | 2.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.2 | 0.5 | GO:0035373 | chondroitin sulfate proteoglycan binding(GO:0035373) |
| 0.2 | 1.2 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.2 | 1.2 | GO:0001601 | peptide YY receptor activity(GO:0001601) |
| 0.2 | 0.9 | GO:0016286 | small conductance calcium-activated potassium channel activity(GO:0016286) |
| 0.2 | 0.7 | GO:0015180 | L-alanine transmembrane transporter activity(GO:0015180) alanine transmembrane transporter activity(GO:0022858) |
| 0.2 | 1.1 | GO:0030550 | acetylcholine receptor inhibitor activity(GO:0030550) |
| 0.2 | 0.7 | GO:0030899 | calcium-dependent ATPase activity(GO:0030899) |
| 0.2 | 4.9 | GO:0045499 | chemorepellent activity(GO:0045499) |
| 0.2 | 0.7 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.2 | 0.9 | GO:0019992 | diacylglycerol binding(GO:0019992) |
| 0.2 | 0.9 | GO:0015016 | [heparan sulfate]-glucosamine N-sulfotransferase activity(GO:0015016) |
| 0.2 | 1.3 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.2 | 0.6 | GO:0004471 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) |
| 0.2 | 1.0 | GO:0004945 | angiotensin type II receptor activity(GO:0004945) |
| 0.2 | 1.0 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.2 | 0.6 | GO:0001226 | RNA polymerase II transcription corepressor binding(GO:0001226) |
| 0.2 | 0.8 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.2 | 1.0 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.2 | 2.4 | GO:0031005 | filamin binding(GO:0031005) |
| 0.2 | 0.4 | GO:0070679 | inositol 1,4,5 trisphosphate binding(GO:0070679) |
| 0.2 | 0.7 | GO:0000983 | transcription factor activity, RNA polymerase II core promoter sequence-specific(GO:0000983) |
| 0.2 | 0.7 | GO:0008934 | inositol monophosphate 1-phosphatase activity(GO:0008934) inositol monophosphate 3-phosphatase activity(GO:0052832) inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.2 | 6.7 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.2 | 4.8 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.2 | 1.5 | GO:0035198 | miRNA binding(GO:0035198) |
| 0.2 | 3.3 | GO:0005112 | Notch binding(GO:0005112) |
| 0.2 | 0.9 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.2 | 0.5 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.2 | 2.7 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.2 | 0.4 | GO:0030284 | estrogen receptor activity(GO:0030284) |
| 0.2 | 1.4 | GO:0097157 | pre-mRNA intronic binding(GO:0097157) |
| 0.2 | 0.7 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.2 | 0.3 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.2 | 1.7 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.2 | 0.5 | GO:0042392 | sphingosine-1-phosphate phosphatase activity(GO:0042392) |
| 0.2 | 5.7 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.2 | 0.5 | GO:0016649 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.2 | 0.6 | GO:0030280 | structural constituent of epidermis(GO:0030280) |
| 0.2 | 0.2 | GO:0008035 | high-density lipoprotein particle binding(GO:0008035) |
| 0.2 | 0.5 | GO:0008900 | hydrogen:potassium-exchanging ATPase activity(GO:0008900) |
| 0.2 | 0.8 | GO:0004111 | creatine kinase activity(GO:0004111) |
| 0.2 | 0.6 | GO:0034190 | apolipoprotein receptor binding(GO:0034190) |
| 0.2 | 0.8 | GO:0008294 | calcium- and calmodulin-responsive adenylate cyclase activity(GO:0008294) |
| 0.2 | 0.6 | GO:0017162 | aryl hydrocarbon receptor binding(GO:0017162) |
| 0.2 | 0.5 | GO:0016716 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, another compound as one donor, and incorporation of one atom of oxygen(GO:0016716) |
| 0.2 | 0.5 | GO:0050692 | DBD domain binding(GO:0050692) |
| 0.1 | 0.7 | GO:0003847 | 1-alkyl-2-acetylglycerophosphocholine esterase activity(GO:0003847) |
| 0.1 | 0.4 | GO:0004995 | tachykinin receptor activity(GO:0004995) |
| 0.1 | 1.0 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) |
| 0.1 | 0.1 | GO:0016775 | phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.1 | 0.7 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 0.1 | 2.4 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.1 | 2.7 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.1 | 0.6 | GO:0047391 | alkylglycerophosphoethanolamine phosphodiesterase activity(GO:0047391) |
| 0.1 | 0.7 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 0.6 | GO:0008113 | peptide-methionine (S)-S-oxide reductase activity(GO:0008113) |
| 0.1 | 0.3 | GO:2001070 | starch binding(GO:2001070) |
| 0.1 | 0.1 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 0.4 | GO:0050510 | N-acetylgalactosaminyl-proteoglycan 3-beta-glucuronosyltransferase activity(GO:0050510) |
| 0.1 | 2.3 | GO:0031489 | myosin V binding(GO:0031489) |
| 0.1 | 0.4 | GO:0031800 | type 3 metabotropic glutamate receptor binding(GO:0031800) |
| 0.1 | 0.7 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.1 | 0.3 | GO:0015018 | galactosylgalactosylxylosylprotein 3-beta-glucuronosyltransferase activity(GO:0015018) |
| 0.1 | 0.8 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.1 | 0.5 | GO:0008486 | diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) |
| 0.1 | 0.8 | GO:0004908 | interleukin-1 receptor activity(GO:0004908) |
| 0.1 | 0.8 | GO:0036122 | BMP binding(GO:0036122) |
| 0.1 | 7.8 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 0.4 | GO:0016917 | GABA receptor activity(GO:0016917) |
| 0.1 | 0.6 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.1 | 3.5 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 0.2 | GO:0004886 | 9-cis retinoic acid receptor activity(GO:0004886) |
| 0.1 | 0.5 | GO:0019145 | aminobutyraldehyde dehydrogenase activity(GO:0019145) 4-trimethylammoniobutyraldehyde dehydrogenase activity(GO:0047105) |
| 0.1 | 1.2 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.1 | 1.1 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.1 | 0.2 | GO:0000403 | Y-form DNA binding(GO:0000403) |
| 0.1 | 0.2 | GO:0044020 | histone methyltransferase activity (H4-R3 specific)(GO:0044020) |
| 0.1 | 0.8 | GO:0034819 | 3-(3-hydroxyphenyl)propionate hydroxylase activity(GO:0008688) 4-chlorobenzaldehyde oxidase activity(GO:0018471) 3,5-xylenol methylhydroxylase activity(GO:0018630) phenylacetate hydroxylase activity(GO:0018631) 4-nitrophenol 4-monooxygenase activity(GO:0018632) dimethyl sulfide monooxygenase activity(GO:0018633) alpha-pinene monooxygenase [NADH] activity(GO:0018634) 1-hydroxy-2-naphthoate hydroxylase activity(GO:0018637) toluene 4-monooxygenase activity(GO:0018638) xylene monooxygenase activity(GO:0018639) dibenzothiophene monooxygenase activity(GO:0018640) 6-hydroxy-3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018641) chlorophenol 4-monooxygenase activity(GO:0018642) carbon disulfide oxygenase activity(GO:0018643) toluene 2-monooxygenase activity(GO:0018644) 1-hydroxy-2-oxolimonene 1,2-monooxygenase activity(GO:0018646) phenanthrene 1,2-monooxygenase activity(GO:0018647) tetrahydrofuran hydroxylase activity(GO:0018649) styrene monooxygenase activity(GO:0018650) toluene-4-sulfonate monooxygenase activity(GO:0018651) toluene-sulfonate methyl-monooxygenase activity(GO:0018652) 3-methyl-2-oxo-1,2-dihydroquinoline 6-monooxygenase activity(GO:0018653) 2-hydroxy-phenylacetate hydroxylase activity(GO:0018654) 2-oxo-delta3-4,5,5-trimethylcyclopentenylacetyl-CoA 1,2-monooxygenase activity(GO:0018655) phenanthrene 3,4-monooxygenase activity(GO:0018656) toluene 3-monooxygenase activity(GO:0018657) 4-hydroxyphenylacetate,NADH:oxygen oxidoreductase (3-hydroxylating) activity(GO:0018660) limonene monooxygenase activity(GO:0019113) 2-methylnaphthalene hydroxylase activity(GO:0034526) 1-methylnaphthalene hydroxylase activity(GO:0034534) bisphenol A hydroxylase A activity(GO:0034560) salicylate 5-hydroxylase activity(GO:0034785) isobutylamine N-hydroxylase activity(GO:0034791) branched-chain dodecylbenzene sulfonate monooxygenase activity(GO:0034802) 3-HSA hydroxylase activity(GO:0034819) 4-hydroxypyridine-3-hydroxylase activity(GO:0034894) 2-octaprenyl-3-methyl-6-methoxy-1,4-benzoquinol hydroxylase activity(GO:0043719) 6-hydroxynicotinate 3-monooxygenase activity(GO:0043731) thalianol hydroxylase activity(GO:0080014) |
| 0.1 | 0.6 | GO:0031849 | olfactory receptor binding(GO:0031849) |
| 0.1 | 1.6 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.1 | 0.3 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.1 | 0.8 | GO:0043522 | leucine zipper domain binding(GO:0043522) |
| 0.1 | 0.6 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.1 | 0.1 | GO:0004690 | cyclic nucleotide-dependent protein kinase activity(GO:0004690) |
| 0.1 | 0.5 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.1 | 0.3 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.1 | 0.9 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.1 | 0.2 | GO:0042166 | acetylcholine binding(GO:0042166) |
| 0.1 | 1.5 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.1 | 0.9 | GO:0008066 | glutamate receptor activity(GO:0008066) |
| 0.1 | 0.6 | GO:0016421 | CoA carboxylase activity(GO:0016421) |
| 0.1 | 0.4 | GO:0099528 | G-protein coupled acetylcholine receptor activity(GO:0016907) G-protein coupled neurotransmitter receptor activity(GO:0099528) |
| 0.1 | 0.8 | GO:0009881 | photoreceptor activity(GO:0009881) |
| 0.1 | 1.8 | GO:0032452 | histone demethylase activity(GO:0032452) |
| 0.1 | 0.3 | GO:1990254 | keratin filament binding(GO:1990254) |
| 0.1 | 0.3 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.1 | 0.3 | GO:0004667 | prostaglandin-D synthase activity(GO:0004667) |
| 0.1 | 0.1 | GO:0031750 | D3 dopamine receptor binding(GO:0031750) |
| 0.1 | 0.4 | GO:0071614 | linoleic acid epoxygenase activity(GO:0071614) |
| 0.1 | 0.5 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 0.3 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.1 | 0.2 | GO:0002151 | G-quadruplex RNA binding(GO:0002151) |
| 0.1 | 0.3 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.1 | 0.3 | GO:0004924 | oncostatin-M receptor activity(GO:0004924) |
| 0.1 | 0.5 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.1 | 0.4 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.1 | 0.8 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 0.4 | GO:0030957 | Tat protein binding(GO:0030957) |
| 0.1 | 1.2 | GO:0042800 | histone methyltransferase activity (H3-K4 specific)(GO:0042800) |
| 0.1 | 2.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 2.0 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 0.2 | GO:0005030 | neurotrophin receptor activity(GO:0005030) |
| 0.1 | 0.3 | GO:0070051 | fibrinogen binding(GO:0070051) |
| 0.1 | 0.9 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 0.2 | GO:0000386 | second spliceosomal transesterification activity(GO:0000386) |
| 0.1 | 0.2 | GO:0030297 | transmembrane receptor protein tyrosine kinase activator activity(GO:0030297) |
| 0.1 | 0.5 | GO:0016670 | oxidoreductase activity, acting on a sulfur group of donors, oxygen as acceptor(GO:0016670) |
| 0.1 | 0.3 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 0.1 | GO:0004629 | phospholipase C activity(GO:0004629) |
| 0.1 | 1.2 | GO:0008266 | poly(U) RNA binding(GO:0008266) |
| 0.1 | 0.3 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.1 | 0.4 | GO:0030275 | LRR domain binding(GO:0030275) |
| 0.1 | 1.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 0.1 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.1 | 0.1 | GO:0031893 | vasopressin receptor binding(GO:0031893) |
| 0.1 | 0.2 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.1 | 0.1 | GO:0051425 | PTB domain binding(GO:0051425) |
| 0.1 | 1.5 | GO:0042054 | histone methyltransferase activity(GO:0042054) |
| 0.1 | 0.2 | GO:0010314 | phosphatidylinositol-5-phosphate binding(GO:0010314) |
| 0.1 | 0.5 | GO:0008889 | glycerophosphodiester phosphodiesterase activity(GO:0008889) |
| 0.1 | 0.3 | GO:0047499 | calcium-independent phospholipase A2 activity(GO:0047499) |
| 0.1 | 0.8 | GO:0043422 | protein kinase B binding(GO:0043422) |
| 0.1 | 0.1 | GO:0001515 | opioid peptide activity(GO:0001515) |
| 0.1 | 0.2 | GO:0030250 | cyclase activator activity(GO:0010853) guanylate cyclase activator activity(GO:0030250) |
| 0.1 | 0.3 | GO:0004969 | histamine receptor activity(GO:0004969) |
| 0.1 | 0.5 | GO:0031701 | angiotensin receptor binding(GO:0031701) |
| 0.1 | 0.1 | GO:0016015 | morphogen activity(GO:0016015) |
| 0.1 | 0.1 | GO:0097603 | temperature-gated ion channel activity(GO:0097603) |
| 0.1 | 1.1 | GO:0036002 | pre-mRNA binding(GO:0036002) |
| 0.1 | 0.3 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.1 | 0.4 | GO:0043855 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.1 | 0.4 | GO:0001206 | transcriptional repressor activity, RNA polymerase II distal enhancer sequence-specific binding(GO:0001206) |
| 0.1 | 2.3 | GO:0005262 | calcium channel activity(GO:0005262) |
| 0.1 | 4.1 | GO:0004860 | protein kinase inhibitor activity(GO:0004860) |
| 0.1 | 0.3 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.1 | 0.2 | GO:1990226 | histone methyltransferase binding(GO:1990226) |
| 0.1 | 0.5 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 0.2 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.1 | 0.3 | GO:0005272 | sodium channel activity(GO:0005272) |
| 0.1 | 0.1 | GO:0070538 | oleic acid binding(GO:0070538) |
| 0.1 | 0.3 | GO:0070742 | C2H2 zinc finger domain binding(GO:0070742) |
| 0.1 | 0.1 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.1 | 0.2 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.1 | 0.3 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.1 | 0.3 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.1 | 0.3 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.1 | 0.6 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 0.6 | GO:0048018 | receptor agonist activity(GO:0048018) |
| 0.1 | 43.0 | GO:0043565 | sequence-specific DNA binding(GO:0043565) |
| 0.1 | 0.1 | GO:0043175 | RNA polymerase core enzyme binding(GO:0043175) |
| 0.1 | 3.6 | GO:0044212 | transcription regulatory region DNA binding(GO:0044212) |
| 0.1 | 0.2 | GO:0003941 | L-serine ammonia-lyase activity(GO:0003941) |
| 0.1 | 0.2 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 0.3 | GO:0008517 | folic acid transporter activity(GO:0008517) |
| 0.1 | 0.4 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.0 | 0.8 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.2 | GO:0004370 | glycerol kinase activity(GO:0004370) |
| 0.0 | 0.0 | GO:0048408 | epidermal growth factor binding(GO:0048408) |
| 0.0 | 0.6 | GO:0042608 | T cell receptor binding(GO:0042608) |
| 0.0 | 0.2 | GO:0099589 | G-protein coupled serotonin receptor activity(GO:0004993) serotonin receptor activity(GO:0099589) |
| 0.0 | 0.2 | GO:0030229 | very-low-density lipoprotein particle receptor activity(GO:0030229) |
| 0.0 | 0.2 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.0 | 0.1 | GO:0031821 | G-protein coupled serotonin receptor binding(GO:0031821) |
| 0.0 | 0.5 | GO:0001618 | virus receptor activity(GO:0001618) |
| 0.0 | 0.9 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 20.5 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 0.6 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.0 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.0 | 0.0 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.0 | 0.1 | GO:0016901 | oxidoreductase activity, acting on the CH-OH group of donors, quinone or similar compound as acceptor(GO:0016901) |
| 0.0 | 0.4 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.0 | 0.2 | GO:0034713 | type I transforming growth factor beta receptor binding(GO:0034713) |
| 0.0 | 0.2 | GO:0001077 | transcriptional activator activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001077) |
| 0.0 | 0.2 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.0 | 0.4 | GO:0015643 | toxic substance binding(GO:0015643) |
| 0.0 | 0.1 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.0 | 0.1 | GO:0004427 | inorganic diphosphatase activity(GO:0004427) |
| 0.0 | 1.4 | GO:0034483 | heparan sulfate sulfotransferase activity(GO:0034483) |
| 0.0 | 0.2 | GO:0035014 | phosphatidylinositol 3-kinase regulator activity(GO:0035014) |
| 0.0 | 0.2 | GO:0033691 | sialic acid binding(GO:0033691) |
| 0.0 | 2.3 | GO:0005261 | cation channel activity(GO:0005261) |
| 0.0 | 0.1 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.0 | 0.4 | GO:0019203 | carbohydrate phosphatase activity(GO:0019203) |
| 0.0 | 0.1 | GO:0030345 | extracellular matrix structural constituent conferring compression resistance(GO:0030021) structural constituent of tooth enamel(GO:0030345) |
| 0.0 | 0.1 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.0 | 0.6 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.1 | GO:0004800 | thyroxine 5'-deiodinase activity(GO:0004800) |
| 0.0 | 0.1 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.1 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.0 | 0.2 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.0 | GO:0005416 | cation:amino acid symporter activity(GO:0005416) |
| 0.0 | 0.7 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.0 | 3.9 | GO:0003729 | mRNA binding(GO:0003729) |
| 0.0 | 0.1 | GO:0070883 | pre-miRNA binding(GO:0070883) |
| 0.0 | 0.6 | GO:0005086 | ARF guanyl-nucleotide exchange factor activity(GO:0005086) |
| 0.0 | 0.2 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 0.2 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.0 | 0.1 | GO:0051185 | coenzyme transporter activity(GO:0051185) |
| 0.0 | 0.1 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 0.3 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.1 | GO:0019166 | trans-2-enoyl-CoA reductase (NADPH) activity(GO:0019166) |
| 0.0 | 0.5 | GO:1990003 | JUN kinase phosphatase activity(GO:0008579) phosphohistidine phosphatase activity(GO:0008969) inositol-1,3,4-trisphosphate 4-phosphatase activity(GO:0017161) NADP phosphatase activity(GO:0019178) inositol-4,5-bisphosphate 5-phosphatase activity(GO:0030487) 5-amino-6-(5-phosphoribitylamino)uracil phosphatase activity(GO:0043726) phosphatidylinositol-3,5-bisphosphate 5-phosphatase activity(GO:0043813) inositol-1,3,4,5,6-pentakisphosphate 1-phosphatase activity(GO:0052825) inositol-3,4-bisphosphate 4-phosphatase activity(GO:0052828) inositol-1,3,4-trisphosphate 1-phosphatase activity(GO:0052829) inositol-1,3,4,6-tetrakisphosphate 6-phosphatase activity(GO:0052830) inositol-1,3,4,6-tetrakisphosphate 1-phosphatase activity(GO:0052831) phosphatidylinositol-1,4,5-trisphosphate 5-phosphatase activity(GO:0052867) IDP phosphatase activity(GO:1990003) |
| 0.0 | 0.1 | GO:0004165 | dodecenoyl-CoA delta-isomerase activity(GO:0004165) |
| 0.0 | 0.0 | GO:0015172 | acidic amino acid transmembrane transporter activity(GO:0015172) |
| 0.0 | 0.2 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.3 | GO:0001730 | 2'-5'-oligoadenylate synthetase activity(GO:0001730) |
| 0.0 | 0.1 | GO:0042284 | sphingolipid delta-4 desaturase activity(GO:0042284) |
| 0.0 | 0.1 | GO:0004982 | N-formyl peptide receptor activity(GO:0004982) |
| 0.0 | 0.4 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 0.0 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 0.0 | GO:0034988 | Fc-gamma receptor I complex binding(GO:0034988) |
| 0.0 | 0.1 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.2 | GO:0003707 | steroid hormone receptor activity(GO:0003707) |
| 0.0 | 0.0 | GO:0034040 | lipid-transporting ATPase activity(GO:0034040) |
| 0.0 | 0.1 | GO:0001849 | complement component C1q binding(GO:0001849) |
| 0.0 | 0.1 | GO:0050051 | leukotriene-B4 20-monooxygenase activity(GO:0050051) |
| 0.0 | 0.1 | GO:0004946 | bombesin receptor activity(GO:0004946) |
| 0.0 | 0.5 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.1 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.0 | 0.0 | GO:0016769 | transferase activity, transferring nitrogenous groups(GO:0016769) |
| 0.0 | 0.1 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 0.0 | 0.3 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.0 | 0.2 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 3.3 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.0 | 0.0 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.0 | 0.5 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.3 | GO:0005186 | pheromone activity(GO:0005186) |
| 0.0 | 0.1 | GO:0004505 | phenylalanine 4-monooxygenase activity(GO:0004505) |
| 0.0 | 0.1 | GO:0035514 | DNA demethylase activity(GO:0035514) |
| 0.0 | 0.0 | GO:0042165 | neurotransmitter binding(GO:0042165) |
| 0.0 | 0.0 | GO:0050694 | galactose 3-O-sulfotransferase activity(GO:0050694) |
| 0.0 | 0.2 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.0 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.0 | GO:0070815 | procollagen-lysine 5-dioxygenase activity(GO:0008475) peptidyl-lysine 5-dioxygenase activity(GO:0070815) |
| 0.0 | 0.0 | GO:0004676 | 3-phosphoinositide-dependent protein kinase activity(GO:0004676) |
| 0.0 | 0.1 | GO:0033695 | oxidoreductase activity, acting on CH or CH2 groups, quinone or similar compound as acceptor(GO:0033695) caffeine oxidase activity(GO:0034875) |
| 0.0 | 0.2 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 0.2 | GO:0030332 | cyclin binding(GO:0030332) |
| 0.0 | 0.3 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.1 | GO:0016019 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 0.1 | GO:0050649 | testosterone 6-beta-hydroxylase activity(GO:0050649) |
| 0.0 | 0.1 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.0 | GO:0008107 | galactoside 2-alpha-L-fucosyltransferase activity(GO:0008107) alpha-(1,2)-fucosyltransferase activity(GO:0031127) |
| 0.0 | 0.1 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.0 | 0.1 | GO:0008479 | queuine tRNA-ribosyltransferase activity(GO:0008479) |
| 0.0 | 0.0 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.3 | GO:0033038 | bitter taste receptor activity(GO:0033038) |
| 0.0 | 0.1 | GO:0016920 | pyroglutamyl-peptidase activity(GO:0016920) |
| 0.0 | 0.0 | GO:0015563 | uptake transmembrane transporter activity(GO:0015563) |
| 0.0 | 0.0 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 10.1 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.4 | 0.4 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.4 | 9.5 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.3 | 8.5 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.2 | 5.4 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.2 | 3.7 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.2 | 5.0 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.2 | 2.7 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.2 | 3.7 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.2 | 5.2 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.2 | 7.1 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.2 | 1.6 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 3.4 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 1.7 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
| 0.1 | 5.3 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 6.6 | PID NOTCH PATHWAY | Notch signaling pathway |
| 0.1 | 5.4 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.1 | 2.2 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.1 | 1.3 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 1.7 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.1 | 1.1 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 0.9 | ST G ALPHA I PATHWAY | G alpha i Pathway |
| 0.1 | 0.5 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.1 | 1.0 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.1 | 4.3 | PID BETA CATENIN NUC PATHWAY | Regulation of nuclear beta catenin signaling and target gene transcription |
| 0.1 | 1.5 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.4 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.1 | 1.7 | SIG REGULATION OF THE ACTIN CYTOSKELETON BY RHO GTPASES | Genes related to regulation of the actin cytoskeleton |
| 0.1 | 2.5 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 0.1 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.1 | 0.9 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.1 | 0.7 | ST DIFFERENTIATION PATHWAY IN PC12 CELLS | Differentiation Pathway in PC12 Cells; this is a specific case of PAC1 Receptor Pathway. |
| 0.1 | 0.1 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 0.3 | PID ANTHRAX PATHWAY | Cellular roles of Anthrax toxin |
| 0.1 | 2.3 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.1 | 0.2 | PID BETA CATENIN DEG PATHWAY | Degradation of beta catenin |
| 0.1 | 1.0 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.0 | 0.1 | SA G1 AND S PHASES | Cdk2, 4, and 6 bind cyclin D in G1, while cdk2/cyclin E promotes the G1/S transition. |
| 0.0 | 1.0 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 0.5 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.0 | 0.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 0.5 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 0.5 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.0 | 0.2 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.0 | 0.8 | PID CDC42 PATHWAY | CDC42 signaling events |
| 0.0 | 0.0 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.0 | 0.0 | PID TCPTP PATHWAY | Signaling events mediated by TCPTP |
| 0.0 | 0.4 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.2 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.0 | 0.2 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.3 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.0 | 0.1 | ST IL 13 PATHWAY | Interleukin 13 (IL-13) Pathway |
| 0.0 | 0.0 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.5 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.2 | PID AMB2 NEUTROPHILS PATHWAY | amb2 Integrin signaling |
| 0.0 | 0.0 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 10.4 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.7 | 7.5 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.7 | 9.9 | REACTOME UNBLOCKING OF NMDA RECEPTOR GLUTAMATE BINDING AND ACTIVATION | Genes involved in Unblocking of NMDA receptor, glutamate binding and activation |
| 0.6 | 14.8 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.6 | 12.6 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.6 | 4.1 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.6 | 12.0 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.5 | 5.9 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.5 | 5.9 | REACTOME DOPAMINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Dopamine Neurotransmitter Release Cycle |
| 0.5 | 6.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.5 | 4.1 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.5 | 4.5 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.4 | 8.5 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.4 | 5.5 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.4 | 11.7 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.3 | 0.3 | REACTOME REGULATION OF INSULIN SECRETION | Genes involved in Regulation of Insulin Secretion |
| 0.3 | 6.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.3 | 5.7 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.3 | 3.2 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.3 | 6.1 | REACTOME SIGNALING BY ROBO RECEPTOR | Genes involved in Signaling by Robo receptor |
| 0.3 | 0.3 | REACTOME FGFR1 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR1 ligand binding and activation |
| 0.3 | 0.8 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.2 | 0.5 | REACTOME NOREPINEPHRINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Norepinephrine Neurotransmitter Release Cycle |
| 0.2 | 2.8 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.2 | 2.3 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.2 | 2.2 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.2 | 8.1 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.2 | 0.2 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.2 | 3.1 | REACTOME ACTIVATED POINT MUTANTS OF FGFR2 | Genes involved in Activated point mutants of FGFR2 |
| 0.2 | 2.2 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.2 | 0.2 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.2 | 0.2 | REACTOME CREB PHOSPHORYLATION THROUGH THE ACTIVATION OF CAMKII | Genes involved in CREB phosphorylation through the activation of CaMKII |
| 0.2 | 2.6 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.2 | 6.4 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.2 | 0.6 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.2 | 1.2 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.1 | 0.1 | REACTOME SIGNALING BY ERBB2 | Genes involved in Signaling by ERBB2 |
| 0.1 | 0.1 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.1 | 0.4 | REACTOME GLUTAMATE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Glutamate Neurotransmitter Release Cycle |
| 0.1 | 1.2 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.1 | 1.4 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.1 | 0.5 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.1 | 1.7 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |
| 0.1 | 2.3 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.1 | 2.6 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 5.1 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.1 | 1.1 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.1 | 2.6 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.1 | 1.2 | REACTOME TANDEM PORE DOMAIN POTASSIUM CHANNELS | Genes involved in Tandem pore domain potassium channels |
| 0.1 | 0.1 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.1 | 2.7 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 0.1 | REACTOME P130CAS LINKAGE TO MAPK SIGNALING FOR INTEGRINS | Genes involved in p130Cas linkage to MAPK signaling for integrins |
| 0.1 | 0.1 | REACTOME ADP SIGNALLING THROUGH P2RY12 | Genes involved in ADP signalling through P2Y purinoceptor 12 |
| 0.1 | 1.5 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.1 | 1.6 | REACTOME GLUCAGON TYPE LIGAND RECEPTORS | Genes involved in Glucagon-type ligand receptors |
| 0.1 | 0.5 | REACTOME REGULATION OF BETA CELL DEVELOPMENT | Genes involved in Regulation of beta-cell development |
| 0.1 | 0.8 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 1.2 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 0.1 | REACTOME CLASS A1 RHODOPSIN LIKE RECEPTORS | Genes involved in Class A/1 (Rhodopsin-like receptors) |
| 0.1 | 0.1 | REACTOME TRANSCRIPTION | Genes involved in Transcription |
| 0.1 | 0.1 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.1 | 0.9 | REACTOME TRANSMISSION ACROSS CHEMICAL SYNAPSES | Genes involved in Transmission across Chemical Synapses |
| 0.1 | 0.6 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.1 | 1.2 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.1 | 0.7 | REACTOME POST NMDA RECEPTOR ACTIVATION EVENTS | Genes involved in Post NMDA receptor activation events |
| 0.1 | 1.1 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 0.7 | REACTOME TRAFFICKING OF AMPA RECEPTORS | Genes involved in Trafficking of AMPA receptors |
| 0.1 | 0.1 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 0.1 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 0.5 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 1.4 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.0 | 0.7 | REACTOME BRANCHED CHAIN AMINO ACID CATABOLISM | Genes involved in Branched-chain amino acid catabolism |
| 0.0 | 0.0 | REACTOME NEF MEDIATES DOWN MODULATION OF CELL SURFACE RECEPTORS BY RECRUITING THEM TO CLATHRIN ADAPTERS | Genes involved in Nef-mediates down modulation of cell surface receptors by recruiting them to clathrin adapters |
| 0.0 | 0.0 | REACTOME AXON GUIDANCE | Genes involved in Axon guidance |
| 0.0 | 0.0 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.0 | 0.8 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.4 | REACTOME HIGHLY CALCIUM PERMEABLE POSTSYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Highly calcium permeable postsynaptic nicotinic acetylcholine receptors |
| 0.0 | 0.0 | REACTOME JNK C JUN KINASES PHOSPHORYLATION AND ACTIVATION MEDIATED BY ACTIVATED HUMAN TAK1 | Genes involved in JNK (c-Jun kinases) phosphorylation and activation mediated by activated human TAK1 |
| 0.0 | 0.2 | REACTOME FORMATION OF TRANSCRIPTION COUPLED NER TC NER REPAIR COMPLEX | Genes involved in Formation of transcription-coupled NER (TC-NER) repair complex |
| 0.0 | 0.4 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.7 | REACTOME DEGRADATION OF THE EXTRACELLULAR MATRIX | Genes involved in Degradation of the extracellular matrix |
| 0.0 | 0.9 | REACTOME SIGNALING BY NOTCH1 | Genes involved in Signaling by NOTCH1 |
| 0.0 | 0.3 | REACTOME GAP JUNCTION ASSEMBLY | Genes involved in Gap junction assembly |
| 0.0 | 0.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.0 | 0.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.4 | REACTOME G ALPHA S SIGNALLING EVENTS | Genes involved in G alpha (s) signalling events |
| 0.0 | 0.1 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.1 | REACTOME INCRETIN SYNTHESIS SECRETION AND INACTIVATION | Genes involved in Incretin Synthesis, Secretion, and Inactivation |
| 0.0 | 0.1 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.0 | 2.1 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.0 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.0 | 0.0 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 0.0 | 0.0 | REACTOME BILE ACID AND BILE SALT METABOLISM | Genes involved in Bile acid and bile salt metabolism |
| 0.0 | 0.1 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.1 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |